

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119417.4 - phase: 0 /pseudo
(459 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
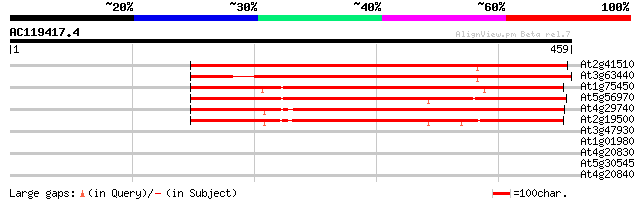
Sequences producing significant alignments: (bits) Value
At2g41510 putative cytokinin oxidase 442 e-124
At3g63440 cytokinin oxidase -like protein 390 e-109
At1g75450 cytokinin oxidase (CKX6) 355 2e-98
At5g56970 cytokinin oxidase 301 4e-82
At4g29740 cytokinin oxidase - like protein 273 1e-73
At2g19500 cytokinin oxidase (CKX2) 272 2e-73
At3g47930 L-galactono-1,4-lactone dehydrogenase - like protein 33 0.25
At1g01980 hypothetical protein 30 2.1
At4g20830 reticuline oxidase -like protein 30 2.8
At5g30545 putative protein 28 8.1
At4g20840 reticuline oxidase - like protein 28 8.1
>At2g41510 putative cytokinin oxidase
Length = 575
Score = 442 bits (1138), Expect = e-124
Identities = 209/312 (66%), Positives = 255/312 (80%), Gaps = 4/312 (1%)
Query: 149 SEEQNGELFYSVLGGLGQFGIITRARILLEPAPTMVKWIRVLYSDFTAFTKDQEKLIFAE 208
SE++N ELF+SVLGGLGQFGIITRARI LEPAP MVKWIRVLYSDF+AF++DQE LI E
Sbjct: 231 SEKRNSELFFSVLGGLGQFGIITRARISLEPAPHMVKWIRVLYSDFSAFSRDQEYLISKE 290
Query: 209 KAFDYIEGFVIKNRTGLLNNWRLSFNPQDPVQASKFKSDGRTLFCLELAKYFNMEETLEV 268
K FDY+EGFVI NRT LLNNWR SF+P D QAS+FKSDG+TL+CLE+ KYFN EE +
Sbjct: 291 KTFDYVEGFVIINRTDLLNNWRSSFSPNDSTQASRFKSDGKTLYCLEVVKYFNPEEASSM 350
Query: 269 NQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVKLRSKGLWDVPHPWLNLFIPKSK 328
+Q+ K LS LN+IPSTLF +EV Y++FLDRVHI+E KLR+KGLW+VPHPWLNL IPKS
Sbjct: 351 DQETGKLLSELNYIPSTLFSSEVPYIEFLDRVHIAERKLRAKGLWEVPHPWLNLLIPKSS 410
Query: 329 IHNFAEVVFGNIVKETSNGPVLIYPVHKSKWDKRTSVVIPDEDIFYLVGFLAS----SSG 384
I+ FA VF NI+ +NGP+LIYPV++SKW K TS++ P+EDIFYLV FL S SSG
Sbjct: 411 IYQFATEVFNNILTSNNNGPILIYPVNQSKWKKHTSLITPNEDIFYLVAFLPSAVPNSSG 470
Query: 385 PDELEHILSQNKRILEYCERAHLGVKQYLPHYTTQEEWQTHYGHKWEIFKQRKSIYDPLA 444
++LE++L QN+R++ +C A+L VKQYLPHY TQ+EW++H+G +WE F QRK YDPLA
Sbjct: 471 KNDLEYLLKQNQRVMNFCAAANLNVKQYLPHYETQKEWKSHFGKRWETFAQRKQAYDPLA 530
Query: 445 ILAPGQGIFSKS 456
ILAPGQ IF K+
Sbjct: 531 ILAPGQRIFQKT 542
>At3g63440 cytokinin oxidase -like protein
Length = 504
Score = 390 bits (1002), Expect = e-109
Identities = 191/316 (60%), Positives = 242/316 (76%), Gaps = 21/316 (6%)
Query: 149 SEEQNGELFYSVLGGLGQFGIITRARILLEPAPTMVKWIRVLYSDFTAFTKDQEKLIFAE 208
++ QN +LF VLGGLGQFGIITRARI LEPAPTM DQE+LI A+
Sbjct: 204 TKRQNSDLFNGVLGGLGQFGIITRARIALEPAPTM----------------DQEQLISAQ 247
Query: 209 -KAFDYIEGFVIKNRTGLLNNWRLSFNPQDPVQASKFKSDGRTLFCLELAKYFNMEETLE 267
FDYIEGFVI NRTGLLN+WRLSF ++P++AS+FK DGRTL+CLELAKY +
Sbjct: 248 GHKFDYIEGFVIINRTGLLNSWRLSFTAEEPLEASQFKFDGRTLYCLELAKYLKQDNKDV 307
Query: 268 VNQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVKLRSKGLWDVPHPWLNLFIPKS 327
+NQ++++ LS L+++ STLF TEV Y FLDRVH+SEVKLRSKG W+VPHPWLNL +P+S
Sbjct: 308 INQEVKETLSELSYVTSTLFTTEVAYEAFLDRVHVSEVKLRSKGQWEVPHPWLNLLVPRS 367
Query: 328 KIHNFAEVVFGNIVKETSNGPVLIYPVHKSKWDKRTSVVIPDEDIFYLVGFLAS----SS 383
KI+ FA VFGNI+ +TSNGPV++YPV+KSKWD +TS V P+E++FYLV L S S+
Sbjct: 368 KINEFARGVFGNILTDTSNGPVIVYPVNKSKWDNQTSAVTPEEEVFYLVAILTSASPGSA 427
Query: 384 GPDELEHILSQNKRILEYCERAHLGVKQYLPHYTTQEEWQTHYGHKWEIFKQRKSIYDPL 443
G D +E IL +N+RILE+ E A +G+KQYLPHYTT+EEW++H+G KW F +RKS YDPL
Sbjct: 428 GKDGVEEILRRNRRILEFSEEAGIGLKQYLPHYTTREEWRSHFGDKWGEFVRRKSRYDPL 487
Query: 444 AILAPGQGIFSKSITF 459
AILAPG IF K++++
Sbjct: 488 AILAPGHRIFQKAVSY 503
>At1g75450 cytokinin oxidase (CKX6)
Length = 540
Score = 355 bits (912), Expect = 2e-98
Identities = 174/312 (55%), Positives = 233/312 (73%), Gaps = 8/312 (2%)
Query: 149 SEEQNGELFYSVLGGLGQFGIITRARILLEPAPTMVKWIRVLYSDFTAFTKDQEKLI--F 206
SEE+N LF+ VLGGLGQFGIITRARI LEPAP V+WIRVLYS F FT+DQE LI
Sbjct: 210 SEEENTRLFHGVLGGLGQFGIITRARISLEPAPQRVRWIRVLYSSFKVFTEDQEYLISMH 269
Query: 207 AEKAFDYIEGFVIKNRTGLLNNWRLSF-NPQDPVQASKFKSDGRTLFCLELAKYFNMEET 265
+ FDY+EGFVI + GL+NNWR SF +P++PV+ S S+G L+CLE+ K ++ ++
Sbjct: 270 GQLKFDYVEGFVIVDE-GLVNNWRSSFFSPRNPVKISSVSSNGSVLYCLEITKNYHDSDS 328
Query: 266 LEVNQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVKLRSKGLWDVPHPWLNLFIP 325
V+Q+++ + LNFIP+++F T++ YVDFLDRVH +E+KLRSK LW+VPHPWLNLF+P
Sbjct: 329 EIVDQEVEILMKKLNFIPTSVFTTDLQYVDFLDRVHKAELKLRSKNLWEVPHPWLNLFVP 388
Query: 326 KSKIHNFAEVVFGNIVKETSNGPVLIYPVHKSKWDKRTSVVIPDEDIFYLVGFLASSSGP 385
KS+I +F + VF I+ ++GP+LIYP++K KWD+R+S V PDE++FYLV L S+
Sbjct: 389 KSRISDFDKGVFKGILGNKTSGPILIYPMNKDKWDERSSAVTPDEEVFYLVALLRSALTD 448
Query: 386 DE----LEHILSQNKRILEYCERAHLGVKQYLPHYTTQEEWQTHYGHKWEIFKQRKSIYD 441
E LE++ QN+RILE+CE+A + VKQYLPH+ TQEEW H+G KW+ F+ K+ +D
Sbjct: 449 GEETQKLEYLKDQNRRILEFCEQAKINVKQYLPHHATQEEWVAHFGDKWDRFRSLKAEFD 508
Query: 442 PLAILAPGQGIF 453
P ILA GQ IF
Sbjct: 509 PRHILATGQRIF 520
>At5g56970 cytokinin oxidase
Length = 523
Score = 301 bits (772), Expect = 4e-82
Identities = 144/310 (46%), Positives = 213/310 (68%), Gaps = 5/310 (1%)
Query: 149 SEEQNGELFYSVLGGLGQFGIITRARILLEPAPTMVKWIRVLYSDFTAFTKDQEKLIFAE 208
S++ N +LF++VLGGLGQFGIITRARI LE AP KW+R LY DF+ FT+DQE++I
Sbjct: 212 SKDMNSDLFFAVLGGLGQFGIITRARIKLEVAPKRAKWLRFLYIDFSEFTRDQERVISKT 271
Query: 209 KAFDYIEGFVIKNRTGLLNNWRLSFNP-QDPVQASKFKSDGRTLFCLELAKYFNMEETLE 267
D++EG ++ + G +NWR ++ P D ++ + R ++CLE+ KY++
Sbjct: 272 DGVDFLEGSIMVDH-GPPDNWRSTYYPPSDHLRIASMVKRHRVIYCLEVVKYYDETSQYT 330
Query: 268 VNQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVKLRSKGLWDVPHPWLNLFIPKS 327
VN+++++ LN + +++ +VTY+DFL+RV E+ L+SKG WDVPHPWLNLF+PK+
Sbjct: 331 VNEEMEELSDSLNHVRGFMYEKDVTYMDFLNRVRTGELNLKSKGQWDVPHPWLNLFVPKT 390
Query: 328 KIHNFAEVVFGNIV--KETSNGPVLIYPVHKSKWDKRTSVVIPDEDIFYLVGFLASSSGP 385
+I F + VF I+ ++GPVL+YP++++KW+ R S IP+ED+FY VGFL S+G
Sbjct: 391 QISKFDDGVFKGIILRNNITSGPVLVYPMNRNKWNDRMSAAIPEEDVFYAVGFL-RSAGF 449
Query: 386 DELEHILSQNKRILEYCERAHLGVKQYLPHYTTQEEWQTHYGHKWEIFKQRKSIYDPLAI 445
D E +N IL++CE A++GV QYLP++++QE W H+G +W IF +RK YDP I
Sbjct: 450 DNWEAFDQENMEILKFCEDANMGVIQYLPYHSSQEGWVRHFGPRWNIFVERKYKYDPKMI 509
Query: 446 LAPGQGIFSK 455
L+PGQ IF K
Sbjct: 510 LSPGQNIFQK 519
>At4g29740 cytokinin oxidase - like protein
Length = 524
Score = 273 bits (699), Expect = 1e-73
Identities = 137/311 (44%), Positives = 207/311 (66%), Gaps = 9/311 (2%)
Query: 149 SEEQNGELFYSVLGGLGQFGIITRARILLEPAPTMVKWIRVLYSDFTAFTKDQEKLIFA- 207
S + N ELFY VLGGLGQFGIITRARI L+ APT VKW R+LYSDF+AF +DQE+LI
Sbjct: 218 SPKLNPELFYGVLGGLGQFGIITRARIALDHAPTRVKWSRILYSDFSAFKRDQERLISMT 277
Query: 208 -EKAFDYIEGFVIKNRTGLLNNWRLSFNP-QDPVQASKFKSDGRTLFCLELAKYFNMEET 265
+ D++EG ++ + G ++ SF P D + + +D R ++ LE+AKY++
Sbjct: 278 NDLGVDFLEGQLMMSN-GFVDT---SFFPLSDQTRVASLVNDHRIIYVLEVAKYYDRTTL 333
Query: 266 LEVNQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVKLRSKGLWDVPHPWLNLFIP 325
++Q I L F P +F +V Y DFL+RV E KLRS GLW+VPHPWLN+F+P
Sbjct: 334 PIIDQVIDTLSRTLGFAPGFMFVQDVPYFDFLNRVRNEEDKLRSLGLWEVPHPWLNIFVP 393
Query: 326 KSKIHNFAE-VVFGNIVKETS-NGPVLIYPVHKSKWDKRTSVVIPDEDIFYLVGFLASSS 383
S+I +F + V+ G ++ +TS +G L YP +++KW+ R S + PDED+FY++G L S+
Sbjct: 394 GSRIQDFHDGVINGLLLNQTSTSGVTLFYPTNRNKWNNRMSTMTPDEDVFYVIGLLQSAG 453
Query: 384 GPDELEHILSQNKRILEYCERAHLGVKQYLPHYTTQEEWQTHYGHKWEIFKQRKSIYDPL 443
G + + + N +++++CE + + +K+YL HYT +E+W H+G KW+ F ++K ++DP
Sbjct: 454 GSQNWQELENLNDKVIQFCENSGIKIKEYLMHYTRKEDWVKHFGPKWDDFLRKKIMFDPK 513
Query: 444 AILAPGQGIFS 454
+L+PGQ IF+
Sbjct: 514 RLLSPGQDIFN 524
>At2g19500 cytokinin oxidase (CKX2)
Length = 501
Score = 272 bits (696), Expect = 2e-73
Identities = 136/311 (43%), Positives = 208/311 (66%), Gaps = 10/311 (3%)
Query: 149 SEEQNGELFYSVLGGLGQFGIITRARILLEPAPTMVKWIRVLYSDFTAFTKDQEKLIFA- 207
S + N ELFY VLGGLGQFGIITRARI+L+ AP KW R+LYSDFT FTKDQE+LI
Sbjct: 195 SRQLNPELFYGVLGGLGQFGIITRARIVLDHAPKRAKWFRMLYSDFTTFTKDQERLISMA 254
Query: 208 -EKAFDYIEGFVIKNRTGLLNNWRLSFNPQDPVQASKFKSDGRTLFCLELAKYFNMEETL 266
+ DY+EG + + G+++ F P D + + ++ LE+AKY++
Sbjct: 255 NDIGVDYLEGQIFLS-NGVVDT--SFFPPSDQSKVADLVKQHGIIYVLEVAKYYDDPNLP 311
Query: 267 EVNQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVKLRSKGLWDVPHPWLNLFIPK 326
+++ I L+++P + +V Y DFL+RVH+ E KLRS GLW++PHPWLNL++PK
Sbjct: 312 IISKVIDTLTKTLSYLPGFISMHDVAYFDFLNRVHVEENKLRSLGLWELPHPWLNLYVPK 371
Query: 327 SKIHNFAEVVFGNIV--KETSNGPVLIYPVHKSKWDKRTSVVIP--DEDIFYLVGFLASS 382
S+I +F V +I+ +++++G L+YP +++KWD R S +IP DED+ Y++G L S+
Sbjct: 372 SRILDFHNGVVKDILLKQKSASGLALLYPTNRNKWDNRMSAMIPEIDEDVIYIIGLLQSA 431
Query: 383 SGPDELEHILSQNKRILEYCERAHLGVKQYLPHYTTQEEWQTHYGHKWEIFKQRKSIYDP 442
+ P +L + S N++I+ +C+ + + +KQYL HYT++E+W H+G KW+ F +RK ++DP
Sbjct: 432 T-PKDLPEVESVNEKIIRFCKDSGIKIKQYLMHYTSKEDWIEHFGSKWDDFSKRKDLFDP 490
Query: 443 LAILAPGQGIF 453
+L+PGQ IF
Sbjct: 491 KKLLSPGQDIF 501
>At3g47930 L-galactono-1,4-lactone dehydrogenase - like protein
Length = 610
Score = 33.5 bits (75), Expect = 0.25
Identities = 67/316 (21%), Positives = 114/316 (35%), Gaps = 61/316 (19%)
Query: 149 SEEQNGELFYSVLGGLGQFGIITRARILLEPAPTMVKWIRVLYSDFTAFTKDQEKLIFAE 208
S E++ ELF+ GLG G++ + +V+ V S+ K+ +KL+ A
Sbjct: 258 SREKDPELFHLARCGLGGLGVVAEVTLQCVARHELVEHTYV--SNLQEIKKNHKKLLSAN 315
Query: 209 KAFDYI----EGFVIKNRTGLLNNWRLSFNPQDPVQASKFKSDG-----RTLFCLELAKY 259
K Y+ V+ ++ W S P+D K+ +D R L+ + KY
Sbjct: 316 KHVKYLYIPYTDTVVVVTCNPVSKW--SGPPKD---KPKYTTDEAVQHVRDLYRESIVKY 370
Query: 260 FNMEETLEVNQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVKLRSKGLWDVP--- 316
+ + + + L+F T + ++ +D L+ VH+++V W
Sbjct: 371 RVQDSGKKSPDSSEPDIQELSF---TELRDKLLALDPLNDVHVAKVNQAEAEFWKKSEGY 427
Query: 317 ---------------HPWLN--------LFIPKSKIHNFAEVVFGNIVKETSNGPVLIYP 353
W++ L P K + E + I KE P I
Sbjct: 428 RVGWSDEILGFDCGGQQWVSESCFPAGTLANPSMKDLEYIEELKKLIEKEAIPAPAPI-- 485
Query: 354 VHKSKWDKRTSVVI------PDEDIFYLVGFLASSSGPDELEHILSQNKRIL-EYCERAH 406
+ +W R+ I ++DIF VG + D Q K I E+ H
Sbjct: 486 --EQRWTARSKSPISPAFSTSEDDIFSWVGIIMYLPTADP-----RQRKDITDEFFHYRH 538
Query: 407 LGVKQYLPHYTTQEEW 422
L KQ ++ E W
Sbjct: 539 LTQKQLWDQFSAYEHW 554
>At1g01980 hypothetical protein
Length = 541
Score = 30.4 bits (67), Expect = 2.1
Identities = 17/46 (36%), Positives = 25/46 (53%), Gaps = 1/46 (2%)
Query: 145 RLLQSEEQNGELFYSVLGGLG-QFGIITRARILLEPAPTMVKWIRV 189
R+L + +LF+++ GG G FG+I +I L P P V RV
Sbjct: 219 RILDRKSMGEDLFWAIGGGGGASFGVILSFKIKLVPVPPRVTVFRV 264
>At4g20830 reticuline oxidase -like protein
Length = 570
Score = 30.0 bits (66), Expect = 2.8
Identities = 14/46 (30%), Positives = 27/46 (58%), Gaps = 1/46 (2%)
Query: 145 RLLQSEEQNGELFYSVLGGLG-QFGIITRARILLEPAPTMVKWIRV 189
R+L + +LF+++ GG G +G++ ++ L P P++V RV
Sbjct: 221 RVLDRKAMGEDLFWAITGGGGGSYGVVLGYKVKLVPVPSVVTVFRV 266
>At5g30545 putative protein
Length = 220
Score = 28.5 bits (62), Expect = 8.1
Identities = 12/44 (27%), Positives = 28/44 (63%), Gaps = 1/44 (2%)
Query: 263 EETLEVNQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVK 306
E+T V + K L+HL+ + L QT+ ++ F++ +H+++++
Sbjct: 163 EQTKAVAEQKDK-LAHLSLVEKYLRQTDPAFLHFIESIHLNQLQ 205
>At4g20840 reticuline oxidase - like protein
Length = 539
Score = 28.5 bits (62), Expect = 8.1
Identities = 14/46 (30%), Positives = 25/46 (53%), Gaps = 1/46 (2%)
Query: 145 RLLQSEEQNGELFYSVLGGLG-QFGIITRARILLEPAPTMVKWIRV 189
++L + +LF+++ GG G FG++ ++ L P P V RV
Sbjct: 220 QILDRKSMGEDLFWAISGGGGASFGVVLGYKVKLVPVPETVTVFRV 265
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.336 0.148 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,938,642
Number of Sequences: 26719
Number of extensions: 415329
Number of successful extensions: 1392
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 1369
Number of HSP's gapped (non-prelim): 11
length of query: 459
length of database: 11,318,596
effective HSP length: 103
effective length of query: 356
effective length of database: 8,566,539
effective search space: 3049687884
effective search space used: 3049687884
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC119417.4


