

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119412.2 - phase: 0
(184 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
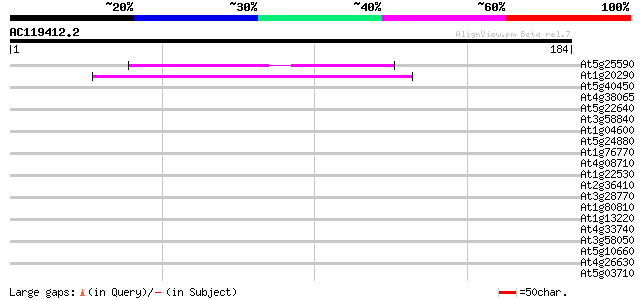
Score E
Sequences producing significant alignments: (bits) Value
At5g25590 unknown protein (At5g25590) 41 4e-04
At1g20290 hypothetical protein 40 7e-04
At5g40450 unknown protein 39 0.002
At4g38065 hypothetical protein 38 0.003
At5g22640 unknown protein 37 0.005
At3g58840 unknown protein 37 0.005
At1g04600 putative myosin heavy chain 37 0.005
At5g24880 glutamic acid-rich protein 37 0.008
At1g76770 putative heat shock protein 37 0.008
At4g08710 putative protein 36 0.010
At1g22530 unknown protein 36 0.010
At2g36410 unknown protein 36 0.013
At3g28770 hypothetical protein 35 0.017
At1g80810 unknown protein 35 0.017
At1g13220 putative nuclear matrix constituent protein 35 0.017
At4g33740 putative protein 35 0.023
At3g58050 putative protein 35 0.023
At5g10660 vacuolar calcium binding protein - like 35 0.030
At4g26630 unknown protein 35 0.030
At5g03710 putative protein 33 0.066
>At5g25590 unknown protein (At5g25590)
Length = 775
Score = 40.8 bits (94), Expect = 4e-04
Identities = 27/87 (31%), Positives = 45/87 (51%), Gaps = 7/87 (8%)
Query: 40 ENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEEFPQERPYTEEAETV 99
EN S+ F+ + EE ++ I ++ + K+VEE+EP PE+ EE E
Sbjct: 222 ENQSSHFQFNEEDDEEEEEEERSGIYRKKSGSGKVVEEMEPKTPEK-------VEEEEEE 274
Query: 100 ENEEVMEVTEKEDRILIKEESMEEKGK 126
+ EE E E+E+ ++ E ++KGK
Sbjct: 275 DEEEDEEEEEEEEEEVVVEVKKKKKGK 301
Score = 27.3 bits (59), Expect = 4.7
Identities = 16/59 (27%), Positives = 31/59 (52%), Gaps = 3/59 (5%)
Query: 96 AETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVEIDRIIDEICALFNKPKLGRIWT 154
AETVE + V+ R ++ +SM+ +VN++ D++ + AL + ++WT
Sbjct: 456 AETVEKTKAA-VSHLHTRYIVDMQSMDSTVSEVNRLRDDQLYPRLVALVE--GMAKMWT 511
>At1g20290 hypothetical protein
Length = 497
Score = 40.0 bits (92), Expect = 7e-04
Identities = 31/105 (29%), Positives = 48/105 (45%)
Query: 28 IQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEEFP 87
+QA + ++E EE + EE ++ EE+ EE + EE E EE
Sbjct: 361 VQAEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 420
Query: 88 QERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVE 132
+E EE E E EE E E+E+ +EE EE+ ++ + E
Sbjct: 421 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 465
Score = 39.7 bits (91), Expect = 0.001
Identities = 32/109 (29%), Positives = 49/109 (44%)
Query: 19 EMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEEL 78
E +E+ + + + ++E EE + EE ++ EE+ EE + EE
Sbjct: 375 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 434
Query: 79 EPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKK 127
E EE +E EE E E EE E E+E+ +EE EE+ KK
Sbjct: 435 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEKK 483
Score = 38.9 bits (89), Expect = 0.002
Identities = 32/116 (27%), Positives = 52/116 (44%)
Query: 17 QMEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVE 76
Q E +E+ + + + ++E EE + EE ++ EE+ EE + E
Sbjct: 362 QAEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 421
Query: 77 ELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVE 132
E E EE +E EE E E EE E E+E+ +EE EE+ ++ + E
Sbjct: 422 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 477
Score = 38.1 bits (87), Expect = 0.003
Identities = 32/115 (27%), Positives = 51/115 (43%)
Query: 19 EMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEEL 78
E +E+ + + + ++E EE + EE ++ EE+ EE + EE
Sbjct: 372 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 431
Query: 79 EPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVEI 133
E EE +E EE E E EE E E+E+ +EE EE+ ++ K I
Sbjct: 432 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEKKYMI 486
Score = 37.7 bits (86), Expect = 0.004
Identities = 30/104 (28%), Positives = 46/104 (43%)
Query: 29 QANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEEFPQ 88
+ I ++E EE + EE ++ EE+ EE + EE E EE +
Sbjct: 357 EGTIVQAEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 416
Query: 89 ERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVE 132
E EE E E EE E E+E+ +EE EE+ ++ + E
Sbjct: 417 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 460
Score = 37.7 bits (86), Expect = 0.004
Identities = 31/117 (26%), Positives = 53/117 (44%)
Query: 19 EMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEEL 78
E +E+ + + + ++E EE + EE ++ EE+ EE + EE
Sbjct: 366 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 425
Query: 79 EPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVEIDR 135
E EE +E EE E E EE E E+E+ +EE EE+ ++ + E ++
Sbjct: 426 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEK 482
Score = 37.4 bits (85), Expect = 0.005
Identities = 30/108 (27%), Positives = 48/108 (43%)
Query: 25 IMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPE 84
I+ + + ++E EE + EE ++ EE+ EE + EE E E
Sbjct: 360 IVQAEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 419
Query: 85 EFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVE 132
E +E EE E E EE E E+E+ +EE EE+ ++ + E
Sbjct: 420 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 467
Score = 37.4 bits (85), Expect = 0.005
Identities = 32/114 (28%), Positives = 51/114 (44%)
Query: 38 KQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEEFPQERPYTEEAE 97
++E EE + EE ++ EE+ EE + EE E EE +E EE E
Sbjct: 381 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 440
Query: 98 TVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVEIDRIIDEICALFNKPKLGR 151
E EE E E+E+ +EE EE+ ++ + E + ++ + K K R
Sbjct: 441 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEKKYMIVKKKKKKR 494
Score = 37.4 bits (85), Expect = 0.005
Identities = 31/114 (27%), Positives = 51/114 (44%)
Query: 19 EMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEEL 78
E +E+ + + + ++E EE + EE ++ EE+ EE + EE
Sbjct: 365 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 424
Query: 79 EPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVE 132
E EE +E EE E E EE E E+E+ +EE EE+ ++ + E
Sbjct: 425 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 478
Score = 27.7 bits (60), Expect = 3.6
Identities = 20/62 (32%), Positives = 30/62 (48%)
Query: 71 TMKMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNK 130
+M V E + EE +E EE E E EE E E+E+ +EE EE+ ++ +
Sbjct: 351 SMVHVGEGTIVQAEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 410
Query: 131 VE 132
E
Sbjct: 411 EE 412
>At5g40450 unknown protein
Length = 2910
Score = 38.9 bits (89), Expect = 0.002
Identities = 29/116 (25%), Positives = 55/116 (47%), Gaps = 6/116 (5%)
Query: 2 INMQDTNQRLKNLSFQMEMMQEQIMDIQANIQSTHGKQEN---NSNTFEEVGKIVEEGVD 58
+++++ N +N+ + EQ Q NI ++E N +F V K ++ +D
Sbjct: 2682 VSVEEKNNTSENIDHEAAKEIEQEEGKQTNIVKEEIREEEKEINQESFNNV-KETDDAID 2740
Query: 59 DTEEDI--ILEECSTMKMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKED 112
T+ +I I S K ++ EP + Q+R T E ++EN ++ E +K+D
Sbjct: 2741 KTQPEIRDIESLSSVSKTQDKPEPEYEVPNQQKREITNEVPSLENSKIEEELQKKD 2796
Score = 32.7 bits (73), Expect = 0.11
Identities = 32/117 (27%), Positives = 51/117 (43%), Gaps = 29/117 (24%)
Query: 40 ENNSNTFEEVGKIVEEGVDDTEEDIILEECSTM-KMVEELEPLHPEEFPQERPYTEEAET 98
E+ S EE K V+E +++ + I L E + + V +L PL E +P +E ET
Sbjct: 943 ESPSEVLEESSKTVDEKIEEKTDSIELGEIAQEERSVTDLTPLQEES---SQPNEQEKET 999
Query: 99 -----------VENEEVMEV--------------TEKEDRILIKEESMEEKGKKVNK 130
V+++EV+EV E E+ IKE E+ +K+ K
Sbjct: 1000 KLEKHEPTNEEVKSDEVIEVLSASPSKELEGETVVEAENIENIKENEEEQAAEKIQK 1056
Score = 32.3 bits (72), Expect = 0.15
Identities = 22/81 (27%), Positives = 40/81 (49%), Gaps = 3/81 (3%)
Query: 61 EEDIILEECSTMKMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEES 120
+E IIL + + EE PEE E+P EE +T E + ++ED I +
Sbjct: 120 DEKIILSDVTLENKKEEDTTGKPEEVSVEKPVIEEDQT---EAKHSLEQEEDIGNISKVL 176
Query: 121 MEEKGKKVNKVEIDRIIDEIC 141
+ KV++ +I++ ++ +C
Sbjct: 177 TDTTPVKVDEYDIEKSLNSVC 197
Score = 31.2 bits (69), Expect = 0.33
Identities = 28/124 (22%), Positives = 55/124 (43%), Gaps = 9/124 (7%)
Query: 25 IMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDD-TEEDIILEECSTMKMVEELEPLH- 82
++D+++ Q K E+ S EE K V+E +++ EE++ L + + LE
Sbjct: 852 LLDVKSVEQMQKPKLESPSEVSEETSKTVDEKIEEKPEEEVTLYQEGQVDGSYGLETKEE 911
Query: 83 ----PEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESM---EEKGKKVNKVEIDR 135
PE E EE ++ + + T + +++E S E+ +K + +E+
Sbjct: 912 TVSVPESIELEEQPQEERSVIDPTPLQKPTLESPSEVLEESSKTVDEKIEEKTDSIELGE 971
Query: 136 IIDE 139
I E
Sbjct: 972 IAQE 975
Score = 29.3 bits (64), Expect = 1.2
Identities = 30/113 (26%), Positives = 53/113 (46%), Gaps = 16/113 (14%)
Query: 39 QENNSNTFEEVGKIVEEGVDDTEEDIIL--EECSTMKMVEELEPLHPEEFPQERPYTEEA 96
+EN+S T K E +++TE + L EE +LE E+ QE P T +A
Sbjct: 1994 EENDSETLIAEAKKGNEEINETERTVALDHEEEFVNHEAPKLEETKDEK-SQEIPETAKA 2052
Query: 97 ETVENEEVME------------VTEKEDRILIK-EESMEEKGKKVNKVEIDRI 136
++ + V++K+D+ + EE +EE+ K+ +KV+ + I
Sbjct: 2053 TETTIDQTLPIGTSQADQTPSLVSDKDDQTPKQVEEILEEETKETHKVQAEDI 2105
Score = 27.7 bits (60), Expect = 3.6
Identities = 35/137 (25%), Positives = 63/137 (45%), Gaps = 31/137 (22%)
Query: 12 KNLSFQMEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECST 71
++++ + E +QE+ + +A+++ + ++ SN EE ++E V ++C T
Sbjct: 616 EDIAPKTEEIQERPSESKASLEPKE-EVDHISNETEEHEHVLERDV---------QQCET 665
Query: 72 MKM--VEELEPLHPEEFPQERPYTEEAET--------------VENEEVMEVTEK----- 110
++ VE E P +E TEEAET V+ E+ E TE
Sbjct: 666 IESEAVETKEDTQPSLDLKEDKETEEAETFKTVFSSDEVRSSAVQEEQFGEHTEPCSSEI 725
Query: 111 EDRILIKEESMEEKGKK 127
+D KEES+E K ++
Sbjct: 726 KDESHGKEESVEVKSQE 742
>At4g38065 hypothetical protein
Length = 1050
Score = 38.1 bits (87), Expect = 0.003
Identities = 30/118 (25%), Positives = 56/118 (47%), Gaps = 14/118 (11%)
Query: 2 INMQDTNQRLKNLSFQMEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTE 61
I MQ +R K + + Q Q+ + K + N +EE V + +D+T
Sbjct: 509 IQMQKEVERFKEMVEESSRFQTQMQE----------KMKEAENDYEEKLLQVCDALDNTN 558
Query: 62 EDIILEECSTMKMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEE 119
D++ E + + ++E L +E+ E ET E +E++E +EK R+L++E+
Sbjct: 559 IDLVAEREKVVSLTRQIESLGT---VKEKNLVMEKETQEYKEMLEESEK-CRVLLEEQ 612
>At5g22640 unknown protein
Length = 871
Score = 37.4 bits (85), Expect = 0.005
Identities = 20/84 (23%), Positives = 46/84 (53%), Gaps = 5/84 (5%)
Query: 40 ENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEEFPQERPYTEEAETV 99
E +EE +++++ ++ E +I LEE +E+++ ++ +E TE T
Sbjct: 596 EEKKKAYEERKEMIQQELELVEAEICLEEA-----IEDMDEELKKKEQEEEKKTEMGLTE 650
Query: 100 ENEEVMEVTEKEDRILIKEESMEE 123
E+E+V+ KE++++ +E ++E
Sbjct: 651 EDEDVLVPVYKEEKVVTAKEKIQE 674
>At3g58840 unknown protein
Length = 318
Score = 37.4 bits (85), Expect = 0.005
Identities = 40/163 (24%), Positives = 67/163 (40%), Gaps = 43/163 (26%)
Query: 5 QDTNQRLKNLSFQMEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVE---------- 54
++ +RL+ L+ ++E M+ D++A + G+ E +EE K +E
Sbjct: 44 RELKERLERLTGEIEEMK----DVEAEMNQRFGEMEKEIEEYEEEKKALEAISTRAVELE 99
Query: 55 --------------EGVDDTEEDIILEECSTMKMVEELEPLHPEEFPQERPYTEEAETVE 100
GVD T E++ + + ++VE+LE E + E + V
Sbjct: 100 TEVSNLHDDLITSLNGVDKTAEEVAELKKALAEIVEKLEGCEKEAEGLRKDRAEVEKRVR 159
Query: 101 NEE----VMEVTEKEDRILIKEESMEEKGKKVNKVEIDRIIDE 139
+ E V+EV E MEEK KK+ E R ID+
Sbjct: 160 DLERKIGVLEVRE-----------MEEKSKKLRSEEEMREIDD 191
>At1g04600 putative myosin heavy chain
Length = 1730
Score = 37.4 bits (85), Expect = 0.005
Identities = 31/139 (22%), Positives = 60/139 (42%), Gaps = 18/139 (12%)
Query: 4 MQDTNQRLKNLSFQMEMMQEQIMD---IQANIQSTHGKQENNSNTFEEVGKIVEEGVDDT 60
+QD ++ LS +EM + + ++ ++ S K + + +EE+ KI EE + D
Sbjct: 979 LQDMQLEIEELSKGLEMTNDLAAENEQLKESVSSLQNKIDESERKYEEISKISEERIKDE 1038
Query: 61 -----EEDIILEECSTMKM----------VEELEPLHPEEFPQERPYTEEAETVENEEVM 105
+ II E K+ ++EL+ H E P +E + + E V
Sbjct: 1039 VPVIDQSAIIKLETENQKLKALVSSMEEKIDELDRKHDETSPNITEKLKEDVSFDYEIVS 1098
Query: 106 EVTEKEDRILIKEESMEEK 124
+ + +R+ S+E+K
Sbjct: 1099 NLEAENERLKALVGSLEKK 1117
Score = 34.7 bits (78), Expect = 0.030
Identities = 28/127 (22%), Positives = 63/127 (49%), Gaps = 7/127 (5%)
Query: 15 SFQMEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKM 74
S ++E +Q + DI+ ++ T + + + V ++ + DT+E E
Sbjct: 919 SQEIEALQSVLTDIKLQLRDTQETKSKEISDLQSVLTDIKLQLRDTQETKSKEISDLQSA 978
Query: 75 VEELEPLHPEEFPQERPYTEEAETVENEEVME-VTEKEDRILIKEESMEEKGKKVNKVEI 133
+++++ L EE + T + ENE++ E V+ +++I + E K ++++K+
Sbjct: 979 LQDMQ-LEIEELSKGLEMTNDL-AAENEQLKESVSSLQNKI----DESERKYEEISKISE 1032
Query: 134 DRIIDEI 140
+RI DE+
Sbjct: 1033 ERIKDEV 1039
Score = 30.8 bits (68), Expect = 0.43
Identities = 27/134 (20%), Positives = 59/134 (43%), Gaps = 7/134 (5%)
Query: 18 MEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEE 77
+E E++ + +++ + NNS +E GK + + + ED ++ K+ +E
Sbjct: 1100 LEAENERLKALVGSLEKKINESGNNSTDEQEEGKYILKE-ESLTEDASIDNERVKKLADE 1158
Query: 78 LEPLHP------EEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKV 131
+ L+ ++ + EEA + E + + + E ++ + SM+ +KV+ +
Sbjct: 1159 NKDLNDLVSSLEKKIDETEKKYEEASRLCEERLKQALDAETGLIDLKTSMQRLEEKVSDM 1218
Query: 132 EIDRIIDEICALFN 145
E I AL N
Sbjct: 1219 ETAEQIRRQQALVN 1232
>At5g24880 glutamic acid-rich protein
Length = 443
Score = 36.6 bits (83), Expect = 0.008
Identities = 33/133 (24%), Positives = 59/133 (43%), Gaps = 8/133 (6%)
Query: 5 QDTNQRLKNLSFQMEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDI 64
+D N + L ++ ++ I + ++ Q N S E+V K ++E ++T E +
Sbjct: 261 KDCNAVVAELEEKLIKNEDDIEEKTEEMKEQDNNQANKSEEEEDVKKKIDE--NETPEKV 318
Query: 65 ILEECSTMKMVEELEPLHPEEFPQERPYTEEAETVENEEVM-----EVTEKEDRILIKEE 119
E ++ VEE EE +E E E E E+V E E+E++ +K +
Sbjct: 319 DTES-KEVESVEETTQEKEEEVKEEGKERVEEEEKEKEKVKEDDQKEKVEEEEKEKVKGD 377
Query: 120 SMEEKGKKVNKVE 132
+EK K+ E
Sbjct: 378 EEKEKVKEEESAE 390
Score = 36.2 bits (82), Expect = 0.010
Identities = 36/144 (25%), Positives = 68/144 (47%), Gaps = 12/144 (8%)
Query: 3 NMQDTNQRLKNLSFQMEMMQEQIMDIQANIQSTHGKQENNSNT--FEEVGKIVEEGVDDT 60
++Q ++ ++ ++ E + Q +QAN +S K E++ T VGK V +
Sbjct: 212 HLQVSDHDIEEPKYEKEEKEVQEKVVQAN-ESVEEKAESSGPTPVASPVGKDCNAVVAEL 270
Query: 61 EEDIILEECSTMKMVEELEPLHPEEFPQERPYTEEAETV-----ENEEVMEVTEKEDRIL 115
EE +I E + EE++ E+ + +EE E V ENE +V + +
Sbjct: 271 EEKLIKNEDDIEEKTEEMK----EQDNNQANKSEEEEDVKKKIDENETPEKVDTESKEVE 326
Query: 116 IKEESMEEKGKKVNKVEIDRIIDE 139
EE+ +EK ++V + +R+ +E
Sbjct: 327 SVEETTQEKEEEVKEEGKERVEEE 350
>At1g76770 putative heat shock protein
Length = 244
Score = 36.6 bits (83), Expect = 0.008
Identities = 29/79 (36%), Positives = 42/79 (52%), Gaps = 5/79 (6%)
Query: 51 KIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEK 110
KI EE ++ ++ I+EE T + E E + E P+E EEAE + EE EV E+
Sbjct: 136 KIEEEDEEEEMKEPIVEE-KTEEKTEPEEEIKEETKPEEE--NEEAEEPQREEEEEVVEE 192
Query: 111 --EDRILIKEESMEEKGKK 127
D KEE +E+K +K
Sbjct: 193 GTRDHEGKKEEEIEDKPRK 211
>At4g08710 putative protein
Length = 715
Score = 36.2 bits (82), Expect = 0.010
Identities = 38/135 (28%), Positives = 55/135 (40%), Gaps = 4/135 (2%)
Query: 30 ANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVE-ELEPLHPEEFPQ 88
A ++ +E T E I +E E +E + + E L PE +
Sbjct: 25 ATKETAPATKETAPATKETAPTITKETAPTKETAPATKETAPTRTEEPSLTEQDPENVEE 84
Query: 89 ERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVEIDRIIDEICALFNKPK 148
E EE E E EE E +E+ +EE EEK ++ N + D +L +P
Sbjct: 85 EESEEEEKEEEEKEEEEEEEGEEEEE--EEEEEEEKEEEENVGGEESSDDSTRSLGKRPP 142
Query: 149 LGRIW-TPHQLYFKF 162
RIW HQL +KF
Sbjct: 143 PMRIWMRRHQLQWKF 157
>At1g22530 unknown protein
Length = 683
Score = 36.2 bits (82), Expect = 0.010
Identities = 26/96 (27%), Positives = 49/96 (50%), Gaps = 8/96 (8%)
Query: 37 GKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEEF--PQERPYTE 94
GK+ S +F+E G + E + + E++ + E ++ E L+ EF P P
Sbjct: 71 GKEILQSESFKEEGYLASE-LQEAEKNALAELKELVR-----EALNKREFTAPPPPPAPV 124
Query: 95 EAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNK 130
+ E VE ++ E EK++ + +E+S+E + K+ K
Sbjct: 125 KEEKVEEKKTEETEEKKEEVKTEEKSLEAETKEEEK 160
>At2g36410 unknown protein
Length = 195
Score = 35.8 bits (81), Expect = 0.013
Identities = 25/136 (18%), Positives = 64/136 (46%), Gaps = 6/136 (4%)
Query: 7 TNQRLKNLSFQMEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIIL 66
T L + + ++++ M+++ IQ+ G+ E + + + +E D +++ +
Sbjct: 61 TRSALSAFRAKEDEIEKRRMEVRERIQAQLGRVEQETKRLSTIREELESMADPMRKEVSV 120
Query: 67 EECSTMKMVEELEPLHPEEFPQERPYTEEAETV--ENEEVMEVTEKEDRILIKEESMEEK 124
+ +EL+PL +ER Y E +T +N E +++ K L++ E + +
Sbjct: 121 VRKKIDSVNKELKPLGSTVQKKEREYKEALDTFNEKNREKVQLITK----LMEMEQLVGE 176
Query: 125 GKKVNKVEIDRIIDEI 140
+K+ ++++ + I
Sbjct: 177 SEKLRMIKLEELSKSI 192
>At3g28770 hypothetical protein
Length = 2081
Score = 35.4 bits (80), Expect = 0.017
Identities = 30/102 (29%), Positives = 49/102 (47%), Gaps = 5/102 (4%)
Query: 39 QENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEEFPQERPYTEEAET 98
+E S T EE K ++ D E+ EE + K EE L ++ +E +E+E
Sbjct: 1016 EEKKSKTKEEAKKEKKKSQDKKREEKDSEERKSKKEKEESRDLKAKKKEEETKEKKESEN 1075
Query: 99 VENEEVMEVTEKED-RILIKEESMEEKGK----KVNKVEIDR 135
++++ + E ED + + KEE +EK K K K E D+
Sbjct: 1076 HKSKKKEDKKEHEDNKSMKKEEDKKEKKKHEESKSRKKEEDK 1117
Score = 30.4 bits (67), Expect = 0.56
Identities = 20/94 (21%), Positives = 47/94 (49%), Gaps = 6/94 (6%)
Query: 39 QENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEEFPQERPYTEEAET 98
+ENN ++ E+ +E D+++D +++ +E ++ E ++ + +
Sbjct: 699 KENNKDSMEDKKLENKESQTDSKDDKSVDD------KQEEAQIYGGESKDDKSVEAKGKK 752
Query: 99 VENEEVMEVTEKEDRILIKEESMEEKGKKVNKVE 132
E++E + E+R+ KEE+++ K+ KVE
Sbjct: 753 KESKENKKTKTNENRVRNKEENVQGNKKESEKVE 786
>At1g80810 unknown protein
Length = 826
Score = 35.4 bits (80), Expect = 0.017
Identities = 32/131 (24%), Positives = 65/131 (49%), Gaps = 5/131 (3%)
Query: 11 LKNLSFQMEMM-QEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVD-DTEEDIILEE 68
LK L+ + + +EQ + ++A +QE + ++ + ++GV+ +T+E+ ++
Sbjct: 689 LKELNAETDRTAEEQEVSLEAESDDRSEEQEYEDDCSDKKEQSQDKGVEAETKEEE--KQ 746
Query: 69 CSTMKMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKV 128
+ E E EE P+ R T++ E E EE E+ ED ++E +++K
Sbjct: 747 YPNSEGESEGEDSESEEEPKWRE-TDDMEDDEEEEEEEIDHMEDEAEEEKEEVDDKEASA 805
Query: 129 NKVEIDRIIDE 139
N EI++ +E
Sbjct: 806 NMSEIEKEEEE 816
Score = 26.6 bits (57), Expect = 8.1
Identities = 30/96 (31%), Positives = 41/96 (42%), Gaps = 22/96 (22%)
Query: 34 STHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEEFPQERPYT 93
++ G+ E + EE K E DD E+D EE M +E E
Sbjct: 749 NSEGESEGEDSESEEEPKWRE--TDDMEDDEEEEEEEIDHMEDEAE-------------- 792
Query: 94 EEAETVENEEV---MEVTEKEDRILIKEESMEEKGK 126
EE E V+++E M EKE+ +EE EEK K
Sbjct: 793 EEKEEVDDKEASANMSEIEKEEE---EEEEDEEKRK 825
>At1g13220 putative nuclear matrix constituent protein
Length = 1128
Score = 35.4 bits (80), Expect = 0.017
Identities = 36/148 (24%), Positives = 64/148 (42%), Gaps = 20/148 (13%)
Query: 12 KNLSFQMEMM---QEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILE- 67
K LS + + + +E + D+Q I+ + EE K +E ++ EE + L+
Sbjct: 465 KRLSLEKQQLLSDKESLEDLQQEIEKIRAEMTKKEEMIEEECKSLEIKKEEREEYLRLQS 524
Query: 68 ------------ECSTMKMVEELEPLHPEEFPQERPYTEEAETVENEE---VMEVTEKED 112
E K VE L+ E F +E +E + V N+E + E EK +
Sbjct: 525 ELKSQIEKSRVHEEFLSKEVENLKQ-EKERFEKEWEILDEKQAVYNKERIRISEEKEKFE 583
Query: 113 RILIKEESMEEKGKKVNKVEIDRIIDEI 140
R + E +K + +V+I + +D+I
Sbjct: 584 RFQLLEGERLKKEESALRVQIMQELDDI 611
Score = 30.8 bits (68), Expect = 0.43
Identities = 29/138 (21%), Positives = 61/138 (44%), Gaps = 15/138 (10%)
Query: 4 MQDTNQRLKNLSFQMEMMQEQIMDIQANIQSTHGKQENNSNT-FEEVGKIVEEGVDDTEE 62
+Q+ N +++ S + ++++ A + S +G+ + N + K+ E +E
Sbjct: 173 IQEENSKIRLSS------EAKLVEANALVASVNGRSSDVENKIYSAESKLAEATRKSSEL 226
Query: 63 DIILEECSTMKMVEELEPL--------HPEEFPQERPYTEEAETVENEEVMEVTEKEDRI 114
+ L+E T + V + E L + F ++R Y E E + +TE++ +
Sbjct: 227 KLRLKEVETRESVLQQERLSFTKERESYEGTFQKQREYLNEWEKKLQGKEESITEQKRNL 286
Query: 115 LIKEESMEEKGKKVNKVE 132
+EE + E KK+ E
Sbjct: 287 NQREEKVNEIEKKLKLKE 304
Score = 28.9 bits (63), Expect = 1.6
Identities = 30/111 (27%), Positives = 57/111 (51%), Gaps = 18/111 (16%)
Query: 32 IQSTHGKQENNSNTFEEV-----GKIVEEGVDDTEEDIILEECSTMKMVE-ELEPLHPEE 85
+Q T +EN FEE G +++ +DD +E + KM+E ELE EE
Sbjct: 345 LQITLLAKENELRAFEEKLIAREGTEIQKLIDDQKEVL------GSKMLEFELEC---EE 395
Query: 86 FPQ--ERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVEID 134
+ ++ + E +E ++V E+ E+++ + ++M +K +VN+ E+D
Sbjct: 396 IRKSLDKELQRKIEELERQKV-EIDHSEEKLEKRNQAMNKKFDRVNEKEMD 445
>At4g33740 putative protein
Length = 227
Score = 35.0 bits (79), Expect = 0.023
Identities = 34/134 (25%), Positives = 65/134 (48%), Gaps = 14/134 (10%)
Query: 15 SFQMEMMQEQIMDIQANIQSTHGKQEN-------NSNTFEEVGKIVEEGVDDTEEDIILE 67
++ + ++ + + + + +HG +E N N EEV K EE V + +E+ +
Sbjct: 78 NYNQKENEKHVEEDEDEEEISHGGEEKEKKSKVENGNHEEEVEKDEEEEVAEDDEEDKNK 137
Query: 68 ECSTMKMVEELEPLHPEEFPQERPYTEEA-ETVENEEVMEVTEKEDRILIKEESMEEKGK 126
+ + +E E H E+ E+ ++ A +T +++E +E EKE + +EK K
Sbjct: 138 QGEEVAEEDEEENKHEEDEIDEQDQSKNAGDTDKDDETLE-EEKESGM----SENDEKEK 192
Query: 127 KVNKV-EIDRIIDE 139
+ N EID +DE
Sbjct: 193 ETNHADEIDMTVDE 206
>At3g58050 putative protein
Length = 1209
Score = 35.0 bits (79), Expect = 0.023
Identities = 22/69 (31%), Positives = 35/69 (49%), Gaps = 18/69 (26%)
Query: 79 EPLHPEEFPQERPYTE--------EAETVENEEVME----------VTEKEDRILIKEES 120
EP+ P+++P+ Y+E +AE +E+ V E VTEK D I +K+E+
Sbjct: 845 EPMEPKKYPRSNSYSEVTVRCSTFKAEEIEDAIVAENSSDLLSQCKVTEKLDNIKLKDEN 904
Query: 121 MEEKGKKVN 129
E G+ N
Sbjct: 905 SMESGETKN 913
>At5g10660 vacuolar calcium binding protein - like
Length = 407
Score = 34.7 bits (78), Expect = 0.030
Identities = 29/124 (23%), Positives = 62/124 (49%), Gaps = 13/124 (10%)
Query: 22 QEQIMDIQANIQ-STHGKQ--ENNSNTFEEVGKIVEE--------GVDDTEEDIILEECS 70
+E+I+ ++ ++ S HG++ E + + F + + EE ++ + ++I E+ S
Sbjct: 206 EEEIVKVETDVHISDHGEEPKEEDKDQFAQPDESGEEKETSPVAASTEEQKGELIDEDKS 265
Query: 71 TMKMVEELEPLHPEE--FPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKV 128
T ++ E EP + EE +E ++++ EN E + T + + EES EE+ ++
Sbjct: 266 TEQIEEPKEPENIEENNSEEEEEVKKKSDDEENSETVATTTDMNEAVNVEESKEEEKEEA 325
Query: 129 NKVE 132
E
Sbjct: 326 EVKE 329
>At4g26630 unknown protein
Length = 763
Score = 34.7 bits (78), Expect = 0.030
Identities = 24/85 (28%), Positives = 38/85 (44%), Gaps = 1/85 (1%)
Query: 27 DIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEEF 86
DIQ +G ++ N+ +E G +V+E T+ D +E K VE +E E+
Sbjct: 162 DIQHEADKANGTKDGNTGDIKEEGTLVDED-KGTDMDEKVENGDENKQVENVEGKEKEDK 220
Query: 87 PQERPYTEEAETVENEEVMEVTEKE 111
+ + EA E +E EKE
Sbjct: 221 EENKTKEVEAAKAEVDESKVEDEKE 245
>At5g03710 putative protein
Length = 81
Score = 33.5 bits (75), Expect = 0.066
Identities = 23/64 (35%), Positives = 32/64 (49%)
Query: 76 EELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVEIDR 135
EE E EE +E EE E E EE E E+E+ +EE EE+ ++ + E DR
Sbjct: 9 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEDR 68
Query: 136 IIDE 139
+E
Sbjct: 69 EREE 72
Score = 31.2 bits (69), Expect = 0.33
Identities = 23/66 (34%), Positives = 31/66 (46%)
Query: 54 EEGVDDTEEDIILEECSTMKMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDR 113
EE ++ EE+ EE + EE E EE +E EE E E EE E E+E+
Sbjct: 7 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 66
Query: 114 ILIKEE 119
+EE
Sbjct: 67 DREREE 72
Score = 30.8 bits (68), Expect = 0.43
Identities = 22/69 (31%), Positives = 32/69 (45%)
Query: 56 GVDDTEEDIILEECSTMKMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRIL 115
G ++ EE+ EE + EE E EE +E EE E E EE E E+E+
Sbjct: 5 GEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 64
Query: 116 IKEESMEEK 124
++ EE+
Sbjct: 65 EEDREREER 73
Score = 30.4 bits (67), Expect = 0.56
Identities = 20/60 (33%), Positives = 30/60 (49%)
Query: 76 EELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVEIDR 135
EE E EE +E EE E E EE E E+E+ +EE EE+ ++ ++ +R
Sbjct: 14 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEDREREER 73
Score = 30.0 bits (66), Expect = 0.73
Identities = 20/57 (35%), Positives = 28/57 (49%)
Query: 76 EELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVE 132
EE E EE +E EE E E EE E E+E+ +EE EE+ ++ + E
Sbjct: 6 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 62
Score = 30.0 bits (66), Expect = 0.73
Identities = 20/57 (35%), Positives = 28/57 (49%)
Query: 76 EELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVE 132
EE E EE +E EE E E EE E E+E+ +EE EE+ ++ + E
Sbjct: 8 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 64
Score = 30.0 bits (66), Expect = 0.73
Identities = 20/57 (35%), Positives = 28/57 (49%)
Query: 76 EELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVE 132
EE E EE +E EE E E EE E E+E+ +EE EE+ ++ + E
Sbjct: 7 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 63
Score = 28.5 bits (62), Expect = 2.1
Identities = 20/61 (32%), Positives = 29/61 (46%)
Query: 72 MKMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKV 131
M+ E E EE +E EE E E EE E E+E+ +EE EE+ ++ +
Sbjct: 1 MEFWGEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 60
Query: 132 E 132
E
Sbjct: 61 E 61
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.132 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,301,724
Number of Sequences: 26719
Number of extensions: 202127
Number of successful extensions: 1495
Number of sequences better than 10.0: 197
Number of HSP's better than 10.0 without gapping: 46
Number of HSP's successfully gapped in prelim test: 153
Number of HSP's that attempted gapping in prelim test: 1096
Number of HSP's gapped (non-prelim): 367
length of query: 184
length of database: 11,318,596
effective HSP length: 93
effective length of query: 91
effective length of database: 8,833,729
effective search space: 803869339
effective search space used: 803869339
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC119412.2


