

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
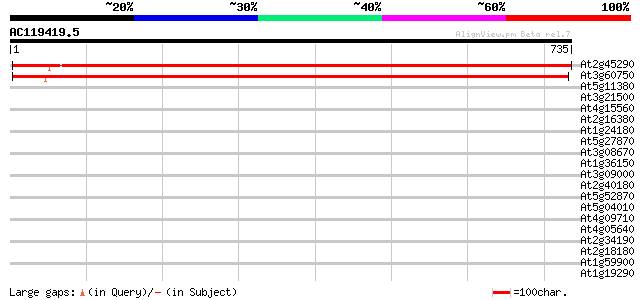
Query= AC119419.5 - phase: 0
(735 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g45290 putative transketolase precursor 1191 0.0
At3g60750 transketolase - like protein 1176 0.0
At5g11380 1-D-deoxyxylulose 5-phosphate synthase - like protein 41 0.002
At3g21500 1-D-deoxyxylulose 5-phosphate synthase, putative 41 0.002
At4g15560 1-D-deoxyxylulose 5-phosphate synthase, putative 41 0.003
At2g16380 putative phosphatidylinositol/phosphatidylcholine tran... 33 0.44
At1g24180 pyruvate dehydrogenase E1 alpha subunit 33 0.75
At5g27870 pectin methyl-esterase - like protein 31 2.8
At3g08670 hypothetical protein 31 2.8
At1g36150 hypothetical protein 31 2.8
At3g09000 unknown protein 30 3.7
At2g40180 protein phosphatase 2C (AthPP2C5) 30 3.7
At5g52870 unknown protein (At5g52870) 30 4.8
At5g04010 putative protein 30 4.8
At4g09710 RNA-directed DNA polymerase -like protein 30 4.8
At4g05640 putative protein 30 4.8
At2g34190 putative membrane transporter 30 4.8
At2g18180 putative phosphatidylinositol/phophatidylcholine trans... 30 4.8
At1g59900 pyruvate dehydrogenase E1 alpha subunit, putative 30 4.8
At1g19290 hypothetical protein, 5' partial 30 6.3
>At2g45290 putative transketolase precursor
Length = 741
Score = 1191 bits (3080), Expect = 0.0
Identities = 586/741 (79%), Positives = 643/741 (86%), Gaps = 10/741 (1%)
Query: 4 SSSTSLYLTNPHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQS---------HHLR 54
+S++SL L+ LTR S + S+P+ L T T+ S H LR
Sbjct: 2 ASTSSLALSQALLTRAISHNGSENCVSIPAFSALKSTSPRTSGTISSRRRNASTISHSLR 61
Query: 55 HTAIHASVSAPPSTTTDSSLVEKSVNTIRFLAVDSVEKANSGHPGLPMGCAPMGHVLYDE 114
A+V A T++DSSLV+KSVNTIRFLA+D+VEKA SGHPGLPMGCAPM H+LYDE
Sbjct: 62 PLVRAAAVEAIV-TSSDSSLVDKSVNTIRFLAIDAVEKAKSGHPGLPMGCAPMSHILYDE 120
Query: 115 VMRYNPKNPFWFNRDRFVLSAGHGCMLQYALLHLAGYDSVKVEDLKQFRQWESRTPGHPE 174
VM+YNPKNP+WFNRDRFVLSAGHGCMLQYALLHLAGYDSV+ EDLK FRQW S+TPGHPE
Sbjct: 121 VMKYNPKNPYWFNRDRFVLSAGHGCMLQYALLHLAGYDSVREEDLKSFRQWGSKTPGHPE 180
Query: 175 NFETPGIEVTTGPLGQGIANAVGLALAEKHLAARFNKPDNEIIDHYTYCILGDGCQMEGI 234
NFETPG+E TTGPLGQGIANAVGLALAEKHLAARFNKPDNEI+DHYTY ILGDGCQMEGI
Sbjct: 181 NFETPGVEATTGPLGQGIANAVGLALAEKHLAARFNKPDNEIVDHYTYSILGDGCQMEGI 240
Query: 235 ANEACSLAGHWGLGKLIAFYDDNHISIDGDTEIAFTESVEKRFEGLGWHVIWVKNGNTGF 294
+NE CSLAGHWGLGKLIAFYDDNHISIDGDT+IAFTESV+KRFE LGWHVIWVKNGN G+
Sbjct: 241 SNEVCSLAGHWGLGKLIAFYDDNHISIDGDTDIAFTESVDKRFEALGWHVIWVKNGNNGY 300
Query: 295 DEIRAAIKEAKAVKDRPTLIKFTTTIGYGSPNKSNSYSVHGSALGAKEVDATRNNLGWPY 354
DEIRAAI+EAKAV D+PTLIK TTTIGYGSPNK+NSYSVHG+ALG KEV+ATRNNLGWPY
Sbjct: 301 DEIRAAIREAKAVTDKPTLIKVTTTIGYGSPNKANSYSVHGAALGEKEVEATRNNLGWPY 360
Query: 355 EPFHVPEDVKKHWSRHIREGAALESEWNAKFADYEKKYKEEAAVLKSIISGDLPAGWEKA 414
EPFHVPEDVK HWSRH EGAALE++WNAKFA YEKKY EEAA LKSIISG+LP GWEKA
Sbjct: 361 EPFHVPEDVKSHWSRHTPEGAALEADWNAKFAAYEKKYPEEAAELKSIISGELPVGWEKA 420
Query: 415 LPTYTPEIPADATRNLSQQNLNALANVLPGLIGGSADLASSNMTLLKKFGDFQSDSPAER 474
LPTYTP+ P DATRNLSQQ LNALA +PG +GGSADLASSNMT+LK FG+FQ +P ER
Sbjct: 421 LPTYTPDSPGDATRNLSQQCLNALAKAVPGFLGGSADLASSNMTMLKAFGNFQKATPEER 480
Query: 475 NVRFGVREHGMGAICNGIALHSRGLIPYCATFFVFTDYMRGAIRLSALSEAGVIYVMTHD 534
N+RFGVREHGMGAICNGIALHS G IPYCATFFVFTDYMR A+R+SALSEAGVIYVMTHD
Sbjct: 481 NLRFGVREHGMGAICNGIALHSPGFIPYCATFFVFTDYMRAAMRISALSEAGVIYVMTHD 540
Query: 535 SIGLGEDGPTHQPIEHLASFRAMPNILMLRPADGNETAGAYKVAVLNRKRPSILALSRQK 594
SIGLGEDGPTHQPIEHL+SFRAMPNI+M RPADGNETAGAYK+AV RK PS+LALSRQK
Sbjct: 541 SIGLGEDGPTHQPIEHLSSFRAMPNIMMFRPADGNETAGAYKIAVTKRKTPSVLALSRQK 600
Query: 595 LPNLAGTSIEGVEKGGYIVSDNSTGNKPDVILIGTGSELEIAYKAGEDLRKEGKAVRVVS 654
LP L GTSIE VEKGGY +SDNSTGNKPDVILIGTGSELEIA +A E LR++GK+VRVVS
Sbjct: 601 LPQLPGTSIESVEKGGYTISDNSTGNKPDVILIGTGSELEIAAQAAEKLREQGKSVRVVS 660
Query: 655 FVSWELFDEQSQAYKESVLPTAVTARVSIEAGSTFGWEKIVGSKGKAIGIDRFGASAPAG 714
FV WELFDEQS AYKESVLP+ V+ARVSIEAGSTFGW KIVG KGK+IGID FGASAPAG
Sbjct: 661 FVCWELFDEQSDAYKESVLPSDVSARVSIEAGSTFGWGKIVGGKGKSIGIDTFGASAPAG 720
Query: 715 TIYKEFGITKEAVIAAAKELI 735
+YKEFGIT EA++ AAK LI
Sbjct: 721 KLYKEFGITIEAMVEAAKSLI 741
>At3g60750 transketolase - like protein
Length = 741
Score = 1176 bits (3041), Expect = 0.0
Identities = 580/738 (78%), Positives = 641/738 (86%), Gaps = 10/738 (1%)
Query: 4 SSSTSLYLTNPHLTRHTSSPSTTRLSSLPSTITLNRTPTPTT---------HLSQSHHLR 54
+S++SL L+ L R S + + SLP+ L T + + ++++ LR
Sbjct: 2 ASTSSLALSQALLARAISHHGSDQRGSLPAFSGLKSTGSRASASSRRRIAQSMTKNRSLR 61
Query: 55 HTAIHASVSAPPSTTTDSSLVEKSVNTIRFLAVDSVEKANSGHPGLPMGCAPMGHVLYDE 114
+ A+ TTDSS+V+KSVN+IRFLA+D+VEKA SGHPGLPMGCAPM H+LYDE
Sbjct: 62 -PLVRAAAVETVEPTTDSSIVDKSVNSIRFLAIDAVEKAKSGHPGLPMGCAPMAHILYDE 120
Query: 115 VMRYNPKNPFWFNRDRFVLSAGHGCMLQYALLHLAGYDSVKVEDLKQFRQWESRTPGHPE 174
VMRYNPKNP+WFNRDRFVLSAGHGCML YALLHLAGYDSV+ EDLKQFRQW S+TPGHPE
Sbjct: 121 VMRYNPKNPYWFNRDRFVLSAGHGCMLLYALLHLAGYDSVQEEDLKQFRQWGSKTPGHPE 180
Query: 175 NFETPGIEVTTGPLGQGIANAVGLALAEKHLAARFNKPDNEIIDHYTYCILGDGCQMEGI 234
NFETPGIEVTTGPLGQGIANAVGLALAEKHLAARFNKPD E++DHYTY ILGDGCQMEGI
Sbjct: 181 NFETPGIEVTTGPLGQGIANAVGLALAEKHLAARFNKPDAEVVDHYTYAILGDGCQMEGI 240
Query: 235 ANEACSLAGHWGLGKLIAFYDDNHISIDGDTEIAFTESVEKRFEGLGWHVIWVKNGNTGF 294
+NEACSLAGHWGLGKLIAFYDDNHISIDGDTEIAFTE+V++RFE LGWHVIWVKNGNTG+
Sbjct: 241 SNEACSLAGHWGLGKLIAFYDDNHISIDGDTEIAFTENVDQRFEALGWHVIWVKNGNTGY 300
Query: 295 DEIRAAIKEAKAVKDRPTLIKFTTTIGYGSPNKSNSYSVHGSALGAKEVDATRNNLGWPY 354
DEIRAAIKEAK V D+PTLIK TTTIGYGSPNK+NSYSVHG+ALG KEV+ATRNNLGWPY
Sbjct: 301 DEIRAAIKEAKTVTDKPTLIKVTTTIGYGSPNKANSYSVHGAALGEKEVEATRNNLGWPY 360
Query: 355 EPFHVPEDVKKHWSRHIREGAALESEWNAKFADYEKKYKEEAAVLKSIISGDLPAGWEKA 414
EPF VP+DVK HWSRH EGA LES+W+AKFA YEKKY EEA+ LKSII+G+LPAGWEKA
Sbjct: 361 EPFQVPDDVKSHWSRHTPEGATLESDWSAKFAAYEKKYPEEASELKSIITGELPAGWEKA 420
Query: 415 LPTYTPEIPADATRNLSQQNLNALANVLPGLIGGSADLASSNMTLLKKFGDFQSDSPAER 474
LPTYTPE P DATRNLSQQ LNALA V+PG +GGSADLASSNMTLLK FGDFQ +P ER
Sbjct: 421 LPTYTPESPGDATRNLSQQCLNALAKVVPGFLGGSADLASSNMTLLKAFGDFQKATPEER 480
Query: 475 NVRFGVREHGMGAICNGIALHSRGLIPYCATFFVFTDYMRGAIRLSALSEAGVIYVMTHD 534
N+RFGVREHGMGAICNGIALHS GLIPYCATFFVFTDYMRGA+R+SALSEAGVIYVMTHD
Sbjct: 481 NLRFGVREHGMGAICNGIALHSPGLIPYCATFFVFTDYMRGAMRISALSEAGVIYVMTHD 540
Query: 535 SIGLGEDGPTHQPIEHLASFRAMPNILMLRPADGNETAGAYKVAVLNRKRPSILALSRQK 594
SIGLGEDGPTHQPIEH+ASFRAMPN LM RPADGNETAGAYK+AV RK PSILALSRQK
Sbjct: 541 SIGLGEDGPTHQPIEHIASFRAMPNTLMFRPADGNETAGAYKIAVTKRKTPSILALSRQK 600
Query: 595 LPNLAGTSIEGVEKGGYIVSDNSTGNKPDVILIGTGSELEIAYKAGEDLRKEGKAVRVVS 654
LP+L GTSIEGVEKGGY +SD+S+GNKPDVILIGTGSELEIA +A E LRK+GK VRVVS
Sbjct: 601 LPHLPGTSIEGVEKGGYTISDDSSGNKPDVILIGTGSELEIAAQAAEVLRKDGKTVRVVS 660
Query: 655 FVSWELFDEQSQAYKESVLPTAVTARVSIEAGSTFGWEKIVGSKGKAIGIDRFGASAPAG 714
FV WELFDEQS YKESVLP+ V+ARVSIEA STFGW KIVG KGK+IGI+ FGASAPA
Sbjct: 661 FVCWELFDEQSDEYKESVLPSDVSARVSIEAASTFGWGKIVGGKGKSIGINSFGASAPAP 720
Query: 715 TIYKEFGITKEAVIAAAK 732
+YKEFGIT EAV+ AAK
Sbjct: 721 LLYKEFGITVEAVVDAAK 738
>At5g11380 1-D-deoxyxylulose 5-phosphate synthase - like protein
Length = 700
Score = 41.2 bits (95), Expect = 0.002
Identities = 46/182 (25%), Positives = 69/182 (37%), Gaps = 34/182 (18%)
Query: 487 AICNGIALHSRGLIPYCATFFVF-----------TDYMRGAIRLSALSEAGVIYVMTHDS 535
A+ L S GL P+C F D R A+R ++ AG++
Sbjct: 434 AVTFSAGLSSGGLKPFCIIPSAFLQRAYDQVVHDVDRQRKAVRF-VITSAGLV------- 485
Query: 536 IGLGEDGPTHQPIEHLASFRAMPNILMLRPADGNETAGAYKVAVLNRKRPSILALSRQKL 595
G DGP +A ++PN++ + PAD +E A RP R +
Sbjct: 486 ---GSDGPVQCGAFDIAFMSSLPNMIAMAPADEDELVNMVATAAYVTDRPVCFRFPRGSI 542
Query: 596 PNL-----AGTSIEGVEKGGYIVSDNSTGNKPDVILIGTGSELEIAYKAGEDLRKEGKAV 650
N+ G IE + +G +V DV L+G G+ ++ A L K G V
Sbjct: 543 VNMNYLVPTGLPIE-IGRGRVLVEGQ------DVALLGYGAMVQNCLHAHSLLSKLGLNV 595
Query: 651 RV 652
V
Sbjct: 596 TV 597
>At3g21500 1-D-deoxyxylulose 5-phosphate synthase, putative
Length = 628
Score = 41.2 bits (95), Expect = 0.002
Identities = 49/198 (24%), Positives = 79/198 (39%), Gaps = 26/198 (13%)
Query: 466 FQSDSPAERNVRFGVREHGMGAICNGIALHSRGLIPYCATFFVFTDYMRGA----IRLSA 521
F+S P R G+ E G+A GL P+C +++ +M+ A +
Sbjct: 380 FESRFPT-RCFDVGIAEQHAVTFAAGLACE--GLKPFCT---IYSSFMQRAYDQVVHDVD 433
Query: 522 LSEAGVIYVMTHDSIGLGEDGPTHQPIEHLASFRAMPNILMLRPADGNETAGAYKVAVLN 581
L + V + + + +G DGPTH + +PN++++ P+D E A
Sbjct: 434 LQKLPVRFAIDRAGL-MGADGPTHCGAFDVTFMACLPNMIVMAPSDEAELFNMVATAAAI 492
Query: 582 RKRPSILALSRQKLPNLAGTSIEGVEKG-------GYIVSDNSTGNKPDVILIGTGSELE 634
RPS R N G S+ KG G I+ D V L+G GS ++
Sbjct: 493 DDRPSCFRYHR---GNGIGVSLPPGNKGVPLQIGRGRILRDGER-----VALLGYGSAVQ 544
Query: 635 IAYKAGEDLRKEGKAVRV 652
+A L + G + V
Sbjct: 545 RCLEAASMLSERGLKITV 562
Score = 36.6 bits (83), Expect = 0.052
Identities = 75/338 (22%), Positives = 124/338 (36%), Gaps = 79/338 (23%)
Query: 21 SSPSTTRLSSLPSTITL-------NRTPTPTTHLSQSHHLRHTAIHASVSAPPSTTTDSS 73
SS S+T+ S + +T NR PTP L T H P + S
Sbjct: 24 SSMSSTKYSKVRATTFSEKGEYYSNRPPTP---------LLDTINH------PMHMKNLS 68
Query: 74 LVEKSV--NTIRFLAVDSVEKANSGHPGLPMGCAPMGHVLYDEVMRYNPKNPFWFN--RD 129
+ E V + +R + +V K GH G +G + L+ + FN D
Sbjct: 69 IKELKVLSDELRSDVIFNVSKTG-GHLGSNLGVVELTVALH-----------YIFNTPHD 116
Query: 130 RFVLSAGHGCMLQYALLHLAGYDSVKVEDLKQFRQWESRTPGHPENFETPGIEVTTGPLG 189
+ + GH L G K++ ++Q T ++ G ++ L
Sbjct: 117 KILWDVGHQSYPHKILTGRRG----KMKTIRQTNGLSGYTKRRESEHDSFGTGHSSTTLS 172
Query: 190 QGIANAVGLALAEKHLAARFNKPDNEIIDHYTYCILGDGCQMEGIANEACSLAGHWGLGK 249
G+ AVG L + N ++ ++GDG G A EA + AG+
Sbjct: 173 AGLGMAVGRDLKGMN---------NSVVS-----VIGDGAMTAGQAYEAMNNAGYLHSNM 218
Query: 250 LIAFYDDNHIS-----IDGDTEIA----------------FTESVEKRFEGLGWHVIWVK 288
++ D+ +S +DG T+ E+ FE LG+H +
Sbjct: 219 IVILNDNKQVSLPTANLDGPTQPVGALSCALSRLQSNCGMIRETSSTLFEELGFHYVGPV 278
Query: 289 NGNTGFDEIRAAIKEAKAVKD-RPTLIKFTTTIGYGSP 325
+G+ D++ + ++ K+ K P LI T G G P
Sbjct: 279 DGH-NIDDLVSILETLKSTKTIGPVLIHVVTEKGRGYP 315
>At4g15560 1-D-deoxyxylulose 5-phosphate synthase, putative
Length = 717
Score = 40.8 bits (94), Expect = 0.003
Identities = 47/178 (26%), Positives = 73/178 (40%), Gaps = 25/178 (14%)
Query: 487 AICNGIALHSRGLIPYCATFFVFTDYMRGA----IRLSALSEAGVIYVMTHDSIGL-GED 541
A+ L GL P+CA +++ +M+ A + L + V + M D GL G D
Sbjct: 453 AVTFAAGLACEGLKPFCA---IYSSFMQRAYDQVVHDVDLQKLPVRFAM--DRAGLVGAD 507
Query: 542 GPTHQPIEHLASFRAMPNILMLRPADGNETAGAYKVAVLNRKRPSILALSRQKLPNLAGT 601
GPTH + +PN++++ P+D + AV RPS R N G
Sbjct: 508 GPTHCGAFDVTFMACLPNMIVMAPSDEADLFNMVATAVAIDDRPSCFRYPR---GNGIGV 564
Query: 602 SIEGVEKG-------GYIVSDNSTGNKPDVILIGTGSELEIAYKAGEDLRKEGKAVRV 652
++ KG G I+ + V L+G GS ++ A L + G V V
Sbjct: 565 ALPPGNKGVPIEIGKGRILKEGER-----VALLGYGSAVQSCLGAAVMLEERGLNVTV 617
>At2g16380 putative phosphatidylinositol/phosphatidylcholine
transfer protein
Length = 582
Score = 33.5 bits (75), Expect = 0.44
Identities = 25/82 (30%), Positives = 42/82 (50%), Gaps = 4/82 (4%)
Query: 576 KVAVL-NRKRPSIL-ALSRQKLPNLAGTSIEGVEKGGYIVSDNSTGNKPDVILIGTGSEL 633
K+ VL N+ +P +L A+ +LP G +KGG + SD N P+++ I E
Sbjct: 269 KIHVLGNKYQPKLLEAIDASELPYFFGGLCTCADKGGCLRSDKGPWNDPELLKIARNPEA 328
Query: 634 EIAYKAGED--LRKEGKAVRVV 653
+ + ED L +EG ++ +V
Sbjct: 329 RFSTISEEDYLLVEEGTSMSMV 350
>At1g24180 pyruvate dehydrogenase E1 alpha subunit
Length = 393
Score = 32.7 bits (73), Expect = 0.75
Identities = 37/162 (22%), Positives = 70/162 (42%), Gaps = 18/162 (11%)
Query: 186 GPLGQGIANAVGLALAEKHLAARFNKPDNEIIDHYTYCILGDGCQMEGIANEACSLAGHW 245
G +G I GLA A+K +NK + T+ + GDG +G EA +++ W
Sbjct: 169 GIVGAQIPLGCGLAFAQK-----YNKDEA-----VTFALYGDGAANQGQLFEALNISALW 218
Query: 246 GLGKLIAFYDDNHISIDGDTEIAFTESVEKRFEGLGWHVIWVKNGNTGFDEIRAAIKEAK 305
L ++ ++NH + T + + + G +V +K ++ A K AK
Sbjct: 219 DLPAILV-CENNHYGMGTAT---WRSAKSPAYFKRGDYVPGLKVDGMDALAVKQACKFAK 274
Query: 306 --AVKDRPTLIKFTT--TIGYGSPNKSNSYSVHGSALGAKEV 343
A+K+ P +++ T G+ + ++Y G ++V
Sbjct: 275 EHALKNGPIILEMDTYRYHGHSMSDPGSTYRTRDEISGVRQV 316
>At5g27870 pectin methyl-esterase - like protein
Length = 732
Score = 30.8 bits (68), Expect = 2.8
Identities = 17/54 (31%), Positives = 23/54 (42%)
Query: 12 TNPHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHASVSAP 65
TN +T + S +TT S PST+ T P HL + + S S P
Sbjct: 569 TNSTVTGSSLSSNTTESSDSPSTVVTPSTSPPAGHLGSPSDTPSSVVSPSTSLP 622
>At3g08670 hypothetical protein
Length = 567
Score = 30.8 bits (68), Expect = 2.8
Identities = 34/108 (31%), Positives = 48/108 (43%), Gaps = 12/108 (11%)
Query: 1 MSSSSSTSLYLTNPHLTRHTSSPSTTRLS---SLPSTITLNRTPTP---TTHLSQSHHLR 54
+S S S S Y H +R S S TR S S S+ T R+P+ T+ S S ++R
Sbjct: 137 LSVSQSESGY----HSSRPARSSSVTRPSISTSQYSSFTSGRSPSSILNTSSASVSSYIR 192
Query: 55 HTA--IHASVSAPPSTTTDSSLVEKSVNTIRFLAVDSVEKANSGHPGL 100
++ +S SA PST T +S +S R S + P L
Sbjct: 193 PSSPSSRSSSSARPSTPTRTSSASRSSTPSRIRPGSSSSSMDKARPSL 240
>At1g36150 hypothetical protein
Length = 256
Score = 30.8 bits (68), Expect = 2.8
Identities = 20/87 (22%), Positives = 43/87 (48%), Gaps = 3/87 (3%)
Query: 13 NPHLTRHTSSPSTTRLSSL-PSTITLNRTPTPTTHLSQSHHLRHTAIHASVSAPPSTTTD 71
+P + T+SPS+ + ++ PS+ ++ +P P H S + H++ S S+PP++ +
Sbjct: 133 SPPVESPTTSPSSAKSPAITPSSPAVSHSPPPVRH--SSPPVSHSSPPVSHSSPPTSRSS 190
Query: 72 SSLVEKSVNTIRFLAVDSVEKANSGHP 98
++ S V +V + + P
Sbjct: 191 PAVSHSSPVVAASSPVKAVSSSTASSP 217
>At3g09000 unknown protein
Length = 541
Score = 30.4 bits (67), Expect = 3.7
Identities = 26/83 (31%), Positives = 42/83 (50%), Gaps = 8/83 (9%)
Query: 3 SSSSTSLYLTNPHL-TRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHAS 61
SSS +S + P TR +++P+T+ + + + +R+ TPT+ + TA A+
Sbjct: 148 SSSGSSRSTSRPATPTRRSTTPTTSTSRPVTTRASNSRSSTPTSRATL------TAARAT 201
Query: 62 VS-APPSTTTDSSLVEKSVNTIR 83
S A P TTT SS +S R
Sbjct: 202 TSTAAPRTTTTSSGSARSATPTR 224
>At2g40180 protein phosphatase 2C (AthPP2C5)
Length = 390
Score = 30.4 bits (67), Expect = 3.7
Identities = 29/103 (28%), Positives = 42/103 (40%), Gaps = 6/103 (5%)
Query: 551 LASFRAMPNILML-----RPADGNETAGAYKVAVLNRKRPSILAL-SRQKLPNLAGTSIE 604
L+S R+ P L + R +T G+ VL RKRP +L L + + + T+ E
Sbjct: 53 LSSPRSSPPKLTMVACPPRKPKETKTTGSDSETVLKRKRPPMLDLTAAPTVASWCSTTRE 112
Query: 605 GVEKGGYIVSDNSTGNKPDVILIGTGSELEIAYKAGEDLRKEG 647
EKG +V G G +E Y A D +G
Sbjct: 113 TAEKGAEVVEAEEDGYYSVYCKRGRRGPMEDRYFAAVDRNDDG 155
>At5g52870 unknown protein (At5g52870)
Length = 326
Score = 30.0 bits (66), Expect = 4.8
Identities = 21/76 (27%), Positives = 34/76 (44%)
Query: 253 FYDDNHISIDGDTEIAFTESVEKRFEGLGWHVIWVKNGNTGFDEIRAAIKEAKAVKDRPT 312
F+ N+ +I EI +ES G + + G F E++ ++ + K K++
Sbjct: 9 FWLTNNTTIKPRREIRISESAVDSTTGSEDPDLDLCEGEDSFFELKISLSDFKTPKEKQR 68
Query: 313 LIKFTTTIGYGSPNKS 328
L TTT Y NKS
Sbjct: 69 LETTTTTTTYSVSNKS 84
>At5g04010 putative protein
Length = 274
Score = 30.0 bits (66), Expect = 4.8
Identities = 28/108 (25%), Positives = 43/108 (38%), Gaps = 15/108 (13%)
Query: 331 YSVHGSALGAKEVDATRNNLGWPY---EPFHVPEDVKKHWSRHIREGAALESEWNAKFAD 387
+ VH SA G+KE + + E F + DV+ S + + WN +
Sbjct: 135 FIVHVSA-GSKEAVVVKQGKDLVFGSNERFQIEADVRD--SGFTADMKDVSMSWNVVLRN 191
Query: 388 YEKKYKEEAAVLKSIISGDLPAGWEKALPTYTPEIPADATRNLSQQNL 435
YE+ + V+KS+ D GW +T E+P R NL
Sbjct: 192 YERMFLMSETVIKSL---DSMIGW------FTDELPGTKNRYCDGSNL 230
>At4g09710 RNA-directed DNA polymerase -like protein
Length = 1274
Score = 30.0 bits (66), Expect = 4.8
Identities = 21/71 (29%), Positives = 30/71 (41%), Gaps = 5/71 (7%)
Query: 287 VKNGNTGFDEIRAAI-----KEAKAVKDRPTLIKFTTTIGYGSPNKSNSYSVHGSALGAK 341
+ G GF EI A + KE ++ R L K+ T + + S S++ HG
Sbjct: 789 INEGGLGFREIEAKLSWRILKEPHSLLSRVLLGKYCNTSSFMDCSASPSFASHGWRGILA 848
Query: 342 EVDATRNNLGW 352
D R LGW
Sbjct: 849 GRDLLRKGLGW 859
>At4g05640 putative protein
Length = 207
Score = 30.0 bits (66), Expect = 4.8
Identities = 22/80 (27%), Positives = 37/80 (45%), Gaps = 16/80 (20%)
Query: 5 SSTSLYLTNPHLT-RHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHASV- 62
S S+ L +P + + T+ P TR +LP TP + ++ HH H+ IH++
Sbjct: 82 SGHSMTLLDPLVEYQFTTPPDNTRPFTLP------HHSTPRSSITSPHHHHHSTIHSTKL 135
Query: 63 --------SAPPSTTTDSSL 74
+PPS + D +L
Sbjct: 136 LDPLVEYHHSPPSASLDRTL 155
>At2g34190 putative membrane transporter
Length = 524
Score = 30.0 bits (66), Expect = 4.8
Identities = 24/81 (29%), Positives = 36/81 (43%), Gaps = 1/81 (1%)
Query: 380 EWNAKFADYEKKYKEEAAVLKSIISGDLPAGWEKALPTYTPEIPADATRNLSQQNLNALA 439
+W A D + AAVL S+I L + TP P +R + Q + L
Sbjct: 271 QWGAPSFDAGHAFAMMAAVLVSLIESTGAFKAAARLASATPPPPHVLSRGIGWQGIGILL 330
Query: 440 NVLPGLIGGSADLASSNMTLL 460
N L G + GS+ ++ N+ LL
Sbjct: 331 NGLFGTLSGSS-VSVENIGLL 350
>At2g18180 putative phosphatidylinositol/phophatidylcholine transfer
protein
Length = 558
Score = 30.0 bits (66), Expect = 4.8
Identities = 24/79 (30%), Positives = 36/79 (45%), Gaps = 7/79 (8%)
Query: 576 KVAVLNRKRPSILA--LSRQKLPNLAGTSIEGVEKGGYIVSDNSTGNKPDVI-LIGTG-- 630
K+ VL K S L + +LP G S + GG + SD N PD++ + G
Sbjct: 264 KIHVLGNKYQSKLLEIIDASELPEFLGGSCTCADNGGCMRSDKGPWNNPDIMKRVNNGDH 323
Query: 631 --SELEIAYKAGEDLRKEG 647
S+ A AGE++ +G
Sbjct: 324 ICSKRSQADNAGENIISQG 342
>At1g59900 pyruvate dehydrogenase E1 alpha subunit, putative
Length = 389
Score = 30.0 bits (66), Expect = 4.8
Identities = 34/161 (21%), Positives = 69/161 (42%), Gaps = 18/161 (11%)
Query: 186 GPLGQGIANAVGLALAEKHLAARFNKPDNEIIDHYTYCILGDGCQMEGIANEACSLAGHW 245
G +G + G+A A+K +NK + T+ + GDG +G EA +++ W
Sbjct: 165 GIVGAQVPLGCGIAFAQK-----YNKEEA-----VTFALYGDGAANQGQLFEALNISALW 214
Query: 246 GLGKLIAFYDDNHISIDGDTEIAFTESVEKRFEGLGWHVIWVKNGNTGFDEIRAAIKEAK 305
L ++ ++++ + A + S KR G +V +K ++ A K AK
Sbjct: 215 DLPAILVCENNHYGMGTAEWRAAKSPSYYKR----GDYVPGLKVDGMDAFAVKQACKFAK 270
Query: 306 --AVKDRPTLIKFTT--TIGYGSPNKSNSYSVHGSALGAKE 342
A++ P +++ T G+ + ++Y G ++
Sbjct: 271 QHALEKGPIILEMDTYRYHGHSMSDPGSTYRTRDEISGVRQ 311
>At1g19290 hypothetical protein, 5' partial
Length = 718
Score = 29.6 bits (65), Expect = 6.3
Identities = 35/113 (30%), Positives = 45/113 (38%), Gaps = 9/113 (7%)
Query: 146 LHLAGYDSVK-------VEDLKQFRQWESRTPGHPENFETPGIEVTTGPLGQGIANAVGL 198
L L GY S+K LK + ES P+ P V + G+ A L
Sbjct: 491 LLLPGYQSLKEFLEASATTCLKTQKIAESVENSTPKKLLVPNNIVYNVAIA-GLCKAGKL 549
Query: 199 ALAEKHLAARFNKPDNEIIDHYTYCILGDGCQMEGIANEACSLAGHWGLGKLI 251
A K L + D I D YTY IL GC + G N+A +L L +I
Sbjct: 550 EDARK-LFSDLLSSDRFIPDEYTYTILIHGCAIAGDINKAFTLRDEMALKGII 601
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.133 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,204,211
Number of Sequences: 26719
Number of extensions: 781465
Number of successful extensions: 2137
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 2121
Number of HSP's gapped (non-prelim): 27
length of query: 735
length of database: 11,318,596
effective HSP length: 107
effective length of query: 628
effective length of database: 8,459,663
effective search space: 5312668364
effective search space used: 5312668364
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 64 (29.3 bits)
Medicago: description of AC119419.5


