

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119414.8 + phase: 0 /pseudo
(427 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
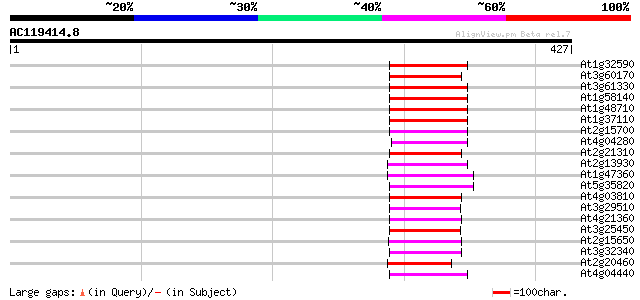
Score E
Sequences producing significant alignments: (bits) Value
At1g32590 hypothetical protein, 5' partial 76 3e-14
At3g60170 putative protein 70 3e-12
At3g61330 copia-type polyprotein 69 6e-12
At1g58140 hypothetical protein 69 6e-12
At1g48710 hypothetical protein 67 2e-11
At1g37110 51 1e-06
At2g15700 copia-like retroelement pol polyprotein 50 2e-06
At4g04280 putative transposon protein 50 3e-06
At2g21310 putative retroelement pol polyprotein 49 4e-06
At2g13930 putative retroelement pol polyprotein 49 4e-06
At1g47360 polyprotein, putative 48 9e-06
At5g35820 copia-like retrotransposable element 48 1e-05
At4g03810 putative retrotransposon protein 48 1e-05
At3g29510 hypothetical protein 48 1e-05
At4g21360 putative transposable element 47 2e-05
At3g25450 hypothetical protein 47 3e-05
At2g15650 putative retroelement pol polyprotein 46 4e-05
At3g32340 hypothetical protein 45 6e-05
At2g20460 putative retroelement pol polyprotein 44 1e-04
At4g04440 putative transposon protein 44 2e-04
>At1g32590 hypothetical protein, 5' partial
Length = 1263
Score = 76.3 bits (186), Expect = 3e-14
Identities = 36/59 (61%), Positives = 46/59 (77%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS YV+M+G GAI+W+S K+PIVTLSTTEAEFV+A+ +C +WLR + IG QE
Sbjct: 1123 KSTSGYVFMLGGGAIAWASKKQPIVTLSTTEAEFVSASYGACQAVWLRNVLEEIGCRQE 1181
>At3g60170 putative protein
Length = 1339
Score = 69.7 bits (169), Expect = 3e-12
Identities = 28/55 (50%), Positives = 44/55 (79%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIG 344
+STS +V+++ +GAI W+S K+P+V LSTTEAE++AAA C+C +WLR++ +G
Sbjct: 1197 RSTSGFVFLMASGAICWASKKQPVVALSTTEAEYIAAAFCACQCVWLRKVLEKLG 1251
>At3g61330 copia-type polyprotein
Length = 1352
Score = 68.6 bits (166), Expect = 6e-12
Identities = 32/59 (54%), Positives = 41/59 (69%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS +V+ +G+ A +W S K+PIVTLST EAE+VAA SC C IWLR + + QE
Sbjct: 1206 KSTSGFVFYIGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQE 1264
>At1g58140 hypothetical protein
Length = 1320
Score = 68.6 bits (166), Expect = 6e-12
Identities = 32/59 (54%), Positives = 41/59 (69%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS +V+ +G+ A +W S K+PIVTLST EAE+VAA SC C IWLR + + QE
Sbjct: 1174 KSTSGFVFYIGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQE 1232
>At1g48710 hypothetical protein
Length = 1352
Score = 66.6 bits (161), Expect = 2e-11
Identities = 31/59 (52%), Positives = 40/59 (67%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS +V+ +G+ A +W S K+PIV LST EAE+VAA SC C IWLR + + QE
Sbjct: 1206 KSTSGFVFYIGDTAFTWMSKKQPIVVLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQE 1264
>At1g37110
Length = 1356
Score = 51.2 bits (121), Expect = 1e-06
Identities = 20/59 (33%), Positives = 37/59 (61%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
+S S YV+ +G +SW + +P+V +STTEAE++A A + +W++ + +G Q+
Sbjct: 1218 RSISGYVFTIGGNTVSWKASLQPVVAMSTTEAEYIALAEAAKEAMWIKGLLQDMGMQQD 1276
>At2g15700 copia-like retroelement pol polyprotein
Length = 1166
Score = 50.4 bits (119), Expect = 2e-06
Identities = 24/59 (40%), Positives = 35/59 (58%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
+S + YV+ VG ISW S + +V LSTTEAE++A IWL+ + S +G Q+
Sbjct: 1068 RSVTGYVFKVGGNTISWKSSLQSVVALSTTEAEYMALTEAVKEAIWLKGLCSELGFKQD 1126
>At4g04280 putative transposon protein
Length = 1104
Score = 49.7 bits (117), Expect = 3e-06
Identities = 23/58 (39%), Positives = 33/58 (56%)
Query: 291 STSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
S S YV+ VG +SW S +P+V LSTTEAE++A +W+R + + G E
Sbjct: 1000 SISGYVFTVGGNIVSWKSCLQPVVALSTTEAEYIALTKAVKEAMWIRNLLDDMMLGTE 1057
>At2g21310 putative retroelement pol polyprotein
Length = 838
Score = 49.3 bits (116), Expect = 4e-06
Identities = 20/55 (36%), Positives = 34/55 (61%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIG 344
+ST+ Y + VG +SW S +P+V LS+TEAE++A + WL+ + + +G
Sbjct: 701 RSTTGYTFTVGGNIVSWRSCLQPVVALSSTEAEYIALTEAAKEAYWLKDLMNELG 755
>At2g13930 putative retroelement pol polyprotein
Length = 1335
Score = 49.3 bits (116), Expect = 4e-06
Identities = 21/61 (34%), Positives = 35/61 (56%)
Query: 288 TEKSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQ 347
T +S + YV+ VG ISW S + +V +S+TEAE++A +WL+ + +G Q
Sbjct: 1196 TRRSITGYVFTVGGNTISWKSKLQKVVAISSTEAEYMALTEAVKEALWLKGFAAELGHSQ 1255
Query: 348 E 348
+
Sbjct: 1256 D 1256
>At1g47360 polyprotein, putative
Length = 1182
Score = 48.1 bits (113), Expect = 9e-06
Identities = 23/67 (34%), Positives = 38/67 (56%), Gaps = 1/67 (1%)
Query: 288 TEKSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQ 347
T +S S YV+ +G I W S + +V LSTTE+E++A +WL+ + + +G Q
Sbjct: 1043 TRRSISGYVFTIGGNTIIWKSKLQKVVALSTTESEYMALTEAVKEALWLKGLANELGFPQ 1102
Query: 348 -EIEHSC 353
++E C
Sbjct: 1103 KDVEVHC 1109
>At5g35820 copia-like retrotransposable element
Length = 1342
Score = 47.8 bits (112), Expect = 1e-05
Identities = 25/65 (38%), Positives = 37/65 (56%), Gaps = 1/65 (1%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIG-EGQE 348
+S + +V+ G ISW S + +V LSTTEAE++A A IWLR + + +G E
Sbjct: 1205 RSITGFVFTAGGNTISWKSGLQRVVALSTTEAEYMALAEAVKEAIWLRGLAAEMGFEQDA 1264
Query: 349 IEHSC 353
+E C
Sbjct: 1265 VEVMC 1269
>At4g03810 putative retrotransposon protein
Length = 964
Score = 47.8 bits (112), Expect = 1e-05
Identities = 20/55 (36%), Positives = 35/55 (63%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIG 344
+S S + + + GA+SW S K+ V STTEAE++AA+ + +W+R+ + +G
Sbjct: 822 RSQSGFFFCLNGGAVSWKSTKQSTVADSTTEAEYIAASEAAKEVVWIRKFITELG 876
>At3g29510 hypothetical protein
Length = 1158
Score = 47.8 bits (112), Expect = 1e-05
Identities = 24/54 (44%), Positives = 32/54 (58%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHI 343
KS + YV++ G AISW SMK+ I S+ AE +A S +WLR +T HI
Sbjct: 1011 KSQTGYVFIHGGTAISWRSMKQTIAATSSNHAEILAMHEASRECVWLRSMTQHI 1064
>At4g21360 putative transposable element
Length = 1308
Score = 47.0 bits (110), Expect = 2e-05
Identities = 21/55 (38%), Positives = 32/55 (58%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIG 344
+S S YV+ VG +SW S + +V LS+T+AEF+A IW+R + +G
Sbjct: 1172 RSISGYVFTVGGNTVSWKSSLQHVVALSSTQAEFIALTEAVKEAIWIRGLLEDMG 1226
>At3g25450 hypothetical protein
Length = 1343
Score = 46.6 bits (109), Expect = 3e-05
Identities = 20/54 (37%), Positives = 35/54 (64%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHI 343
KST +++ + + I+W S K+ +VTLS+ EAEF+AA + IWL+ + + +
Sbjct: 1188 KSTGGHIFYLNDCPITWCSQKQQVVTLSSCEAEFMAATEAAKQAIWLQELLAEV 1241
>At2g15650 putative retroelement pol polyprotein
Length = 1347
Score = 45.8 bits (107), Expect = 4e-05
Identities = 21/56 (37%), Positives = 32/56 (56%)
Query: 289 EKSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIG 344
+KST+ YV+ +G W S K+ V ST EAE++A + + IWL+R+ G
Sbjct: 1206 KKSTTGYVFTLGLAMFCWQSCKQQTVAQSTAEAEYIAVCAATNQAIWLQRLFEDFG 1261
>At3g32340 hypothetical protein
Length = 871
Score = 45.4 bits (106), Expect = 6e-05
Identities = 20/55 (36%), Positives = 33/55 (59%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIG 344
+S +V+ GA+SW S K+ IV ST EAE++AA+ + W+R+ + +G
Sbjct: 729 QSDPGFVFCPNGGAVSWKSSKQSIVADSTIEAEYIAASEAAKEAAWIRKFITELG 783
>At2g20460 putative retroelement pol polyprotein
Length = 1461
Score = 44.3 bits (103), Expect = 1e-04
Identities = 21/49 (42%), Positives = 30/49 (60%)
Query: 288 TEKSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWL 336
+ +ST+ + VG+ ISW S K+P V+ S+ EAE+ A A SC WL
Sbjct: 1317 SRRSTTGFTMFVGSSLISWRSKKQPTVSRSSAEAEYRALALASCEMAWL 1365
>At4g04440 putative transposon protein
Length = 570
Score = 43.9 bits (102), Expect = 2e-04
Identities = 19/59 (32%), Positives = 33/59 (55%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
+ST+ YV+ VG +SW S + + + TTEAE+VA +W++ + +G Q+
Sbjct: 445 RSTTGYVFTVGGNTVSWKSNLQNTIAIPTTEAEYVAITEAVKEALWMQGLLKDMGVKQD 503
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.372 0.166 0.671
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,514,662
Number of Sequences: 26719
Number of extensions: 247462
Number of successful extensions: 1046
Number of sequences better than 10.0: 66
Number of HSP's better than 10.0 without gapping: 66
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 977
Number of HSP's gapped (non-prelim): 69
length of query: 427
length of database: 11,318,596
effective HSP length: 102
effective length of query: 325
effective length of database: 8,593,258
effective search space: 2792808850
effective search space used: 2792808850
T: 11
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC119414.8


