

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
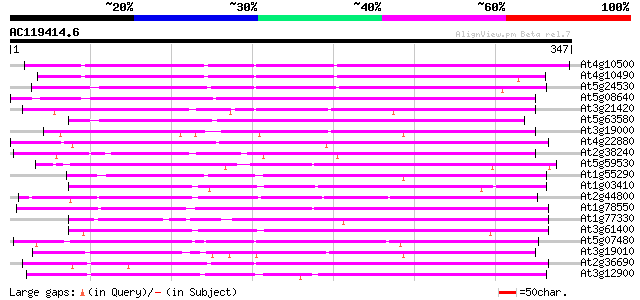
Query= AC119414.6 - phase: 0
(347 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At4g10500 putative Fe(II)/ascorbate oxidase 255 2e-68
At4g10490 putative flavanone 3-beta-hydroxylase 250 9e-67
At5g24530 flavanone 3-hydroxylase-like protein 230 1e-60
At5g08640 flavonol synthase (FLS) (sp|Q96330) 209 1e-54
At3g21420 unknown protein 201 6e-52
At5g63580 flavonol synthase 199 2e-51
At3g19000 unknown protein 192 3e-49
At4g22880 putative leucoanthocyanidin dioxygenase (LDOX) 191 4e-49
At2g38240 putative anthocyanidin synthase 187 7e-48
At5g59530 1-aminocyclopropane-1-carboxylate oxidase - like protein 185 3e-47
At1g55290 leucoanthocyanidin dioxygenase 2, putative 183 1e-46
At1g03410 putative 1-aminocyclopropane-1-carboxylate oxidase 183 1e-46
At2g44800 putative flavonol synthase 183 1e-46
At1g78550 unknown protein 183 1e-46
At1g77330 unknown protein 182 2e-46
At3g61400 1-aminocyclopropane-1-carboxylate oxidase-like protein 182 3e-46
At5g07480 putative protein 180 9e-46
At3g19010 oxidase like protein 179 2e-45
At2g36690 putative giberellin beta-hydroxylase 179 2e-45
At3g12900 hypothetical protein 178 3e-45
>At4g10500 putative Fe(II)/ascorbate oxidase
Length = 349
Score = 255 bits (652), Expect = 2e-68
Identities = 142/339 (41%), Positives = 199/339 (57%), Gaps = 8/339 (2%)
Query: 10 SSPLSLTPNFILPEHKRPHLSEVKYL-DSIPIIDLSYCDGNNPSSLEVIHKISKACEEFG 68
+S + + N++ P RP+LSEV+ DSIP+IDL D + P+ ++ +++ AC +G
Sbjct: 15 ASSVHIPSNYVRPISDRPNLSEVESSGDSIPLIDLR--DLHGPNRAVIVQQLASACSTYG 72
Query: 69 FFQIVNHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVDGGEKVK 128
FFQI NHGVPD KM FF ER S D TK RL + G +KV
Sbjct: 73 FFQIKNHGVPDTTVNKMQTVAREFFHQPESERVKHYSADPTKTTRLSTSFNV--GADKVL 130
Query: 129 LWSECFAHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDC 188
W + +PI+D I+ P +RE EYA V +LV RLL IS LGLE D
Sbjct: 131 NWRDFLRLHCFPIEDFIEEWPSS-PISFREVTAEYATSVRALVLRLLEAISESLGLESDH 189
Query: 189 LLKKLGEQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVNKDGKWIS 248
+ LG+ + A N+YPPCP+PELT GL H D +TVLLQ +VSGLQV KD KW++
Sbjct: 190 ISNILGKHAQHMA-FNYYPPCPEPELTYGLPGHKDPTVITVLLQDQVSGLQVFKDDKWVA 248
Query: 249 IPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPIHEL 308
+ IPN F++N+ DQ++V+SN +YKSVLHRAV N R+S+ FY P+ + +IGP HEL
Sbjct: 249 VSPIPNTFIVNIGDQMQVISNDKYKSVLHRAVVNTENERLSIPTFYFPSTDAVIGPAHEL 308
Query: 309 IDEEHP-PKYRNYHFSKFLEEFFNQEGTRRIVKEVFELP 346
++E+ YR Y F ++ ++F+N+ + F+ P
Sbjct: 309 VNEQDSLAIYRTYPFVEYWDKFWNRSLATASCLDAFKAP 347
>At4g10490 putative flavanone 3-beta-hydroxylase
Length = 348
Score = 250 bits (638), Expect = 9e-67
Identities = 141/317 (44%), Positives = 191/317 (59%), Gaps = 9/317 (2%)
Query: 18 NFILPEHKRPHLSEVKYL-DSIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIVNHG 76
N++ P RP +SEV+ DSIP+IDL D + P+ ++I++ + AC GFFQI NHG
Sbjct: 21 NYVRPVSDRPKMSEVQTSGDSIPLIDLH--DLHGPNRADIINQFAHACSSCGFFQIKNHG 78
Query: 77 VPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSECFAH 136
VP++ KMM A FF + ER S D K RL + EKV W +
Sbjct: 79 VPEETIKKMMNAAREFFRQSESERVKHYSADTKKTTRLSTSFNV--SKEKVSNWRDFLRL 136
Query: 137 PWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKLGEQ 196
YPI+D I P +RE EYA V +LV LL IS LGL +D + +G+
Sbjct: 137 HCYPIEDFINEWPST-PISFREVTAEYATSVRALVLTLLEAISESLGLAKDRVSNTIGKH 195
Query: 197 PRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVNKDGKWISIPCIPNAF 256
+ A N+YP CP PELT GL H D N +TVLLQ EVSGLQV KDGKWI++ +PN F
Sbjct: 196 GQHMA-INYYPRCPQPELTYGLPGHKDANLITVLLQDEVSGLQVFKDGKWIAVNPVPNTF 254
Query: 257 VINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPIHELIDEEH--P 314
++NL DQ++V+SN +YKSVLHRAV N+ RIS+ FY P+ + +I P ELI+EE P
Sbjct: 255 IVNLGDQMQVISNEKYKSVLHRAVVNSDMERISIPTFYCPSEDAVISPAQELINEEEDSP 314
Query: 315 PKYRNYHFSKFLEEFFN 331
YRN+ ++++ E+F++
Sbjct: 315 AIYRNFTYAEYFEKFWD 331
>At5g24530 flavanone 3-hydroxylase-like protein
Length = 341
Score = 230 bits (586), Expect = 1e-60
Identities = 131/323 (40%), Positives = 194/323 (59%), Gaps = 13/323 (4%)
Query: 14 SLTPNFILPEHKRPHLSEVKYLDSIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIV 73
+L N++ P RP LSEV L+ P+IDLS D + +I +I +AC FGFFQ++
Sbjct: 14 TLPENYVRPISDRPRLSEVSQLEDFPLIDLSSTDRSF-----LIQQIHQACARFGFFQVI 68
Query: 74 NHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSEC 133
NHGV Q+ +M+ FF ++ EE+ L S D TK RL + E+V W +
Sbjct: 69 NHGVNKQIIDEMVSVAREFFSMSMEEKMKLYSDDPTKTTRLSTSFNVKK--EEVNNWRDY 126
Query: 134 FAHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKL 193
YPI + P ++E ++Y++EV + ++ LIS LGLE+D + K L
Sbjct: 127 LRLHCYPIHKYVNEWPSN-PPSFKEIVSKYSREVREVGFKIEELISESLGLEKDYMKKVL 185
Query: 194 GEQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQ-SEVSGLQVNKDGKWISIPCI 252
GEQ + A N+YPPCP+PELT GL HTD NALT+LLQ + V GLQ+ DG+W ++
Sbjct: 186 GEQGQHMAV-NYYPPCPEPELTYGLPAHTDPNALTILLQDTTVCGLQILIDGQWFAVNPH 244
Query: 253 PNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIG---PIHELI 309
P+AFVIN+ DQ++ LSNG YKSV HRAVTN PR+S+A F P ++ P+ E
Sbjct: 245 PDAFVINIGDQLQALSNGVYKSVWHRAVTNTENPRLSVASFLCPADCAVMSPAKPLWEAE 304
Query: 310 DEEHPPKYRNYHFSKFLEEFFNQ 332
D+E P Y+++ ++++ ++F+++
Sbjct: 305 DDETKPVYKDFTYAEYYKKFWSR 327
>At5g08640 flavonol synthase (FLS) (sp|Q96330)
Length = 336
Score = 209 bits (533), Expect = 1e-54
Identities = 119/327 (36%), Positives = 184/327 (55%), Gaps = 12/327 (3%)
Query: 1 MESFQLANESSPLSLTPNFILPEHKRPHLSEVKY-LDSIPIIDLSYCDGNNPSSLEVIHK 59
+ S L E+ PL FI E ++P ++ + +IP++DLS +P V
Sbjct: 9 ISSSSLLTEAIPLE----FIRSEKEQPAITTFRGPTPAIPVVDLS-----DPDEESVRRA 59
Query: 60 ISKACEEFGFFQIVNHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYL 119
+ KA EE+G FQ+VNHG+P ++ ++ FFEL E+E ++ +++K++ + L
Sbjct: 60 VVKASEEWGLFQVVNHGIPTELIRRLQDVGRKFFELPSSEKESVAKPEDSKDIEGYGTKL 119
Query: 120 QVDGGEKVKLWSECFAHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLIS 179
Q D K K W + H +P + K +YRE EYA V L LL ++S
Sbjct: 120 QKDPEGK-KAWVDHLFHRIWPPSCVNYRFWPKNPPEYREVNEEYAVHVKKLSETLLGILS 178
Query: 180 IGLGLEEDCLLKKLG-EQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGL 238
GLGL+ D L + LG E + N+YPPCP P+L +G+ HTDL+ +T+L+ +EV GL
Sbjct: 179 DGLGLKRDALKEGLGGEMAEYMMKINYYPPCPRPDLALGVPAHTDLSGITLLVPNEVPGL 238
Query: 239 QVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNP 298
QV KD W IP+A ++++ DQI LSNGRYK+VLHR + + R+S +F P
Sbjct: 239 QVFKDDHWFDAEYIPSAVIVHIGDQILRLSNGRYKNVLHRTTVDKEKTRMSWPVFLEPPR 298
Query: 299 ETIIGPIHELIDEEHPPKYRNYHFSKF 325
E I+GP+ EL +++PPK++ + F +
Sbjct: 299 EKIVGPLPELTGDDNPPKFKPFAFKDY 325
>At3g21420 unknown protein
Length = 364
Score = 201 bits (510), Expect = 6e-52
Identities = 129/329 (39%), Positives = 186/329 (56%), Gaps = 20/329 (6%)
Query: 9 ESSPLSLTPNFILPEHKR----PHLSEVKYLDSIPIIDLSYCDG-NNPSSLEVIHKISKA 63
+S P + FI E++R L IP+IDLS +N I K+S+A
Sbjct: 22 KSKPNKVPERFIREEYERGVVVSSLKTHHLHHQIPVIDLSKLSKPDNDDFFFEILKLSQA 81
Query: 64 CEEFGFFQIVNHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVDG 123
CE++GFFQ++NHG+ +V + + + FF++ EE++ T Y
Sbjct: 82 CEDWGFFQVINHGIEVEVVEDIEEVASEFFDMPLEEKKKYPMEPGTVQ----GYGQAFIF 137
Query: 124 GEKVKL-WSECFA---HPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLIS 179
E KL W FA HP P +L P K ++ E+ Y+KE+ L +RLL I+
Sbjct: 138 SEDQKLDWCNMFALGVHP--PQIRNPKLWPSK-PARFSESLEGYSKEIRELCKRLLKYIA 194
Query: 180 IGLGLEEDCLLKKLGEQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVS--G 237
I LGL+E+ + GE Q + N+YPPC P+L +GL+ H+D +ALTVL QS+ S G
Sbjct: 195 ISLGLKEERFEEMFGEAV-QAVRMNYYPPCSSPDLVLGLSPHSDGSALTVLQQSKNSCVG 253
Query: 238 LQVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPN 297
LQ+ KD W+ + +PNA VIN+ D IEVLSNG+YKSV HRAVTN + R+++ FY PN
Sbjct: 254 LQILKDNTWVPVKPLPNALVINIGDTIEVLSNGKYKSVEHRAVTNREKERLTIVTFYAPN 313
Query: 298 PETIIGPIHELIDEE-HPPKYRNYHFSKF 325
E I P+ EL+D+E +P KYR+Y+ +
Sbjct: 314 YEVEIEPMSELVDDETNPCKYRSYNHGDY 342
>At5g63580 flavonol synthase
Length = 310
Score = 199 bits (505), Expect = 2e-51
Identities = 109/283 (38%), Positives = 163/283 (57%), Gaps = 9/283 (3%)
Query: 37 SIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIVNHGVPDQVCTKMMKAITNFFELA 96
+IPIIDLS D V H + K EE+G F +VNHG+P + ++ T FFEL
Sbjct: 18 TIPIIDLSNLDEEL-----VAHAVVKGSEEWGIFHVVNHGIPMDLIQRLKDVGTQFFELP 72
Query: 97 PEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSECFAHPWYPIDDIIQLLPEKIGTQY 156
E++ ++ D +K+ + L+ GE +W+E H +P I K QY
Sbjct: 73 ETEKKAVAKQDGSKDFEGYTTNLKYVKGE---VWTENLFHRIWPPTCINFDYWPKNPPQY 129
Query: 157 REAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKLG-EQPRQRAQSNFYPPCPDPELT 215
RE EY KE L R+L +S GLGL + L++ LG E + N YPP P P+LT
Sbjct: 130 REVIEEYTKETKKLSERILGYLSEGLGLPSEALIQGLGGESTEYVMRINNYPPDPKPDLT 189
Query: 216 MGLNEHTDLNALTVLLQSEVSGLQVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYKSV 275
+G+ EHTD+ +T+++ +EV GLQ+ KD W+ + IP++ +N+ DQI LSNG+YK+V
Sbjct: 190 LGVPEHTDIIGITIIITNEVPGLQIFKDDHWLDVHYIPSSITVNIGDQIMRLSNGKYKNV 249
Query: 276 LHRAVTNNVQPRISMAMFYGPNPETIIGPIHELIDEEHPPKYR 318
LHRA+ + R+S +F NP+ +IGP+ ELI ++P K++
Sbjct: 250 LHRAIVDKENQRMSWPVFVDANPDVVIGPLPELITGDNPSKFK 292
>At3g19000 unknown protein
Length = 352
Score = 192 bits (487), Expect = 3e-49
Identities = 120/327 (36%), Positives = 176/327 (53%), Gaps = 33/327 (10%)
Query: 22 PEHK-RPHLS---EVKYLDSIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIVNHGV 77
PEH+ HL+ + + D IP IDLS + + + +I++AC+ +GFFQ++NHG+
Sbjct: 12 PEHRPNTHLTNSGDFIFSDEIPTIDLSSLEDTHHDKTAIAKEIAEACKRWGFFQVINHGL 71
Query: 78 PDQVCTKMMKAITNFFELAPEEREHLS----------STDNTKNVR----LFNYYLQVDG 123
P + ++ K FF L EE+ + ++TKNVR +F+++LQ
Sbjct: 72 PSALRHRVEKTAAEFFNLTTEEKRKVKRDEVNPMGYHDEEHTKNVRDWKEIFDFFLQD-- 129
Query: 124 GEKVKLWSECFAHPWYPIDDIIQLLPEKIG---TQYREAFTEYAKEVGSLVRRLLSLISI 180
S P D ++ L + + +RE EYA+EV L RLL L+SI
Sbjct: 130 -------STIVPASPEPEDTELRKLTNQWPQNPSHFREVCQEYAREVEKLAFRLLELVSI 182
Query: 181 GLGLEEDCLLKKLGEQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQV 240
LGL D L EQ + N YPPCP+PEL +G+ H D ALTVL Q V GLQV
Sbjct: 183 SLGLPGDRLTGFFNEQT-SFLRFNHYPPCPNPELALGVGRHKDGGALTVLAQDSVGGLQV 241
Query: 241 NK--DGKWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNP 298
++ DG+WI + I +A +IN+ + I+V +N Y S HR V N + R S+ F+ P+
Sbjct: 242 SRRSDGQWIPVKPISDALIINMGNCIQVWTNDEYWSAEHRVVVNTSKERFSIPFFFFPSH 301
Query: 299 ETIIGPIHELIDEEHPPKYRNYHFSKF 325
E I P+ ELI EE+PP Y+ Y++ KF
Sbjct: 302 EANIEPLEELISEENPPCYKKYNWGKF 328
>At4g22880 putative leucoanthocyanidin dioxygenase (LDOX)
Length = 356
Score = 191 bits (486), Expect = 4e-49
Identities = 110/345 (31%), Positives = 198/345 (56%), Gaps = 14/345 (4%)
Query: 1 MESFQLANESSPLSLTPNFILPEHKRPHLSEVKYLDS-------IPIIDLSYCDGNNPSS 53
+E + +S +S+ +I P+ + +++V +L+ +P IDL + ++
Sbjct: 4 VERVESLAKSGIISIPKEYIRPKEELESINDV-FLEEKKEDGPQVPTIDLKNIESDDEKI 62
Query: 54 LE-VIHKISKACEEFGFFQIVNHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNV 112
E I ++ KA ++G ++NHG+P + ++ KA FF L+ EE+E ++ T +
Sbjct: 63 RENCIEELKKASLDWGVMHLINHGIPADLMERVKKAGEEFFSLSVEEKEKYANDQATGKI 122
Query: 113 RLFNYYLQVDGGEKVKLWSECFAHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVR 172
+ + L + +++ W + F H YP + + K + Y EA +EYAK + L
Sbjct: 123 QGYGSKLANNASGQLE-WEDYFFHLAYPEEKRDLSIWPKTPSDYIEATSEYAKCLRLLAT 181
Query: 173 RLLSLISIGLGLEEDCLLKKLG--EQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVL 230
++ +S+GLGLE D L K++G E+ + + N+YP CP PEL +G+ HTD++ALT +
Sbjct: 182 KVFKALSVGLGLEPDRLEKEVGGLEELLLQMKINYYPKCPQPELALGVEAHTDVSALTFI 241
Query: 231 LQSEVSGLQVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISM 290
L + V GLQ+ +GKW++ C+P++ V+++ D +E+LSNG+YKS+LHR + N + RIS
Sbjct: 242 LHNMVPGLQLFYEGKWVTAKCVPDSIVMHIGDTLEILSNGKYKSILHRGLVNKEKVRISW 301
Query: 291 AMFYGPNPETII-GPIHELIDEEHPPKYRNYHFSKFLE-EFFNQE 333
A+F P + I+ P+ E++ E P K+ F++ +E + F +E
Sbjct: 302 AVFCEPPKDKIVLKPLPEMVSVESPAKFPPRTFAQHIEHKLFGKE 346
>At2g38240 putative anthocyanidin synthase
Length = 353
Score = 187 bits (475), Expect = 7e-48
Identities = 113/332 (34%), Positives = 184/332 (55%), Gaps = 20/332 (6%)
Query: 3 SFQLANESSPLSLTPNFILPEHKRP--HLSEVKYLDSIPIIDLSYCDGNNPSSLEVIHKI 60
S Q +++ ++ ++ P H+RP + ++ IP++D++ G P L ++
Sbjct: 11 SVQSLSQTGVPTVPNRYVKPAHQRPVFNTTQSDAGIEIPVLDMNDVWGK-PEGLRLVRS- 68
Query: 61 SKACEEFGFFQIVNHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQ 120
ACEE+GFFQ+VNHGV + ++ A FFEL EE+ +++ +T Y +
Sbjct: 69 --ACEEWGFFQMVNHGVTHSLMERVRGAWREFFELPLEEKRKYANSPDTYE----GYGSR 122
Query: 121 VDGGEKVKL-WSECFAHPWYPIDDIIQLLPEKIGTQ---YREAFTEYAKEVGSLVRRLLS 176
+ + KL WS+ F + P P K +Q RE +Y +EV L RL
Sbjct: 123 LGVVKDAKLDWSDYFFLNYLPSSI---RNPSKWPSQPPKIRELIEKYGEEVRKLCERLTE 179
Query: 177 LISIGLGLEEDCLLKKLGEQPRQRA--QSNFYPPCPDPELTMGLNEHTDLNALTVLLQSE 234
+S LGL+ + L++ LG + A ++NFYP CP P+LT+GL+ H+D +T+LL E
Sbjct: 180 TLSESLGLKPNKLMQALGGGDKVGASLRTNFYPKCPQPQLTLGLSSHSDPGGITILLPDE 239
Query: 235 -VSGLQVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMF 293
V+GLQV + W++I +PNA ++N+ DQ+++LSNG YKSV H+ + N+ R+S+A F
Sbjct: 240 KVAGLQVRRGDGWVTIKSVPNALIVNIGDQLQILSNGIYKSVEHQVIVNSGMERVSLAFF 299
Query: 294 YGPNPETIIGPIHELIDEEHPPKYRNYHFSKF 325
Y P + +GPI EL+ P Y+ F ++
Sbjct: 300 YNPRSDIPVGPIEELVTANRPALYKPIRFDEY 331
>At5g59530 1-aminocyclopropane-1-carboxylate oxidase - like protein
Length = 364
Score = 185 bits (470), Expect = 3e-47
Identities = 110/331 (33%), Positives = 181/331 (54%), Gaps = 22/331 (6%)
Query: 17 PNFILPEHKRPHLSEVKYLDSIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIVNHG 76
P LP+ KR +S+++ IP ID + + + PS ++ K+ A E +GFFQ++NHG
Sbjct: 42 PQDTLPDKKRS-VSDLE----IPTIDFASVNVDTPSREAIVEKVKYAVENWGFFQVINHG 96
Query: 77 VPDQVCTKMMKAITNFFELA-PEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSE--- 132
VP V ++ + F E PE ++ S D TKN ++ + W +
Sbjct: 97 VPLNVLEEIKDGVRRFHEEEDPEVKKSYYSLDFTKNKFAYSSNFDLYSSSPSLTWRDSIS 156
Query: 133 CFAHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKK 192
C+ P P PE++ R+A EY+K V SL L L+S LGL+ + +LK
Sbjct: 157 CYMAPDPPT-------PEELPETCRDAMIEYSKHVLSLGDLLFELLSEALGLKSE-ILKS 208
Query: 193 LGEQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVNKDGKWISIPCI 252
+ ++YPPCP P+LT+G+++H+D + LTVLLQ + GLQ+ W+ + +
Sbjct: 209 MDCLKSLLMICHYYPPCPQPDLTLGISKHSDNSFLTVLLQDNIGGLQILHQDSWVDVSPL 268
Query: 253 PNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPN---PETIIGPIHELI 309
P A V+N+ D +++++N ++ SV HR + N PRIS+A F+ + T+ GP+ EL+
Sbjct: 269 PGALVVNVGDFLQLITNDKFISVEHRVLANTRGPRISVASFFSSSIRENSTVYGPMKELV 328
Query: 310 DEEHPPKYRNYHFSKFLEEFFNQ--EGTRRI 338
EE+PPKYR+ ++ E +F + +GT +
Sbjct: 329 SEENPPKYRDTTLREYSEGYFKKGLDGTSHL 359
>At1g55290 leucoanthocyanidin dioxygenase 2, putative
Length = 361
Score = 183 bits (465), Expect = 1e-46
Identities = 105/300 (35%), Positives = 164/300 (54%), Gaps = 14/300 (4%)
Query: 36 DSIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIVNHGVPDQVCTKMMKAITNFFEL 95
+SIP+ID+S D + S + A EE+GFFQ++NHGV +V M A FF L
Sbjct: 60 ESIPVIDISNLDEKSVSKA-----VCDAAEEWGFFQVINHGVSMEVLENMKTATHRFFGL 114
Query: 96 APEEREHLSSTDN-TKNVRLFNYYLQVDGGEKVKLWSECFAHPWYPIDDIIQLLPEKIGT 154
EE+ S + + NVR + EK W + + + + QL P+
Sbjct: 115 PVEEKRKFSREKSLSTNVRFGTSFSP--HAEKALEWKDYLSLFFVSEAEASQLWPDSC-- 170
Query: 155 QYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKLGEQPRQRAQSNFYPPCPDPEL 214
R EY E LV++LL + L ++E K+ R N+YP CP+PEL
Sbjct: 171 --RSETLEYMNETKPLVKKLLRFLGENLNVKELDKTKESFFMGSTRINLNYYPICPNPEL 228
Query: 215 TMGLNEHTDLNALTVLLQSEVSGLQVNK--DGKWISIPCIPNAFVINLADQIEVLSNGRY 272
T+G+ H+D+++LT+LLQ E+ GL V G+W+ +P I + VIN+ D ++++SNGRY
Sbjct: 229 TVGVGRHSDVSSLTILLQDEIGGLHVRSLTTGRWVHVPPISGSLVINIGDAMQIMSNGRY 288
Query: 273 KSVLHRAVTNNVQPRISMAMFYGPNPETIIGPIHELIDEEHPPKYRNYHFSKFLEEFFNQ 332
KSV HR + N RIS+ +F P PE++IGP+ E+I+ P Y++ ++ +++ FF +
Sbjct: 289 KSVEHRVLANGSYNRISVPIFVSPKPESVIGPLLEVIENGEKPVYKDILYTDYVKHFFRK 348
>At1g03410 putative 1-aminocyclopropane-1-carboxylate oxidase
Length = 398
Score = 183 bits (465), Expect = 1e-46
Identities = 111/303 (36%), Positives = 167/303 (54%), Gaps = 16/303 (5%)
Query: 37 SIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIVNHGVPDQVCTKMMKAITNFFELA 96
+IP +DL + S V+ KI A E +GFFQ+VNHG+ +V +M + I F E
Sbjct: 93 TIPTVDLKGGSMDLISRRSVVEKIGDAAERWGFFQVVNHGISVEVMERMKEGIRRFHEQD 152
Query: 97 PEEREHLSSTDNTKNVRLFNYYLQVD--GGEKVKLWSECFAHPWYPIDDIIQLLPEKIGT 154
PE ++ S D+T++V YY +D K W + A P +Q LP G
Sbjct: 153 PEVKKRFYSRDHTRDVL---YYSNIDLHTCNKAANWRDTLACYMAPDPPKLQDLPAVCG- 208
Query: 155 QYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKLGEQPRQRAQSNFYPPCPDPEL 214
E EY+K++ +L L L+S LGL + L K +G +YPPCP P+L
Sbjct: 209 ---EIMMEYSKQLMTLGEFLFELLSEALGLNPNHL-KDMGCAKSHIMFGQYYPPCPQPDL 264
Query: 215 TMGLNEHTDLNALTVLLQSEVSGLQVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYKS 274
T+G+++HTD + +T+LLQ + GLQV D W+ + +P A VIN+ D ++++SN ++ S
Sbjct: 265 TLGISKHTDFSFITILLQDNIGGLQVIHDQCWVDVSPVPGALVINIGDLLQLISNDKFIS 324
Query: 275 VLHRAVTN-NVQPRISM----AMFYGPNPETIIGPIHELIDEEHPPKYRNYHFSKFLEEF 329
HR + N + +PRISM + F PNP I GPI EL+ E++P KYR+ ++F F
Sbjct: 325 AEHRVIANGSSEPRISMPCFVSTFMKPNPR-IYGPIKELLSEQNPAKYRDLTITEFSNTF 383
Query: 330 FNQ 332
+Q
Sbjct: 384 RSQ 386
>At2g44800 putative flavonol synthase
Length = 352
Score = 183 bits (464), Expect = 1e-46
Identities = 114/325 (35%), Positives = 176/325 (54%), Gaps = 12/325 (3%)
Query: 6 LANESSPLSLTPNFILPEHKRPHLSEVKYLD--SIPIIDLSYCDGNNPSSLEVIHKISKA 63
L N P + ++LP +RP L ++P+IDLS SL IH+IS A
Sbjct: 14 LTNSGVP-QVPDRYVLPPSQRPALGSSLGTSETTLPVIDLSLLHQPFLRSL-AIHEISMA 71
Query: 64 CEEFGFFQIVNHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVDG 123
C+EFGFFQ++NHG+P V + A T FF+L EE+ L S + + VR Y ++
Sbjct: 72 CKEFGFFQVINHGIPSSVVNDALDAATQFFDLPVEEKMLLVSANVHEPVR---YGTSLNH 128
Query: 124 G-EKVKLWSECFAHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGL 182
++V W + H +P+ I + P Y++ +YA+ L ++L+ IS L
Sbjct: 129 STDRVHYWRDFIKHYSHPLSKWIDMWPSNPPC-YKDKVGKYAEATHLLHKQLIEAISESL 187
Query: 183 GLEEDCLLKKLGEQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVNK 242
GLE++ L +++ E+ Q N YP CP+PE+ +G+ H+D ++LT+LLQS GLQ+
Sbjct: 188 GLEKNYLQEEI-EEGSQVMAVNCYPACPEPEMALGMPPHSDFSSLTILLQSS-KGLQIMD 245
Query: 243 DGK-WISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETI 301
K W+ +P I A ++ L DQ+EV+SNG YKSV+HR N R+S A +
Sbjct: 246 CNKNWVCVPYIEGALIVQLGDQVEVMSNGIYKSVIHRVTVNKEVKRLSFASLHSLPLHKK 305
Query: 302 IGPIHELIDEEHPPKYRNYHFSKFL 326
I P +L++ + P Y + F+ FL
Sbjct: 306 ISPAPKLVNPNNAPAYGEFSFNDFL 330
>At1g78550 unknown protein
Length = 356
Score = 183 bits (464), Expect = 1e-46
Identities = 104/333 (31%), Positives = 184/333 (55%), Gaps = 11/333 (3%)
Query: 5 QLANESSPLSLTPNFI-LPEHKRPHLSEVKYLDSIPIIDLSYCDGNNPSSLEVIHKISKA 63
++ E + ++ P ++ + + K L++ IP+ID++ + E + K+ A
Sbjct: 19 EIVKEKNFTTIPPRYVRVDQEKTEILNDSSLSSEIPVIDMTRLCSVSAMDSE-LKKLDFA 77
Query: 64 CEEFGFFQIVNHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVD- 122
C+++GFFQ+VNHG+ K+ + FF L +E++ L F + QV+
Sbjct: 78 CQDWGFFQLVNHGIDSSFLEKLETEVQEFFNLPMKEKQKLWQRSGE-----FEGFGQVNI 132
Query: 123 GGEKVKL-WSECFAHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIG 181
E KL W + F PI L K+ +RE Y+ EV S+ + L + ++
Sbjct: 133 VSENQKLDWGDMFILTTEPIRSRKSHLFSKLPPPFRETLETYSSEVKSIAKILFAKMASV 192
Query: 182 LGLEEDCLLKKLGEQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQ-SEVSGLQV 240
L ++ + ++ L + Q + N+YPPCP P+ MGL +H+D LT+LLQ ++V GLQ+
Sbjct: 193 LEIKHE-EMEDLFDDVWQSIKINYYPPCPQPDQVMGLTQHSDAAGLTILLQVNQVEGLQI 251
Query: 241 NKDGKWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPET 300
KDGKW+ + + +A V+N+ + +E+++NGRY+S+ HRAV N+ + R+S+AMF+ P ET
Sbjct: 252 KKDGKWVVVKPLRDALVVNVGEILEIITNGRYRSIEHRAVVNSEKERLSVAMFHSPGKET 311
Query: 301 IIGPIHELIDEEHPPKYRNYHFSKFLEEFFNQE 333
II P L+D + +++ ++ + FF Q+
Sbjct: 312 IIRPAKSLVDRQKQCLFKSMSTQEYFDAFFTQK 344
>At1g77330 unknown protein
Length = 307
Score = 182 bits (463), Expect = 2e-46
Identities = 108/305 (35%), Positives = 163/305 (53%), Gaps = 21/305 (6%)
Query: 37 SIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIVNHGVPDQVCTKMMKAITNFFELA 96
+IP+ID S +G + + +I++ACEE+GFFQ+VNHG+P ++ K+ K ++ ++
Sbjct: 2 AIPVIDFSKLNGEERE--KTLSEIARACEEWGFFQLVNHGIPLELLNKVKKLSSDCYKT- 58
Query: 97 PEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSECFAHPWYPIDDIIQLLPEKIGTQY 156
ERE T N V+L N +Q + GEK++ W + ++ + +
Sbjct: 59 --EREEAFKTSNP--VKLLNELVQKNSGEKLENVD------WEDVFTLLDHNQNEWPSNI 108
Query: 157 REAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKLGEQPRQRAQSNF-------YPPC 209
+E EY +EV L +++ ++ LGL + + K E ++ F YPPC
Sbjct: 109 KETMGEYREEVRKLASKMMEVMDENLGLPKGYIKKAFNEGMEDGEETAFFGTKVSHYPPC 168
Query: 210 PDPELTMGLNEHTDLNALTVLLQS-EVSGLQVNKDGKWISIPCIPNAFVINLADQIEVLS 268
P PEL GL HTD + +L Q E GLQV KDG+WI + +PNA VIN DQIEVLS
Sbjct: 169 PHPELVNGLRAHTDAGGVVLLFQDDEYDGLQVLKDGEWIDVQPLPNAIVINTGDQIEVLS 228
Query: 269 NGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPIHELIDEEHPPKYRNYHFSKFLEE 328
NGRYKS HR + R S+A FY P+ + IGP +E KY + F +++
Sbjct: 229 NGRYKSAWHRVLAREEGNRRSIASFYNPSYKAAIGPAAVAEEEGSEKKYPKFVFGDYMDV 288
Query: 329 FFNQE 333
+ NQ+
Sbjct: 289 YANQK 293
>At3g61400 1-aminocyclopropane-1-carboxylate oxidase-like protein
Length = 370
Score = 182 bits (461), Expect = 3e-46
Identities = 110/308 (35%), Positives = 169/308 (54%), Gaps = 18/308 (5%)
Query: 37 SIPIIDLS-----YCDGNNPSSLEVIHKISKACEEFGFFQIVNHGVPDQVCTKMMKAITN 91
+IP IDL+ Y + ++ + ++ KI A E++GFFQ+VNHG+P V K+ + I
Sbjct: 59 TIPTIDLNGGVVYYKNQDSVTRRSMVEKIGDAAEKWGFFQVVNHGIPLDVLEKVKEGIRA 118
Query: 92 FFELAPEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKL-WSECFAHPWYPIDDIIQLLPE 150
F E E ++ S D+T+ + YY +D +K W + P + LPE
Sbjct: 119 FHEQDAELKKRFYSRDHTRKMV---YYSNLDLFTAMKASWRDTMCAYMAPDPPTSEDLPE 175
Query: 151 KIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKLGEQPRQRAQSNFYPPCP 210
G E EYAKE+ +L + L+S LGL LK + +YPPCP
Sbjct: 176 VCG----EIMMEYAKEIMNLGELIFELLSEALGLNNSNHLKDMDCSKSLVLFGQYYPPCP 231
Query: 211 DPELTMGLNEHTDLNALTVLLQSEVSGLQVNKDGK-WISIPCIPNAFVINLADQIEVLSN 269
P+ T+GL++HTD + LT++LQ + GLQV D + WI IP +P A V+NL D ++++SN
Sbjct: 232 QPDHTLGLSKHTDFSFLTIVLQGNLGGLQVLHDKQYWIDIPPVPGALVVNLGDLLQLISN 291
Query: 270 GRYKSVLHRAVTNN-VQPRISMAMFYGP---NPETIIGPIHELIDEEHPPKYRNYHFSKF 325
G++ SV HR + N +PRIS+ F+ + GPI EL+ E++PPKYR+ S+F
Sbjct: 292 GKFISVEHRVIANRAAEPRISVPCFFSTVMRESHRVYGPIKELLSEQNPPKYRDTTISEF 351
Query: 326 LEEFFNQE 333
+ ++E
Sbjct: 352 ASMYASKE 359
>At5g07480 putative protein
Length = 356
Score = 180 bits (457), Expect = 9e-46
Identities = 107/334 (32%), Positives = 178/334 (53%), Gaps = 16/334 (4%)
Query: 3 SFQLANESSPLSL--TPN-FILPEHKRPHLSEVKYLDSIPIIDLSYCDGNNPSSLEVIHK 59
+F++ N + SL P+ +++P +P S +P ID+S G + VI +
Sbjct: 9 TFKMGNSAQERSLPYVPDCYVVPPSSKPCDSNSGI---VPTIDVSRLKGGDDERRGVIQE 65
Query: 60 ISKACEEFGFFQIVNHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYL 119
+S AC+ GFFQIVNHG+ + ++ +FFEL +E++ S D VR ++ L
Sbjct: 66 LSLACQHLGFFQIVNHGINQNILDDALEVANSFFELPAKEKKQFMSNDVYAPVR-YSTSL 124
Query: 120 QVDGGEKVKLWSECFAHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLIS 179
+ DG + ++ W H +P+ I L PE YRE ++ +EV L L+ I+
Sbjct: 125 K-DGLDTIQFWRIFLKHYAHPLHRWIHLWPEN-PPGYREKMGKFCEEVRKLSIELMGAIT 182
Query: 180 IGLGLEEDCLLKKLGEQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQ 239
LGL D L ++ E Q N YPPCPDPE +GL H+D + +T+LLQ+ + GL+
Sbjct: 183 ESLGLGRDYLSSRMDENGMQVMTVNCYPPCPDPETALGLPPHSDYSCITLLLQN-LDGLK 241
Query: 240 V------NKDGKWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMF 293
+ G+W+ +P + +++ D +EVLSNG YKS++H+ N + RIS+A
Sbjct: 242 IFDPMAHGGSGRWVGVPQVTGVLKVHIGDHVEVLSNGLYKSIVHKVTLNEEKTRISLASL 301
Query: 294 YGPNPETIIGPIHELIDEEHPPKYRNYHFSKFLE 327
+ + + EL+++E+P +Y+ F+ FL+
Sbjct: 302 HSLGMDDKMSVPRELVNDENPVRYKESSFNDFLD 335
>At3g19010 oxidase like protein
Length = 349
Score = 179 bits (454), Expect = 2e-45
Identities = 116/330 (35%), Positives = 175/330 (52%), Gaps = 29/330 (8%)
Query: 15 LTPNFI-LPEHKRPHLSEVKYLDSIPIIDLSYCDGNNPSSLE-VIHKISKACEEFGFFQI 72
L P +I PEH+ + IP+IDLS D +P ++ VI +I ACE++GFFQ+
Sbjct: 4 LDPTYIQAPEHRSDSNFIPNQPEEIPVIDLSRLD--DPEDVQNVISEIGDACEKWGFFQV 61
Query: 73 VNHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVDGG--EKVKLW 130
+NHGVP ++ K + FF+L EE+ + D V Y+ DG + VK W
Sbjct: 62 LNHGVPSDARQRVEKTVKMFFDLPMEEKIKVKRDD----VNPVGYH---DGEHTKNVKDW 114
Query: 131 SECF----------AHPWYPIDDIIQLLPEK---IGTQYREAFTEYAKEVGSLVRRLLSL 177
E F P D+ ++L+ K + +REA YA+ L +LL L
Sbjct: 115 KEVFDIYFKDPMVIPSTTDPEDEGLRLVYNKWPQSPSDFREACEVYARHAEKLAFKLLEL 174
Query: 178 ISIGLGLEEDCLLKKLGEQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSG 237
IS+ LGL ++ EQ + N YPPCP P+L +G+ H D + +++L Q +V G
Sbjct: 175 ISLSLGLPKERFHDYFKEQ-MSFFRINRYPPCPRPDLALGVGHHKDADVISLLAQDDVGG 233
Query: 238 LQVNK--DGKWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYG 295
LQV++ DG W I +PNA VIN+ + +E+ +N +Y S HR V N + R S+ F
Sbjct: 234 LQVSRRSDGVWFPIRPVPNALVINIGNCMEIWTNDKYWSAEHRVVVNTTRERYSIPFFLL 293
Query: 296 PNPETIIGPIHELIDEEHPPKYRNYHFSKF 325
P+ + + P+ EL+ E+PPKY+ Y + KF
Sbjct: 294 PSHDVEVKPLEELVSPENPPKYKGYKWGKF 323
>At2g36690 putative giberellin beta-hydroxylase
Length = 392
Score = 179 bits (454), Expect = 2e-45
Identities = 111/357 (31%), Positives = 179/357 (50%), Gaps = 37/357 (10%)
Query: 9 ESSPLSLTPNFILPEHKRPHLSEVKYLDS------IPIIDLSYCDGNNPSSLEVIHKISK 62
E+ + +I PE RP L++ L +P+ID + G P+ V+ I++
Sbjct: 26 ENGLTKVPTKYIWPEPDRPILTKSDKLIKPNKNLKLPLIDFAELLG--PNRPHVLRTIAE 83
Query: 63 ACEEFGFFQI--------------------------VNHGVPDQVCTKMMKAITNFFELA 96
AC+ +GFFQ+ VNHG+ V M+ FFEL
Sbjct: 84 ACKTYGFFQVGSLNPKAHAVISRFLAKQVSYGNGQVVNHGMEGDVSKNMIDVCKRFFELP 143
Query: 97 PEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSECFAHPWYPIDDIIQLLPEKIGTQY 156
EER S+D + VR + Q+ + V W + +P+ D + P + +
Sbjct: 144 YEERSKYMSSDMSAPVRYGTSFNQIK--DNVFCWRDFLKLYAHPLPDYLPHWPSS-PSDF 200
Query: 157 REAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKLGEQPRQRAQSNFYPPCPDPELTM 216
R + YAKE + ++ I L ++ K E+ Q N YPPCP+PELT+
Sbjct: 201 RSSAATYAKETKEMFEMMVKAILESLEIDGSDEAAKELEEGSQVVVVNCYPPCPEPELTL 260
Query: 217 GLNEHTDLNALTVLLQSEVSGLQVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYKSVL 276
G+ H+D LT+LLQ EV GLQ+ +W+++ IP +FV+N+ D +E+ SNGRYKSVL
Sbjct: 261 GMPPHSDYGFLTLLLQDEVEGLQILYRDEWVTVDPIPGSFVVNVGDHLEIFSNGRYKSVL 320
Query: 277 HRAVTNNVQPRISMAMFYGPNPETIIGPIHELIDEEHPPKYRNYHFSKFLEEFFNQE 333
HR + N+ +PRIS+A + +++ P +L+D+ +P +Y + F+ FL+ ++E
Sbjct: 321 HRVLVNSTKPRISVASLHSFPLTSVVKPSPKLVDKHNPSQYMDTDFTTFLQYITSRE 377
>At3g12900 hypothetical protein
Length = 357
Score = 178 bits (452), Expect = 3e-45
Identities = 109/327 (33%), Positives = 172/327 (52%), Gaps = 16/327 (4%)
Query: 11 SPLSLTPN-FILPEHKRPHLSEVKYLDSIPIIDLSYCDGNNPSSLEVIHKISKACEEFGF 69
S LS P F+ P +R + ++ IDLS DG P EV +I +A E GF
Sbjct: 28 SGLSSVPRPFVQPLSERIPTQKALTCEATQPIDLSNLDG--PQHKEVAKQIVEAAETLGF 85
Query: 70 FQIVNHGVPDQVCTKMMKAITNFFELAPEERE-HLSSTDNTKNVRLFNYYLQVDGGEKVK 128
FQ+VNHGV ++ + + FF APEE+ +L +K V+ + V EK
Sbjct: 86 FQVVNHGVSVELLELLKSSAHEFFAQAPEEKSMYLKEVSPSKLVKYGTSF--VPDKEKAI 143
Query: 129 LWSECFAHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLI--SIGLGLEE 186
W + + + + +Q P+ RE E+ +V+ +++++ ++G+ LEE
Sbjct: 144 EWKDYVSMLYTNDSEALQHWPQPC----REVALEFLNSSMEMVKNVVNILMENVGVTLEE 199
Query: 187 DCLLKKLGEQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVNKD-GK 245
+ K G + N+YP CP PELT+G+ H+D+ LTVLLQ + GL V D G+
Sbjct: 200 E---KMNGLMGTKMVNMNYYPTCPSPELTVGVGRHSDMGMLTVLLQDGIGGLYVKLDNGE 256
Query: 246 WISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPI 305
W IP + A VIN+ D +++LSNG+YKS HR T N+ R+S+ +F PNP +GP+
Sbjct: 257 WAEIPPVHGALVINIGDTLQILSNGKYKSAEHRVRTTNIGSRVSVPIFTAPNPSQKVGPL 316
Query: 306 HELIDEEHPPKYRNYHFSKFLEEFFNQ 332
E++ + +Y+ + F ++ FF Q
Sbjct: 317 PEVVKRDGVARYKEFLFQDYMNNFFGQ 343
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.138 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,502,505
Number of Sequences: 26719
Number of extensions: 376662
Number of successful extensions: 1259
Number of sequences better than 10.0: 109
Number of HSP's better than 10.0 without gapping: 102
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 913
Number of HSP's gapped (non-prelim): 127
length of query: 347
length of database: 11,318,596
effective HSP length: 100
effective length of query: 247
effective length of database: 8,646,696
effective search space: 2135733912
effective search space used: 2135733912
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC119414.6


