

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119412.1 - phase: 0
(411 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
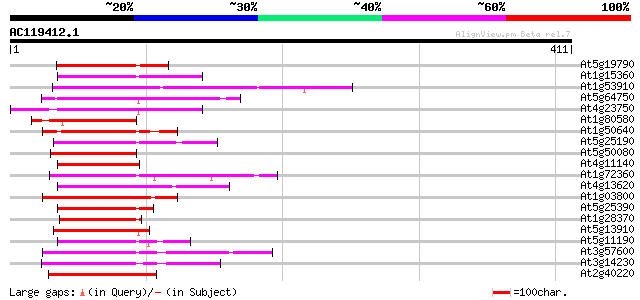
Score E
Sequences producing significant alignments: (bits) Value
At5g19790 AP2 domain containing protein RAP2.11 117 9e-27
At1g15360 ethylene responsive element like protein 90 2e-18
At1g53910 unknown protein 89 4e-18
At5g64750 unknown protein 86 5e-17
At4g23750 putative Ap2 domain protein 86 5e-17
At1g80580 unknown protein 86 5e-17
At1g50640 ethylene responsive element binding factor 3 86 5e-17
At5g25190 ethylene-responsive element - like protein 83 3e-16
At5g50080 putative protein 82 4e-16
At4g11140 unknown protein with Ap2 domain 82 4e-16
At1g72360 putative AP2 domain transcription factor 82 4e-16
At4g13620 putative protein 81 1e-15
At1g03800 unknown protein 80 2e-15
At5g25390 AP2 domain containing protein 80 2e-15
At1g28370 putative ethylene responsive element binding factor 4 ... 80 2e-15
At5g13910 AP2/EREBP-like transcription factor LEAFY PETIOLE (gb|... 80 3e-15
At5g11190 putative protein 80 3e-15
At3g57600 AP2 transcription factor - like protein 80 3e-15
At3g14230 transcription factor EREBP like protein 80 3e-15
At2g40220 AP2 domain transcription factor (ABI4:abscisic acid-in... 80 3e-15
>At5g19790 AP2 domain containing protein RAP2.11
Length = 253
Score = 117 bits (294), Expect = 9e-27
Identities = 59/82 (71%), Positives = 68/82 (81%), Gaps = 2/82 (2%)
Query: 35 RKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFW 94
+ +FVGVRQRPSG+WVAEIKDT QKIR+WLGTF+TAEEAARAYDEAACLLRG+NTRTNF
Sbjct: 19 KTKFVGVRQRPSGKWVAEIKDTTQKIRMWLGTFETAEEAARAYDEAACLLRGSNTRTNF- 77
Query: 95 PCSQSSTSPALSSKITNLLLQR 116
+ + LS KI NLL Q+
Sbjct: 78 -ANHFPNNSQLSLKIRNLLHQK 98
>At1g15360 ethylene responsive element like protein
Length = 199
Score = 90.1 bits (222), Expect = 2e-18
Identities = 49/106 (46%), Positives = 63/106 (59%), Gaps = 1/106 (0%)
Query: 36 KRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFWP 95
K+F GVRQR G WVAEI+ + K R+WLGTF+TAEEAARAYDEAA L+ G N +TNF P
Sbjct: 5 KKFRGVRQRHWGSWVAEIRHPLLKRRIWLGTFETAEEAARAYDEAAVLMSGRNAKTNF-P 63
Query: 96 CSQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSSKYTSLLGSP 141
+ ++T K + ++S SS S+K SP
Sbjct: 64 LNNNNTGETSEGKTDISASSTMSSSTSSSSLSSILSAKLRKCCKSP 109
>At1g53910 unknown protein
Length = 358
Score = 89.0 bits (219), Expect = 4e-18
Identities = 69/222 (31%), Positives = 109/222 (49%), Gaps = 5/222 (2%)
Query: 32 RRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRT 91
R+ + ++ G+RQRP G+W AEI+D + R+WLGTF TAEEAARAYD AA +RG+ +
Sbjct: 119 RKRKNQYRGIRQRPWGKWAAEIRDPREGARIWLGTFKTAEEAARAYDAAARRIRGSKAKV 178
Query: 92 NFWPCSQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSSKYTSLLGSPEERREQQRAN 151
NF + + S S K L + + + N N S + +S + E + Q N
Sbjct: 179 NFPEENMKANSQKRSVKAN--LQKPVAKPNPNPSPALVQNSNISFENMCFMEEKHQVSNN 236
Query: 152 DHDRQEMQQQVDAYGEVSKDFSIDQFTDFLNDHEDYSTSNNELINDTAQFDYISSSFESC 211
++++ M VDA + FS DQ ++ D ++ S+ I I+++ +
Sbjct: 237 NNNQFGMTNSVDAGCNGYQYFSSDQGSNSF-DCSEFGWSDQAPITPDISSAVINNNNSAL 295
Query: 212 LTE--NNGCKEDEMNMESKSNNVTETSSGDSNSEDGEEEVNN 251
E N K M+ E+ NN +S D +ED +N
Sbjct: 296 FFEEANPAKKLKSMDFETPYNNTEWDASLDFLNEDAVTTQDN 337
>At5g64750 unknown protein
Length = 391
Score = 85.5 bits (210), Expect = 5e-17
Identities = 57/160 (35%), Positives = 76/160 (46%), Gaps = 19/160 (11%)
Query: 24 EAASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACL 83
E +S G + R+R+ GVRQRP G+W AEI+D + RVWLGTFD AE AARAYDEAA
Sbjct: 172 ETSSFSGDQP-RRRYRGVRQRPWGKWAAEIRDPFKAARVWLGTFDNAESAARAYDEAALR 230
Query: 84 LRGANTRTNF--------------WPCSQSSTSPALSSKITNLLLQRLKERNMNNSCSSF 129
RG + NF P Q++ S+ + L R +N
Sbjct: 231 FRGNKAKLNFPENVKLVRPASTEAQPVHQTAAQRPTQSRNSGSTTTLLPIRPASNQSVHS 290
Query: 130 SSSKYTSLLGSPEERREQQRANDHDRQEMQQQVDAYGEVS 169
+ L E R+QQ+ H QQ +D Y ++S
Sbjct: 291 QPLMQSYNLSYSEMARQQQQFQQHH----QQSLDLYDQMS 326
>At4g23750 putative Ap2 domain protein
Length = 343
Score = 85.5 bits (210), Expect = 5e-17
Identities = 52/143 (36%), Positives = 74/143 (51%), Gaps = 7/143 (4%)
Query: 1 MARKRKVSEAVEEGSMVWDDMMKEAASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQKI 60
M K++ AV+ S V + + G K+F GVRQRP G+W AEI+D ++++
Sbjct: 90 MESKKRQKRAVKSESTVSPVVSATTTTTG-----EKKFRGVRQRPWGKWAAEIRDPLKRV 144
Query: 61 RVWLGTFDTAEEAARAYDEAACLLRGANTRTNF--WPCSQSSTSPALSSKITNLLLQRLK 118
R+WLGT++TAEEAA YD AA LRG + TNF P + + S + + K
Sbjct: 145 RLWLGTYNTAEEAAMVYDNAAIQLRGPDALTNFSVTPTTATEKKAPPPSPVKKKKKKNNK 204
Query: 119 ERNMNNSCSSFSSSKYTSLLGSP 141
+ + SS S S L SP
Sbjct: 205 SKKSVTASSSISRSSSNDCLCSP 227
>At1g80580 unknown protein
Length = 256
Score = 85.5 bits (210), Expect = 5e-17
Identities = 48/83 (57%), Positives = 55/83 (65%), Gaps = 10/83 (12%)
Query: 17 VWDDMMKEAASLGGARRVRKR------FVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTA 70
V D + KE GG +R RK F+GVR+RP GRW AEI+D I + R WLGTFDTA
Sbjct: 93 VSDHLTKE----GGVKRGRKMPQKTGGFMGVRKRPWGRWSAEIRDRIGRCRHWLGTFDTA 148
Query: 71 EEAARAYDEAACLLRGANTRTNF 93
EEAARAYD AA LRG +TNF
Sbjct: 149 EEAARAYDAAARRLRGTKAKTNF 171
>At1g50640 ethylene responsive element binding factor 3
Length = 225
Score = 85.5 bits (210), Expect = 5e-17
Identities = 50/99 (50%), Positives = 62/99 (62%), Gaps = 9/99 (9%)
Query: 25 AASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLL 84
AA G + +R F GVR+RP GR+ AEI+D +K RVWLGTFD+AEEAARAYD AA L
Sbjct: 17 AAINGSVKEIR--FRGVRKRPWGRFAAEIRDPWKKARVWLGTFDSAEEAARAYDSAARNL 74
Query: 85 RGANTRTNFWPCSQSSTSPALSSKITNLLLQRLKERNMN 123
RG +TNF P SS P NL +++ +N N
Sbjct: 75 RGPKAKTNF-PIDSSSPPP------PNLRFNQIRNQNQN 106
>At5g25190 ethylene-responsive element - like protein
Length = 181
Score = 82.8 bits (203), Expect = 3e-16
Identities = 49/120 (40%), Positives = 70/120 (57%), Gaps = 5/120 (4%)
Query: 33 RVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTN 92
R ++RF GVRQR G WV+EI+ + K R+WLGTF+TAE+AARAYDEAA L+ G RTN
Sbjct: 3 RPQQRFRGVRQRHWGSWVSEIRHPLLKTRIWLGTFETAEDAARAYDEAARLMCGPRARTN 62
Query: 93 FWPCSQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSSKYTSLLGSPEERREQQRAND 152
F P + ++ + S ++ L +L + M +S +K T + R Q +D
Sbjct: 63 F-PYNPNAIPTSSSKLLSATLTAKLHKCYM----ASLQMTKQTQTQTQTQTARSQSADSD 117
>At5g50080 putative protein
Length = 220
Score = 82.4 bits (202), Expect = 4e-16
Identities = 39/63 (61%), Positives = 46/63 (72%)
Query: 31 ARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTR 90
AR ++R+ GVRQRP G+W AEI+D + RVWLGTFDTAE AARAYDEAA RG +
Sbjct: 70 AREKKRRYRGVRQRPWGKWAAEIRDPHRAARVWLGTFDTAEAAARAYDEAALRFRGNKAK 129
Query: 91 TNF 93
NF
Sbjct: 130 LNF 132
>At4g11140 unknown protein with Ap2 domain
Length = 287
Score = 82.4 bits (202), Expect = 4e-16
Identities = 38/60 (63%), Positives = 45/60 (74%)
Query: 36 KRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFWP 95
++F GVRQRP G+W AEI+D +++RVWLGTFDTAEEAA YD AA LRG N NF P
Sbjct: 85 RKFRGVRQRPWGKWAAEIRDPSRRVRVWLGTFDTAEEAAIVYDNAAIQLRGPNAELNFPP 144
>At1g72360 putative AP2 domain transcription factor
Length = 262
Score = 82.4 bits (202), Expect = 4e-16
Identities = 61/190 (32%), Positives = 95/190 (49%), Gaps = 27/190 (14%)
Query: 30 GARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANT 89
G ++ R+ G+R+RP GRW AEI+D I+ +RVWLGTF+TAEEAARAYD A +RGA
Sbjct: 68 GKKKQSSRYKGIRRRPWGRWAAEIRDPIKGVRVWLGTFNTAEEAARAYDLEAKRIRGAKA 127
Query: 90 RTNFWPCSQSSTSPA--------------------LSSKITNLLLQRLKERNMNNSCSSF 129
+ NF P S A +SS ++ L L E N ++
Sbjct: 128 KLNF-PNESSGKRKAKAKTVQQVEENHEADLDVAVVSSAPSSSCLDFLWEENNPDTLLID 186
Query: 130 SSSKYTSLLGSPEERRE---QQRANDHDRQEMQQQVDAYGEVSKDFSIDQFTDFLNDHED 186
+ ++G ++ E + AN+ D + +++ A+ ++ FS FT+ + D
Sbjct: 187 TQWLEDIIMGDANKKHEPNDSEEANNVDASLLSEELLAFENQTEYFSQMPFTE---GNCD 243
Query: 187 YSTSNNELIN 196
STS + L +
Sbjct: 244 SSTSLSSLFD 253
>At4g13620 putative protein
Length = 388
Score = 80.9 bits (198), Expect = 1e-15
Identities = 48/126 (38%), Positives = 72/126 (57%), Gaps = 2/126 (1%)
Query: 36 KRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFWP 95
K + GVRQR G+WVAEI+ + RVWLGTF+TAE+AA AYD AA +LRG NF
Sbjct: 230 KLYRGVRQRHWGKWVAEIRLPRNRTRVWLGTFETAEQAAMAYDTAAYILRGEFAHLNFPD 289
Query: 96 CSQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSSKYTSLLGSPEERREQQRANDHDR 155
S +L I +LL ++++ +++S S S S +G+PE++ + +
Sbjct: 290 LKHQLKSGSLRCMIASLLESKIQQ--ISSSQVSNSPSPPPPKVGTPEQKNHHMKMESGED 347
Query: 156 QEMQQQ 161
M++Q
Sbjct: 348 VMMKKQ 353
>At1g03800 unknown protein
Length = 245
Score = 80.5 bits (197), Expect = 2e-15
Identities = 44/99 (44%), Positives = 60/99 (60%), Gaps = 2/99 (2%)
Query: 25 AASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLL 84
A + GG + R+ GVR+RP GR+ AEI+D ++K RVWLG+F+T EEAARAYD AA
Sbjct: 40 AVNDGGEKSKEVRYRGVRRRPWGRYAAEIRDPVKKKRVWLGSFNTGEEAARAYDSAAIRF 99
Query: 85 RGANTRTNFWPCSQSSTSPALSSKITNLLLQRLKERNMN 123
RG+ TNF S A + + N L + + + N N
Sbjct: 100 RGSKATTNFPLIGYYGISSA--TPVNNNLSETVSDGNAN 136
>At5g25390 AP2 domain containing protein
Length = 189
Score = 80.1 bits (196), Expect = 2e-15
Identities = 41/70 (58%), Positives = 52/70 (73%), Gaps = 1/70 (1%)
Query: 36 KRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFWP 95
K+F GVRQR G WV+EI+ + K RVWLGTFDTAE AARAYD+AA L+ G + +TNF P
Sbjct: 5 KKFRGVRQRQWGSWVSEIRHPLLKRRVWLGTFDTAETAARAYDQAAVLMNGQSAKTNF-P 63
Query: 96 CSQSSTSPAL 105
+S+ S +L
Sbjct: 64 VIKSNGSNSL 73
>At1g28370 putative ethylene responsive element binding factor 4
protein
Length = 166
Score = 80.1 bits (196), Expect = 2e-15
Identities = 39/60 (65%), Positives = 46/60 (76%), Gaps = 1/60 (1%)
Query: 37 RFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFWPC 96
R+ GVR+RP GR+ AEI+D +K RVWLGTFDT EEAARAYD+ A RGA +TNF PC
Sbjct: 19 RYRGVRKRPWGRYAAEIRDPFKKSRVWLGTFDTPEEAARAYDKRAIEFRGAKAKTNF-PC 77
>At5g13910 AP2/EREBP-like transcription factor LEAFY PETIOLE
(gb|AAF32292.1)
Length = 211
Score = 79.7 bits (195), Expect = 3e-15
Identities = 44/74 (59%), Positives = 50/74 (67%), Gaps = 4/74 (5%)
Query: 33 RVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTN 92
+V RF+GVR+RP GR+ AEI+D K R WLGTFDTAEEAA AYD AA +RG RTN
Sbjct: 15 QVGTRFLGVRRRPWGRYAAEIRDPTTKERHWLGTFDTAEEAALAYDRAARSMRGTRARTN 74
Query: 93 F----WPCSQSSTS 102
F P S S TS
Sbjct: 75 FVYSDMPPSSSVTS 88
>At5g11190 putative protein
Length = 189
Score = 79.7 bits (195), Expect = 3e-15
Identities = 47/106 (44%), Positives = 60/106 (56%), Gaps = 14/106 (13%)
Query: 36 KRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFWP 95
++F GVRQR G WV+EI+ + K RVWLGTF+TAE AARAYD+AA L+ G N +TNF P
Sbjct: 5 RKFRGVRQRQWGSWVSEIRHPLLKRRVWLGTFETAEAAARAYDQAALLMNGQNAKTNF-P 63
Query: 96 CSQSS---------TSPALSSKITNLLLQRLKERNMNNSCSSFSSS 132
+S SP +S K L L + SC + S
Sbjct: 64 VVKSEEGSDHVKDVNSPLMSPK----SLSELLNAKLRKSCKDLTPS 105
>At3g57600 AP2 transcription factor - like protein
Length = 277
Score = 79.7 bits (195), Expect = 3e-15
Identities = 57/168 (33%), Positives = 81/168 (47%), Gaps = 9/168 (5%)
Query: 25 AASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLL 84
A GG + ++ GVRQR G+WVAEI++ ++ R+WLG+F TAEEAA AYDEAA L
Sbjct: 15 ARGKGGPQNALCQYRGVRQRTWGKWVAEIREPKKRARLWLGSFATAEEAAMAYDEAALKL 74
Query: 85 RGANTRTNFWPCSQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSSKYTSLLGSPEER 144
G + N P Q +T P+LS+ QR K S F S ++ P
Sbjct: 75 YGHDAYLNL-PHLQRNTRPSLSNS------QRFKWVPSRKFISMFPSCGMLNVNAQPSVH 127
Query: 145 REQQRANDHDRQEMQQQVDAYGEVSKDFSIDQFTDFLNDHEDYSTSNN 192
QQR + + + Q +Y S T FL++ ++N
Sbjct: 128 IIQQRLEELKKTGLLSQ--SYSSSSSSTESKTNTSFLDEKTSKGETDN 173
>At3g14230 transcription factor EREBP like protein
Length = 375
Score = 79.7 bits (195), Expect = 3e-15
Identities = 48/131 (36%), Positives = 68/131 (51%), Gaps = 10/131 (7%)
Query: 24 EAASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACL 83
E A R+ + ++ G+RQRP G+W AEI+D + R WLGTFDTAEEAARAYD AA
Sbjct: 110 EQAEKSSKRKRKNQYRGIRQRPWGKWAAEIRDPRKGSREWLGTFDTAEEAARAYDAAARR 169
Query: 84 LRGANTRTNFWPCSQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSSKYTSLLGSPEE 143
+RG + NF +P++ S+ +R + N S +K +L+ P
Sbjct: 170 IRGTKAKVNF----PEEKNPSVVSQ------KRPSAKTNNLQKSVAKPNKSVTLVQQPTH 219
Query: 144 RREQQRANDHD 154
+Q N D
Sbjct: 220 LSQQYCNNSFD 230
>At2g40220 AP2 domain transcription factor (ABI4:abscisic
acid-insensitive 4 )
Length = 328
Score = 79.7 bits (195), Expect = 3e-15
Identities = 41/79 (51%), Positives = 52/79 (64%)
Query: 29 GGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGAN 88
GG + R+ GVRQR G+WVAEI++ ++ R WLGTF TAE+AARAYD AA L G+
Sbjct: 46 GGPDNSKFRYRGVRQRSWGKWVAEIREPRKRTRKWLGTFATAEDAARAYDRAAVYLYGSR 105
Query: 89 TRTNFWPCSQSSTSPALSS 107
+ N P S SS S + SS
Sbjct: 106 AQLNLTPSSPSSVSSSSSS 124
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.311 0.127 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,621,776
Number of Sequences: 26719
Number of extensions: 433974
Number of successful extensions: 2532
Number of sequences better than 10.0: 279
Number of HSP's better than 10.0 without gapping: 167
Number of HSP's successfully gapped in prelim test: 114
Number of HSP's that attempted gapping in prelim test: 2220
Number of HSP's gapped (non-prelim): 390
length of query: 411
length of database: 11,318,596
effective HSP length: 102
effective length of query: 309
effective length of database: 8,593,258
effective search space: 2655316722
effective search space used: 2655316722
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC119412.1


