

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119411.2 - phase: 0
(302 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
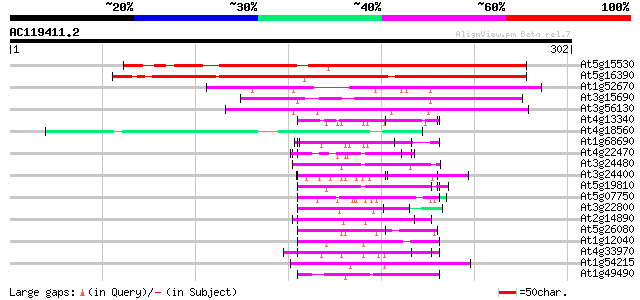
Score E
Sequences producing significant alignments: (bits) Value
At5g15530 biotin carboxyl carrier protein isoform 2 (BCCP2) 198 3e-51
At5g16390 biotin carboxyl carrier protein of acetyl-CoA carboxyl... 193 8e-50
At1g52670 unknown protein 63 2e-10
At3g15690 putative acetyl-CoA carboxylase biotin-containing subunit 62 5e-10
At3g56130 unknown protein 55 6e-08
At4g13340 extensin-like protein 53 2e-07
At4g18560 pherophorin - like protein 53 2e-07
At1g68690 protein kinase, putative 51 7e-07
At4g22470 extensin - like protein 50 1e-06
At3g24480 disease resistance protein, putative 50 1e-06
At3g24400 protein kinase, putative 50 1e-06
At5g19810 proline-rich protein 50 2e-06
At5g07750 putative protein 49 3e-06
At3g22800 hypothetical protein 49 4e-06
At2g14890 arabinogalactan-protein AGP9 49 4e-06
At5g26080 extensin - like protein 48 6e-06
At1g12040 leucine-rich repeat/extensin 1 (LRX1) 48 7e-06
At4g33970 extensin-like protein 47 1e-05
At1g54215 putative protein 47 1e-05
At1g49490 hypothetical protein 47 1e-05
>At5g15530 biotin carboxyl carrier protein isoform 2 (BCCP2)
Length = 255
Score = 198 bits (503), Expect = 3e-51
Identities = 117/219 (53%), Positives = 137/219 (62%), Gaps = 24/219 (10%)
Query: 62 WRSQALPNEVATIENSSNSVPVLINEPNGALPKEKDNHNGKPPGPSASTDASSVSTFMDQ 121
WR +A NEV SNS P+ NG K N P + +S FM +
Sbjct: 50 WRLRATTNEVV-----SNSTPMT----NGGYMNGKAKTNVPEP--------AELSEFMAK 92
Query: 122 VSELVKLVDSRDIMELELKQAGYELMIRKKEALQPPPVSQQPFPYPAYP--PSYQAPLPP 179
VS L+KLVDS+DI+ELELKQ E++IRKKEALQ Q P P Y P A
Sbjct: 93 VSGLLKLVDSKDIVELELKQLDCEIVIRKKEALQ-----QAVPPAPVYHSMPPVMADFSM 147
Query: 180 PPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLKCPMAGTFYRCPGPGEPPFVKVGDQV 239
PP + PPS P++ P +SSHP LK PMAGTFYR PGPGEPPFVKVGD+V
Sbjct: 148 PPAQPVALPPSPTPTSTPATAKPTSAPSSSHPPLKSPMAGTFYRSPGPGEPPFVKVGDKV 207
Query: 240 QKGQVVCIIEAMKLMNEIEADRSGTIVEVLVEDGKPVSV 278
QKGQ+VCIIEAMKLMNEIEA++SGTI+E+L EDGKPVSV
Sbjct: 208 QKGQIVCIIEAMKLMNEIEAEKSGTIMELLAEDGKPVSV 246
>At5g16390 biotin carboxyl carrier protein of acetyl-CoA carboxylase
precursor (BCCP) (sp|Q42533)
Length = 280
Score = 193 bits (491), Expect = 8e-50
Identities = 114/227 (50%), Positives = 149/227 (65%), Gaps = 13/227 (5%)
Query: 56 RKQFSVWRSQALPNEVATIENSSNSVPVLINEPNGALPKEKDN-HNGKPPGPSASTDASS 114
R + V ++Q+ N+V+T ++S ++ P+ A KEK++ P +T+ S
Sbjct: 54 RSSYPVVKAQS--NKVST---GASSNAAKVDGPSSAEGKEKNSLKESSASSPELATE-ES 107
Query: 115 VSTFMDQVSELVKLVDSRDIMELELKQAGYELMIRKKEALQPP--PVSQQPFPYPAYPPS 172
+S F+ QV+ LVKLVDSRDI+EL+LKQ EL+IRKKEAL P P S P P
Sbjct: 108 ISEFLTQVTTLVKLVDSRDIVELQLKQLDCELVIRKKEALPQPQAPASYVMMQQPNQPSY 167
Query: 173 YQAPLPPPPPVVASTPPSAPPSNVVPALP-PAKTNASSHPQLKCPMAGTFYRCPGPGEPP 231
Q PP P A+ PS P S P+ P PAK SS P +K PMAGTFYR P PGEPP
Sbjct: 168 AQQMAPPAAPAAAAPAPSTPASLPPPSPPTPAK---SSLPTVKSPMAGTFYRSPAPGEPP 224
Query: 232 FVKVGDQVQKGQVVCIIEAMKLMNEIEADRSGTIVEVLVEDGKPVSV 278
F+KVGD+VQKGQV+CI+EAMKLMNEIE+D +GT+V+++ EDGKPVS+
Sbjct: 225 FIKVGDKVQKGQVLCIVEAMKLMNEIESDHTGTVVDIVAEDGKPVSL 271
>At1g52670 unknown protein
Length = 274
Score = 63.2 bits (152), Expect = 2e-10
Identities = 65/208 (31%), Positives = 89/208 (42%), Gaps = 46/208 (22%)
Query: 107 SASTDASSVST-FMDQVSELVKLV----DSRDIMELELKQAGYELMIRKK--EALQPPPV 159
S T ++ V+T M + SE+ L+ DS I E ELK G+ L + +K + PPP
Sbjct: 75 SDETKSTVVNTHLMPKSSEVEALISEITDSSSIAEFELKLGGFRLYVARKLTDESSPPPQ 134
Query: 160 SQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSN---VVPALPPAKTNASS--HPQ-- 212
QP V AS P +N +L KT++SS PQ
Sbjct: 135 QIQPV------------------VAASATPEGVHTNGSATSSSLAITKTSSSSADRPQTL 176
Query: 213 -----------LKCPMAGTFYRCP---GPGEPPFVKVGDQVQKGQVVCIIEAMKLMNEIE 258
L+ P G F R G P K D V++GQV+C IE + +E
Sbjct: 177 ANKAADQGLVILQSPTVGYFRRSKTIKGKRTPTICKEKDIVKEGQVLCYIEQLGGQIPVE 236
Query: 259 ADRSGTIVEVLVEDGKPVSVGMVLSLTL 286
+D SG IV++L EDG+PV L L
Sbjct: 237 SDVSGEIVKILREDGEPVGYNDALITVL 264
>At3g15690 putative acetyl-CoA carboxylase biotin-containing subunit
Length = 263
Score = 61.6 bits (148), Expect = 5e-10
Identities = 49/159 (30%), Positives = 73/159 (45%), Gaps = 20/159 (12%)
Query: 125 LVKLVDSRDIMELELKQAGYELMIRKKEA----LQPPPVSQQPFPYPAYPPSYQAPLPPP 180
+ ++ DS I E ELK G+ L + + A LQPPP PA S P
Sbjct: 98 VTEICDSSSIAEFELKLGGFRLYVARNIADNSSLQPPPT-------PAVTASNATTESP- 149
Query: 181 PPVVASTPPSAPPSNVVPALPPAKTNASSHPQLKCPMAGTFYRCP---GPGEPPFVKVGD 237
+ SA +++ + P + L+ P G F R G P K D
Sbjct: 150 -----ESNGSASSTSLAISKPASSAADQGLMILQSPKVGFFRRSKTIKGKRLPSSCKEKD 204
Query: 238 QVQKGQVVCIIEAMKLMNEIEADRSGTIVEVLVEDGKPV 276
QV++GQ++C IE + IE+D +G +V++L EDG+PV
Sbjct: 205 QVKEGQILCYIEQLGGQFPIESDVTGEVVKILREDGEPV 243
>At3g56130 unknown protein
Length = 281
Score = 54.7 bits (130), Expect = 6e-08
Identities = 45/179 (25%), Positives = 81/179 (45%), Gaps = 16/179 (8%)
Query: 117 TFMDQVSELV-KLVDSRDIMELELKQAGYELMIRKK--EALQPPPVSQ-QPF---PYPAY 169
TF +V LV ++ D ++ L+LK +E+ +++K A P PV+ P P P+
Sbjct: 87 TFPKEVEALVHEMCDETEVAVLQLKVGDFEMNLKRKIGAATNPIPVADISPTVAPPIPSE 146
Query: 170 PPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAK------TNASSHPQLKCPMAGTFYR 223
P + A P P + P PAK + ++++ + P G F R
Sbjct: 147 PMNKSASSAPSPSQAKPSSEKVSPFKNTSYGKPAKLAALEASGSTNYVLVTSPAVGKFQR 206
Query: 224 C---PGPGEPPFVKVGDQVQKGQVVCIIEAMKLMNEIEADRSGTIVEVLVEDGKPVSVG 279
G + P K GD +++GQV+ + + + +D +G ++++L +DG V G
Sbjct: 207 SRTVKGKKQSPSCKEGDAIKEGQVIGYLHQLGTELPVTSDVAGEVLKLLSDDGDSVGYG 265
>At4g13340 extensin-like protein
Length = 760
Score = 53.1 bits (126), Expect = 2e-07
Identities = 30/76 (39%), Positives = 32/76 (41%), Gaps = 1/76 (1%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLKC 215
PPP P P P Y P +P PPPPPV + PP PP PP SS P
Sbjct: 467 PPPPPPPPPPPPVYSPPPPSPPPPPPPVYSPPPPPPPPPPPPVYSPPPPPVYSSPPPPPS 526
Query: 216 PMAGTFYRCPGPGEPP 231
P Y C P PP
Sbjct: 527 PAPTPVY-CTRPPPPP 541
Score = 50.8 bits (120), Expect = 9e-07
Identities = 29/77 (37%), Positives = 31/77 (39%), Gaps = 1/77 (1%)
Query: 156 PPPVSQQPFPYPAY-PPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLK 214
PPP P P P Y PP P PPPPPV + PPS PP PP P +
Sbjct: 451 PPPPPPPPPPPPVYSPPPPPPPPPPPPPVYSPPPPSPPPPPPPVYSPPPPPPPPPPPPVY 510
Query: 215 CPMAGTFYRCPGPGEPP 231
P Y P P P
Sbjct: 511 SPPPPPVYSSPPPPPSP 527
Score = 50.4 bits (119), Expect = 1e-06
Identities = 32/79 (40%), Positives = 32/79 (40%), Gaps = 9/79 (11%)
Query: 156 PPPVSQQPFPYPAYPPSYQA---PLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQ 212
PPPV P P P PP Y P PPPPP V S PP PP PP S P
Sbjct: 432 PPPVYSPPPPPPPPPPVYSPPPPPPPPPPPPVYSPPPPPPPP------PPPPPVYSPPPP 485
Query: 213 LKCPMAGTFYRCPGPGEPP 231
P Y P P PP
Sbjct: 486 SPPPPPPPVYSPPPPPPPP 504
Score = 46.2 bits (108), Expect = 2e-05
Identities = 27/76 (35%), Positives = 30/76 (38%), Gaps = 6/76 (7%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLKC 215
PP ++ P P P P P PPPPP V S PP PP PP + P
Sbjct: 420 PPTLTSPPPPSPPPPVYSPPPPPPPPPPVYSPPPPPPPP------PPPPVYSPPPPPPPP 473
Query: 216 PMAGTFYRCPGPGEPP 231
P Y P P PP
Sbjct: 474 PPPPPVYSPPPPSPPP 489
Score = 43.1 bits (100), Expect = 2e-04
Identities = 28/78 (35%), Positives = 32/78 (40%), Gaps = 9/78 (11%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPL---PPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQ 212
P PVS P P P Y P P PPPPP + +PP PP + PP SS P
Sbjct: 595 PTPVSSPP-PTPVYSPPPPPPCIEPPPPPPCIEYSPPPPPPVVHYSSPPPPPVYYSSPPP 653
Query: 213 LKCPMAGTFYRCPGPGEP 230
+Y P P P
Sbjct: 654 -----PPVYYSSPPPPPP 666
Score = 41.6 bits (96), Expect = 5e-04
Identities = 31/88 (35%), Positives = 36/88 (40%), Gaps = 14/88 (15%)
Query: 156 PPPV-SQQPFPYPAYPPSYQAPLPPPPPVVASTPP---SAPPSNVVPALPPAKTNASSHP 211
PPPV S P P P PP + PPPPPV +S PP AP PP ++ P
Sbjct: 491 PPPVYSPPPPPPPPPPPPVYS--PPPPPVYSSPPPPPSPAPTPVYCTRPPPPPPHSPPPP 548
Query: 212 QLKCPMAGTFY--------RCPGPGEPP 231
Q P +Y P P PP
Sbjct: 549 QFSPPPPEPYYYSSPPPPHSSPPPHSPP 576
Score = 41.2 bits (95), Expect = 7e-04
Identities = 30/83 (36%), Positives = 33/83 (39%), Gaps = 14/83 (16%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSA-------PPSNVVPALPPAKTNAS 208
PPP S P P+ PP Y PPPPP S+PP PP + P PP S
Sbjct: 570 PPPHSPPP-PHSPPPPIYPYLSPPPPPTPVSSPPPTPVYSPPPPPPCIEPPPPPPCIEYS 628
Query: 209 SHPQLKCPMAGTFYRCPGPGEPP 231
P P Y P P PP
Sbjct: 629 PPP----PPPVVHYSSPPP--PP 645
Score = 41.2 bits (95), Expect = 7e-04
Identities = 22/47 (46%), Positives = 25/47 (52%), Gaps = 3/47 (6%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPP 202
PPPV P P PP Y + PPPPPV S+PP PP + PP
Sbjct: 632 PPPVVHYSSPPP--PPVYYSS-PPPPPVYYSSPPPPPPVHYSSPPPP 675
Score = 41.2 bits (95), Expect = 7e-04
Identities = 29/82 (35%), Positives = 33/82 (39%), Gaps = 11/82 (13%)
Query: 156 PPPVSQQPFPYPAY----PPSYQAPLPPPPPVVASTPPSAPPSNVVPAL--PPAKTNASS 209
PPP P P P Y PP + +P P PP PP +PP + P L PP T SS
Sbjct: 546 PPPQFSPPPPEPYYYSSPPPPHSSPPPHSPP-----PPHSPPPPIYPYLSPPPPPTPVSS 600
Query: 210 HPQLKCPMAGTFYRCPGPGEPP 231
P C P PP
Sbjct: 601 PPPTPVYSPPPPPPCIEPPPPP 622
Score = 39.3 bits (90), Expect = 0.003
Identities = 25/56 (44%), Positives = 30/56 (52%), Gaps = 5/56 (8%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHP 211
PPPV P P P Y +P PPPPPV S+PP PP + PP+ + SS P
Sbjct: 643 PPPVYYSSPPPP--PVYYSSP-PPPPPVHYSSPP--PPEVHYHSPPPSPVHYSSPP 693
Score = 38.9 bits (89), Expect = 0.003
Identities = 29/75 (38%), Positives = 33/75 (43%), Gaps = 11/75 (14%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVV-ASTPPSAPPSNVVPALPPAKTNASSHPQLK 214
PPP + P P PP + PPPPPVV S+PP PP + PP SS P
Sbjct: 612 PPPCIEPPPP----PPCIEYSPPPPPPVVHYSSPP--PPPVYYSSPPPPPVYYSSPP--- 662
Query: 215 CPMAGTFYRCPGPGE 229
P Y P P E
Sbjct: 663 -PPPPVHYSSPPPPE 676
Score = 38.1 bits (87), Expect = 0.006
Identities = 28/77 (36%), Positives = 34/77 (43%), Gaps = 7/77 (9%)
Query: 156 PPPV--SQQPFPYPAYPPSYQAPLPPPPPVVASTPP---SAPPSNVVPALPPAKTNASSH 210
P PV ++ P P P PP Q PPP P S+PP S+PP + P PP +
Sbjct: 529 PTPVYCTRPPPPPPHSPPPPQFSPPPPEPYYYSSPPPPHSSPPPHSPP--PPHSPPPPIY 586
Query: 211 PQLKCPMAGTFYRCPGP 227
P L P T P P
Sbjct: 587 PYLSPPPPPTPVSSPPP 603
Score = 37.7 bits (86), Expect = 0.008
Identities = 28/87 (32%), Positives = 34/87 (38%), Gaps = 12/87 (13%)
Query: 153 ALQPPPVSQQPFPYPAYPPSYQAPLPP------PPPVVASTPPSAPPS--NVVPALPPAK 204
++ P P P P PPS +P PP PP + + PPS PP + P PP
Sbjct: 391 SVSPRPPVVTPLP----PPSLPSPPPPAPIFSTPPTLTSPPPPSPPPPVYSPPPPPPPPP 446
Query: 205 TNASSHPQLKCPMAGTFYRCPGPGEPP 231
S P P Y P P PP
Sbjct: 447 PVYSPPPPPPPPPPPPVYSPPPPPPPP 473
Score = 37.0 bits (84), Expect = 0.013
Identities = 35/100 (35%), Positives = 42/100 (42%), Gaps = 27/100 (27%)
Query: 156 PPPV--SQQPFP---YPAYPPS---YQAP----------LPPPPPVVASTPP-----SAP 192
PPPV S P P Y + PPS Y +P PPP PVV +PP +P
Sbjct: 664 PPPVHYSSPPPPEVHYHSPPPSPVHYSSPPPPPSAPCEESPPPAPVVHHSPPPPMVHHSP 723
Query: 193 PSNVVPALPPAKTNASSHPQLKCPMAGTFYRCPGPGEPPF 232
P V+ PP + P P+ G Y P P PPF
Sbjct: 724 PPPVIHQSPPPPSPEYEGP--LPPVIGVSYASPPP--PPF 759
Score = 35.8 bits (81), Expect = 0.029
Identities = 27/87 (31%), Positives = 35/87 (40%), Gaps = 10/87 (11%)
Query: 156 PPPV--SQQPFPYPAY-----PPSYQAPLPPPPPVVASTPPSAP--PSNVVPALPPAKTN 206
PPPV S P P P + PP PPP PV S+PP P P P P +
Sbjct: 653 PPPVYYSSPPPPPPVHYSSPPPPEVHYHSPPPSPVHYSSPPPPPSAPCEESPPPAPVVHH 712
Query: 207 ASSHPQL-KCPMAGTFYRCPGPGEPPF 232
+ P + P ++ P P P +
Sbjct: 713 SPPPPMVHHSPPPPVIHQSPPPPSPEY 739
>At4g18560 pherophorin - like protein
Length = 637
Score = 52.8 bits (125), Expect = 2e-07
Identities = 52/204 (25%), Positives = 76/204 (36%), Gaps = 21/204 (10%)
Query: 20 LKSQTSSRNVQLGFMKSQSF-GSLSCDLVSNGIQCLERKQFSVWRSQALPNEVATIENSS 78
L+ SS + SQ F G + SN I+ L+R V + LP + EN++
Sbjct: 163 LRKLVSSESDDHALSVSQRFQGLMDVSAKSNLIRSLKR----VGSLRNLPEPITNQENTN 218
Query: 79 NSVPVLINEPNGALPKEKDNHNGKPPGPSASTDASSVSTFMDQVSELVKLVDSRDIMELE 138
S+ + K++ + T++SS+ST +V + K R I
Sbjct: 219 KSISSSGDADGDIYRKDEIESYSRSSNSEELTESSSLSTVRSRVPRVPKPPPKRSI---- 274
Query: 139 LKQAGYELMIRKKEALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVP 198
L + PPP P P P PP PPPP V + PP PP
Sbjct: 275 ------SLGDSTENRADPPPQKSIPPPPPPPPPPLLQQPPPPPSVSKAPPPPPPPP---- 324
Query: 199 ALPPAKTNASSHPQLKCPMAGTFY 222
PP + +S + P FY
Sbjct: 325 --PPKSLSIASAKVRRVPEVVEFY 346
Score = 28.5 bits (62), Expect = 4.7
Identities = 13/29 (44%), Positives = 17/29 (57%), Gaps = 1/29 (3%)
Query: 176 PLPPPPPVVASTPPSAPPSNVVPALPPAK 204
PLPPPPP PPS+ + P + P+K
Sbjct: 29 PLPPPPPP-PLKPPSSGSATTKPPINPSK 56
>At1g68690 protein kinase, putative
Length = 708
Score = 51.2 bits (121), Expect = 7e-07
Identities = 32/80 (40%), Positives = 36/80 (45%), Gaps = 2/80 (2%)
Query: 154 LQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSA-PPSNVVPALPPA-KTNASSHP 211
L PP S P P P P + PPP PV+ S PPSA PP +VP LP + AS P
Sbjct: 72 LPKPPESSSPPPQPVIPSPPPSTSPPPQPVIPSPPPSASPPPALVPPLPSSPPPPASVPP 131
Query: 212 QLKCPMAGTFYRCPGPGEPP 231
P R P P P
Sbjct: 132 PRPSPSPPILVRSPPPSVRP 151
Score = 47.0 bits (110), Expect = 1e-05
Identities = 31/84 (36%), Positives = 39/84 (45%), Gaps = 20/84 (23%)
Query: 156 PPPVSQQPFPY------PAYPPSYQAPLPPPPPVVASTPPSA--PPSNVVPALPPAKTNA 207
PPPV+ P P P P ++ PPP PV+ S PPS PP V+P+ PP +A
Sbjct: 53 PPPVTTSPPPVANGAPPPPLPKPPESSSPPPQPVIPSPPPSTSPPPQPVIPSPPP---SA 109
Query: 208 SSHPQLKCPMAGTFYRCPGPGEPP 231
S P L P+ P PP
Sbjct: 110 SPPPALVPPL---------PSSPP 124
Score = 44.3 bits (103), Expect = 8e-05
Identities = 24/52 (46%), Positives = 26/52 (49%), Gaps = 1/52 (1%)
Query: 157 PPVSQQPFPYPAYPPSYQAPLPP-PPPVVASTPPSAPPSNVVPALPPAKTNA 207
PPVS P PP A P PPPV + PPSAPP N P PP T +
Sbjct: 8 PPVSNSPPVTSPPPPLNNATSPATPPPVTSPLPPSAPPPNRAPPPPPPVTTS 59
Score = 43.5 bits (101), Expect = 1e-04
Identities = 29/77 (37%), Positives = 36/77 (46%), Gaps = 19/77 (24%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPP------PVVASTPP----------SAPPSNVVPA 199
PPPV+ P P P+ PP +AP PPPP PV PP S PP V+P+
Sbjct: 32 PPPVTS-PLP-PSAPPPNRAPPPPPPVTTSPPPVANGAPPPPLPKPPESSSPPPQPVIPS 89
Query: 200 LPPAKTNASSHPQLKCP 216
PP T+ P + P
Sbjct: 90 -PPPSTSPPPQPVIPSP 105
Score = 42.0 bits (97), Expect = 4e-04
Identities = 24/67 (35%), Positives = 32/67 (46%), Gaps = 6/67 (8%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPP-----PPPVVASTPPSAPPSNVVP-ALPPAKTNASS 209
P P S +P P PP P PP PP S+PP +PP +VP + P++ N +
Sbjct: 202 PSPPSDRPSQSPPPPPEDTKPQPPRRSPNSPPPTFSSPPRSPPEILVPGSNNPSQNNPTL 261
Query: 210 HPQLKCP 216
P L P
Sbjct: 262 RPPLDAP 268
Score = 41.6 bits (96), Expect = 5e-04
Identities = 25/65 (38%), Positives = 28/65 (42%), Gaps = 8/65 (12%)
Query: 157 PPVSQQPFPY-----PAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHP 211
PPV+ P P PA PP +PLPP P PP PP V PP N + P
Sbjct: 14 PPVTSPPPPLNNATSPATPPPVTSPLPPSAPPPNRAPPPPPP---VTTSPPPVANGAPPP 70
Query: 212 QLKCP 216
L P
Sbjct: 71 PLPKP 75
Score = 40.4 bits (93), Expect = 0.001
Identities = 24/80 (30%), Positives = 33/80 (41%), Gaps = 4/80 (5%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLKC 215
P P S++P P PPS + PPPP +PPS P + PP + P +
Sbjct: 173 PSPPSERPTQSPPSPPSERPTQSPPPP----SPPSPPSDRPSQSPPPPPEDTKPQPPRRS 228
Query: 216 PMAGTFYRCPGPGEPPFVKV 235
P + P PP + V
Sbjct: 229 PNSPPPTFSSPPRSPPEILV 248
Score = 40.0 bits (92), Expect = 0.002
Identities = 26/78 (33%), Positives = 35/78 (44%), Gaps = 2/78 (2%)
Query: 154 LQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQL 213
++ PP S +P P PPS + PPPP S PPS P+ P+ PP++ S P
Sbjct: 142 VRSPPPSVRPIQSPPPPPSDRPTQSPPPPSPPS-PPSERPTQSPPS-PPSERPTQSPPPP 199
Query: 214 KCPMAGTFYRCPGPGEPP 231
P + P PP
Sbjct: 200 SPPSPPSDRPSQSPPPPP 217
Score = 40.0 bits (92), Expect = 0.002
Identities = 28/79 (35%), Positives = 35/79 (43%), Gaps = 6/79 (7%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLP--PPPPVVASTP-PSAPPSNVVPALPPAKTNASSHPQ 212
P PV P P + PP+ PLP PPPP P PS P +V + PP+ S P
Sbjct: 98 PQPVIPSPPPSASPPPALVPPLPSSPPPPASVPPPRPSPSPPILVRSPPPSVRPIQSPPP 157
Query: 213 LKCPMAGTFYRCPGPGEPP 231
P + + P P PP
Sbjct: 158 ---PPSDRPTQSPPPPSPP 173
Score = 37.0 bits (84), Expect = 0.013
Identities = 25/84 (29%), Positives = 35/84 (40%), Gaps = 12/84 (14%)
Query: 156 PPPVSQQPFPYPA--------YPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNA 207
PPP + P P P+ PP P+ PPP + P +PP P+ PP++
Sbjct: 123 PPPPASVPPPRPSPSPPILVRSPPPSVRPIQSPPPPPSDRPTQSPPPPSPPS-PPSERPT 181
Query: 208 SSHPQLKCPMAGTFYRCPGPGEPP 231
S P P + + P P PP
Sbjct: 182 QSPPS---PPSERPTQSPPPPSPP 202
Score = 36.2 bits (82), Expect = 0.022
Identities = 24/78 (30%), Positives = 30/78 (37%), Gaps = 2/78 (2%)
Query: 156 PPPVSQQPFPYPAYPP-SYQAPLP-PPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQL 213
PPP P P PP S P P P PP++ +PP + P PP+ S P
Sbjct: 111 PPPALVPPLPSSPPPPASVPPPRPSPSPPILVRSPPPSVRPIQSPPPPPSDRPTQSPPPP 170
Query: 214 KCPMAGTFYRCPGPGEPP 231
P + P PP
Sbjct: 171 SPPSPPSERPTQSPPSPP 188
>At4g22470 extensin - like protein
Length = 375
Score = 50.4 bits (119), Expect = 1e-06
Identities = 28/72 (38%), Positives = 35/72 (47%), Gaps = 11/72 (15%)
Query: 156 PPPVSQQPFPYPAYPPSYQA-------PLPPP----PPVVASTPPSAPPSNVVPALPPAK 204
P P QP P P PP++Q P PPP PP VA+TPP+ PP + P L P +
Sbjct: 42 PGPPPPQPDPQPPTPPTFQPAPPANDQPPPPPQSTSPPPVATTPPALPPKPLPPPLSPPQ 101
Query: 205 TNASSHPQLKCP 216
T P + P
Sbjct: 102 TTPPPPPAITPP 113
Score = 50.1 bits (118), Expect = 1e-06
Identities = 25/65 (38%), Positives = 34/65 (51%), Gaps = 2/65 (3%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPP--PPPVVASTPPSAPPSNVVPALPPAKTNASSHPQL 213
PPP + P P PA P P P PPP +A+TPP+ PP + P L P +T P +
Sbjct: 105 PPPPAITPPPPPAITPPLSPPPPAITPPPPLATTPPALPPKPLPPPLSPPQTTPPPPPAI 164
Query: 214 KCPMA 218
P++
Sbjct: 165 TPPLS 169
Score = 43.5 bits (101), Expect = 1e-04
Identities = 25/68 (36%), Positives = 30/68 (43%), Gaps = 5/68 (7%)
Query: 153 ALQPPPVSQQPFPYPAYPPSYQAPLPP----PPPVVASTPPSAPPSNVVPALPPAKTNAS 208
A PP + +P P P PP P PP PPP A TPP +PP + PP T
Sbjct: 82 ATTPPALPPKPLPPPLSPPQTTPPPPPAITPPPPP-AITPPLSPPPPAITPPPPLATTPP 140
Query: 209 SHPQLKCP 216
+ P P
Sbjct: 141 ALPPKPLP 148
Score = 43.1 bits (100), Expect = 2e-04
Identities = 26/60 (43%), Positives = 31/60 (51%), Gaps = 8/60 (13%)
Query: 152 EALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHP 211
++ PPPV+ P PA PP PLPPP +TPP PP + P PPA T S P
Sbjct: 74 QSTSPPPVATTP---PALPPK---PLPPPLSPPQTTPP--PPPAITPPPPPAITPPLSPP 125
Score = 37.4 bits (85), Expect = 0.010
Identities = 18/41 (43%), Positives = 22/41 (52%), Gaps = 5/41 (12%)
Query: 153 ALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPP 193
A PP + +P P P PP PPPPP + TPP +PP
Sbjct: 136 ATTPPALPPKPLPPPLSPPQ---TTPPPPPAI--TPPLSPP 171
Score = 37.0 bits (84), Expect = 0.013
Identities = 21/47 (44%), Positives = 25/47 (52%), Gaps = 8/47 (17%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPP 202
PPP++ P PA PP PLPPP +TPP PP + P L P
Sbjct: 132 PPPLATTP---PALPPK---PLPPPLSPPQTTPP--PPPAITPPLSP 170
Score = 33.1 bits (74), Expect = 0.19
Identities = 16/37 (43%), Positives = 18/37 (48%), Gaps = 2/37 (5%)
Query: 157 PPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPP 193
P +S P P P PPPPP A TPP +PP
Sbjct: 258 PTISPPPLPPQTLKPPPPQTTPPPPP--AITPPLSPP 292
Score = 33.1 bits (74), Expect = 0.19
Identities = 26/72 (36%), Positives = 28/72 (38%), Gaps = 5/72 (6%)
Query: 163 PFPYPAYPPSYQAPLPPPPPVVASTPPS---APPSNVVPALPPAKTNASSHPQLKCPMAG 219
P P P P PP P TPP+ APP+N P PP T S P P A
Sbjct: 31 PPPPPCICICNPGPPPPQPDPQPPTPPTFQPAPPANDQPPPPPQST--SPPPVATTPPAL 88
Query: 220 TFYRCPGPGEPP 231
P P PP
Sbjct: 89 PPKPLPPPLSPP 100
Score = 32.0 bits (71), Expect = 0.42
Identities = 23/64 (35%), Positives = 27/64 (41%), Gaps = 14/64 (21%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLKC 215
PPP P P PP+ PP P+ PP +PP P PPA T P L
Sbjct: 125 PPPAITPPPPLATTPPAL-----PPKPLP---PPLSPPQTTPPP-PPAIT-----PPLSP 170
Query: 216 PMAG 219
P+ G
Sbjct: 171 PLVG 174
Score = 29.6 bits (65), Expect = 2.1
Identities = 17/40 (42%), Positives = 20/40 (49%), Gaps = 4/40 (10%)
Query: 163 PFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPP 202
P P + PP L PPPP +TPP PP + P L P
Sbjct: 256 PSPTISPPPLPPQTLKPPPP--QTTPP--PPPAITPPLSP 291
>At3g24480 disease resistance protein, putative
Length = 494
Score = 50.1 bits (118), Expect = 1e-06
Identities = 35/89 (39%), Positives = 39/89 (43%), Gaps = 13/89 (14%)
Query: 153 ALQPPPVSQQPFPYPAYPPSYQAPL--PPPPPVVASTPPSAPPSNVVPALPPAKTNASSH 210
AL PPP P P P Y P P+ PPP P V S PP PPS + PP + SS
Sbjct: 409 ALPPPPPPSPPLPPPVYSPPPSPPVFSPPPSPPVYSPPP--PPSIHYSSPPPPPVHHSSP 466
Query: 211 PQLK-------CPMAGTFYRCPGPGEPPF 232
P P+ G Y P P PPF
Sbjct: 467 PPPSPEFEGPLPPVIGVSYASPPP--PPF 493
Score = 34.7 bits (78), Expect = 0.065
Identities = 28/81 (34%), Positives = 32/81 (38%), Gaps = 17/81 (20%)
Query: 157 PPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPP-----SAPPSNVVPALPPAKTNASSHP 211
PP+ P P P P PP PP V S PP S PPS V + PP + S P
Sbjct: 405 PPIVALPPPPP--------PSPPLPPPVYSPPPSPPVFSPPPSPPVYSPPPPPSIHYSSP 456
Query: 212 QLKCPMAGTFYRCPGPGEPPF 232
P + P P P F
Sbjct: 457 ----PPPPVHHSSPPPPSPEF 473
>At3g24400 protein kinase, putative
Length = 694
Score = 50.1 bits (118), Expect = 1e-06
Identities = 36/101 (35%), Positives = 45/101 (43%), Gaps = 12/101 (11%)
Query: 156 PPPVSQQPFP--YPAYPPSYQAPL-----PPPPPVVASTPPSAPPSNVVPALPPAKTNAS 208
PPP P P P+ PPS +PL PPP P STPP +PP P PP + +
Sbjct: 134 PPPSPSIPSPPLTPSPPPS--SPLRPSSPPPPSPATPSTPPRSPPPPSTPT-PPPRVGSL 190
Query: 209 SHPQLKCPMAGTFYRCPG--PGEPPFVKVGDQVQKGQVVCI 247
S P P G P PG P + ++ KG +V I
Sbjct: 191 SPPPPASPSGGRSPSTPSTTPGSSPPAQSSKELSKGAMVGI 231
Score = 45.4 bits (106), Expect = 4e-05
Identities = 24/53 (45%), Positives = 28/53 (52%), Gaps = 6/53 (11%)
Query: 156 PPP---VSQQPFPYPAYPPSYQAPLPPPPPVVA---STPPSAPPSNVVPALPP 202
PPP S P P P PP P+ PPPP A + PP PP+ + PALPP
Sbjct: 5 PPPGGTPSPPPQPLPIPPPPQPLPVTPPPPPTALPPALPPPPPPTALPPALPP 57
Score = 44.7 bits (104), Expect = 6e-05
Identities = 23/55 (41%), Positives = 27/55 (48%), Gaps = 6/55 (10%)
Query: 155 QPPPVSQQPFPYPAYPP----SYQAPLPPPPPVVASTP--PSAPPSNVVPALPPA 203
QP P+ P P P PP + LPPPPP A P P PP VP +PP+
Sbjct: 16 QPLPIPPPPQPLPVTPPPPPTALPPALPPPPPPTALPPALPPPPPPTTVPPIPPS 70
Score = 43.5 bits (101), Expect = 1e-04
Identities = 28/89 (31%), Positives = 35/89 (38%), Gaps = 14/89 (15%)
Query: 156 PPPVSQQPFP------YPAYPPSYQAPLPP----PPPVVASTPPSAP----PSNVVPALP 201
PPP++ P P P PS P PP PPP + +PP P PS P+ P
Sbjct: 75 PPPLTPSPLPPSPTTPSPPLTPSPTTPSPPLTPSPPPAITPSPPLTPSPLPPSPTTPSPP 134
Query: 202 PAKTNASSHPQLKCPMAGTFYRCPGPGEP 230
P + S P P + R P P
Sbjct: 135 PPSPSIPSPPLTPSPPPSSPLRPSSPPPP 163
Score = 40.8 bits (94), Expect = 0.001
Identities = 20/47 (42%), Positives = 25/47 (52%), Gaps = 1/47 (2%)
Query: 170 PPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLKCP 216
PP P+PPPP + TPP PP+ + PALPP + P L P
Sbjct: 13 PPPQPLPIPPPPQPLPVTPPP-PPTALPPALPPPPPPTALPPALPPP 58
Score = 40.4 bits (93), Expect = 0.001
Identities = 24/59 (40%), Positives = 27/59 (45%), Gaps = 6/59 (10%)
Query: 156 PPPVSQQPFPYPAYPPSYQAP-LPPPPPVVASTP-----PSAPPSNVVPALPPAKTNAS 208
PPP + P P PP+ P LPPPPP P PS PP LPP+ T S
Sbjct: 33 PPPTALPPALPPPPPPTALPPALPPPPPPTTVPPIPPSTPSPPPPLTPSPLPPSPTTPS 91
Score = 38.5 bits (88), Expect = 0.005
Identities = 21/60 (35%), Positives = 27/60 (45%), Gaps = 2/60 (3%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPP--PPPVVASTPPSAPPSNVVPALPPAKTNASSHPQL 213
PPP + P P PP+ P+PP P P TP PPS P+ P + + P L
Sbjct: 46 PPPTALPPALPPPPPPTTVPPIPPSTPSPPPPLTPSPLPPSPTTPSPPLTPSPTTPSPPL 105
Score = 35.8 bits (81), Expect = 0.029
Identities = 24/64 (37%), Positives = 28/64 (43%), Gaps = 14/64 (21%)
Query: 153 ALQPPPVSQQPFPYPAYPPSYQAPLPP-------PPPVVASTP----PSAPPSNVVPALP 201
AL PPP P P PPS +P PP P P S P P+ P + P+ P
Sbjct: 54 ALPPPP---PPTTVPPIPPSTPSPPPPLTPSPLPPSPTTPSPPLTPSPTTPSPPLTPSPP 110
Query: 202 PAKT 205
PA T
Sbjct: 111 PAIT 114
Score = 33.9 bits (76), Expect = 0.11
Identities = 27/80 (33%), Positives = 33/80 (40%), Gaps = 6/80 (7%)
Query: 157 PPVSQQPFP-YPAYPPSYQAPLPPPPPVVASTP---PSAPPSNVVPALPPAKTNASSHPQ 212
PP++ P P PP +PLPP P S P PS P + P+ PP+ S P
Sbjct: 103 PPLTPSPPPAITPSPPLTPSPLPPSP-TTPSPPPPSPSIPSPPLTPSPPPSSPLRPSSPP 161
Query: 213 LKCPMA-GTFYRCPGPGEPP 231
P T R P P P
Sbjct: 162 PPSPATPSTPPRSPPPPSTP 181
>At5g19810 proline-rich protein
Length = 249
Score = 49.7 bits (117), Expect = 2e-06
Identities = 31/75 (41%), Positives = 35/75 (46%), Gaps = 5/75 (6%)
Query: 156 PPPVSQQPFPYPAY---PPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQ 212
PPPV+ P P P PP PPPPPV+ S PP PP N+ P PP + P
Sbjct: 100 PPPVNLSPPPPPVNLSPPPPPVLLSPPPPPVLLSPPP--PPVNLSPPPPPVLLSPPPPPV 157
Query: 213 LKCPMAGTFYRCPGP 227
L P T R P P
Sbjct: 158 LFSPPPPTVTRPPPP 172
Score = 45.8 bits (107), Expect = 3e-05
Identities = 32/83 (38%), Positives = 38/83 (45%), Gaps = 9/83 (10%)
Query: 156 PPPVSQQPFPYPAY--PPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQL 213
PPPV P P P PP L PPPP V +PP PP+ P PP T + P+
Sbjct: 127 PPPVLLSPPPPPVNLSPPPPPVLLSPPPPPVLFSPP--PPTVTRPPPPPTITRSPPPPR- 183
Query: 214 KCPMAGTFYRCPGPGEPPFVKVG 236
P A +Y+ P PP K G
Sbjct: 184 --PQAAAYYKKTPP--PPPYKYG 202
Score = 45.4 bits (106), Expect = 4e-05
Identities = 30/78 (38%), Positives = 34/78 (43%), Gaps = 5/78 (6%)
Query: 156 PPPVSQQPFPYPAY--PPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQL 213
PPPV+ P P P PP L PPPP V +PP PP N+ P PP + P L
Sbjct: 64 PPPVNLSPPPPPVNLSPPPPPVNLSPPPPPVLLSPP-PPPVNLSPPPPPVNLSPPPPPVL 122
Query: 214 KCPMAGTFYRCPGPGEPP 231
P P P PP
Sbjct: 123 LSPPPPPVLLSPPP--PP 138
Score = 43.9 bits (102), Expect = 1e-04
Identities = 29/78 (37%), Positives = 33/78 (42%), Gaps = 5/78 (6%)
Query: 156 PPPVSQQPFPYPAY---PPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQ 212
PPPV+ P P P PP PPPPPV S PP PP + P PP +
Sbjct: 109 PPPVNLSPPPPPVLLSPPPPPVLLSPPPPPVNLSPPP--PPVLLSPPPPPVLFSPPPPTV 166
Query: 213 LKCPMAGTFYRCPGPGEP 230
+ P T R P P P
Sbjct: 167 TRPPPPPTITRSPPPPRP 184
Score = 40.4 bits (93), Expect = 0.001
Identities = 26/68 (38%), Positives = 30/68 (43%), Gaps = 10/68 (14%)
Query: 154 LQPPP----VSQQPFPYPAYPPSYQAPLPPPPPVVASTPP------SAPPSNVVPALPPA 203
L PPP +S P P PP L PPPP V +PP S PP V+ + PP
Sbjct: 42 LSPPPPPVNISSPPPPVNLSPPPPPVNLSPPPPPVNLSPPPPPVNLSPPPPPVLLSPPPP 101
Query: 204 KTNASSHP 211
N S P
Sbjct: 102 PVNLSPPP 109
Score = 37.0 bits (84), Expect = 0.013
Identities = 30/92 (32%), Positives = 32/92 (34%), Gaps = 16/92 (17%)
Query: 156 PPPVSQQPFPYPAY---PPSYQAPLPPPPPVVASTPP-----SAPPS--------NVVPA 199
PPPV+ P P P PP PPPP V PP S PP P
Sbjct: 136 PPPVNLSPPPPPVLLSPPPPPVLFSPPPPTVTRPPPPPTITRSPPPPRPQAAAYYKKTPP 195
Query: 200 LPPAKTNASSHPQLKCPMAGTFYRCPGPGEPP 231
PP K P P A Y+ P PP
Sbjct: 196 PPPYKYGRVYPPPPPPPQAARSYKRSPPPPPP 227
Score = 32.7 bits (73), Expect = 0.25
Identities = 22/76 (28%), Positives = 29/76 (37%), Gaps = 8/76 (10%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVAST--PPSAPPSNVVPALPPAKTNASSHPQL 213
PP +++ P P +Y PPPPP PP PP A++ S P
Sbjct: 172 PPTITRSPPPPRPQAAAYYKKTPPPPPYKYGRVYPPPPPPPQA------ARSYKRSPPPP 225
Query: 214 KCPMAGTFYRCPGPGE 229
G Y P PG+
Sbjct: 226 PPSKYGRVYSPPPPGK 241
Score = 29.3 bits (64), Expect = 2.7
Identities = 19/42 (45%), Positives = 22/42 (52%), Gaps = 5/42 (11%)
Query: 171 PSYQAPLPPPPPVV-ASTPPSAPPSNVVPALPPAKTNASSHP 211
P+ L PPPP V S+PP PP N+ P PP N S P
Sbjct: 36 PAPLVDLSPPPPPVNISSPP--PPVNLSP--PPPPVNLSPPP 73
>At5g07750 putative protein
Length = 1289
Score = 49.3 bits (116), Expect = 3e-06
Identities = 31/82 (37%), Positives = 35/82 (41%), Gaps = 6/82 (7%)
Query: 156 PPPVSQQPFPYPAYPPSYQA--PLPPPPPVVASTPPSAPP----SNVVPALPPAKTNASS 209
PPP P P P PPSY + P PPPPP S PP PP + P PP + S
Sbjct: 936 PPPSYGSPPPPPPPPPSYGSPPPPPPPPPSYGSPPPPPPPPPGYGSPPPPPPPPPSYGSP 995
Query: 210 HPQLKCPMAGTFYRCPGPGEPP 231
P P + P P PP
Sbjct: 996 PPPPPPPFSHVSSIPPPPPPPP 1017
Score = 48.5 bits (114), Expect = 4e-06
Identities = 30/84 (35%), Positives = 34/84 (39%), Gaps = 4/84 (4%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVA----STPPSAPPSNVVPALPPAKTNASSHP 211
PPP S PPSY +P PPPPP + PP PPS P PP P
Sbjct: 923 PPPFSNAHSVLSPPPPSYGSPPPPPPPPPSYGSPPPPPPPPPSYGSPPPPPPPPPGYGSP 982
Query: 212 QLKCPMAGTFYRCPGPGEPPFVKV 235
P ++ P P PPF V
Sbjct: 983 PPPPPPPPSYGSPPPPPPPPFSHV 1006
Score = 47.8 bits (112), Expect = 7e-06
Identities = 28/76 (36%), Positives = 30/76 (38%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLKC 215
PPP+ P P P AP PPPPP+ PP PP A PP P
Sbjct: 1090 PPPMRGGAPPPPPPPMRGGAPPPPPPPMHGGAPPPPPPPMRGGAPPPPPPPGGRGPGAPP 1149
Query: 216 PMAGTFYRCPGPGEPP 231
P R PGP PP
Sbjct: 1150 PPPPPGGRAPGPPPPP 1165
Score = 46.2 bits (108), Expect = 2e-05
Identities = 33/85 (38%), Positives = 37/85 (42%), Gaps = 13/85 (15%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPV----VASTPPSAPPSNV-----VPALPPAKTN 206
PPP P P P PPSY +P PPPPP V+S PP PP + P PP
Sbjct: 975 PPPGYGSPPPPPPPPPSYGSP-PPPPPPPFSHVSSIPPPPPPPPMHGGAPPPPPPPPMHG 1033
Query: 207 ASSHPQLKCPMAGTFYRCPGPGEPP 231
+ P PM G P P PP
Sbjct: 1034 GAPPPPPPPPMHG---GAPPPPPPP 1055
Score = 46.2 bits (108), Expect = 2e-05
Identities = 30/84 (35%), Positives = 34/84 (39%), Gaps = 8/84 (9%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPP--SNVVPALPPAKTNASSHPQL 213
PPP+ P P P AP PPPPP+ PP PP P PP + + P
Sbjct: 1066 PPPMFGGAQPPPPPPMRGGAPPPPPPPMRGGAPPPPPPPMRGGAPPPPPPPMHGGAPPPP 1125
Query: 214 KCPMAGTFYRCP------GPGEPP 231
PM G P GPG PP
Sbjct: 1126 PPPMRGGAPPPPPPPGGRGPGAPP 1149
Score = 43.9 bits (102), Expect = 1e-04
Identities = 29/79 (36%), Positives = 33/79 (41%), Gaps = 6/79 (7%)
Query: 156 PPPVSQQPFPYPAYPPSYQ-APLPPPPPVV--ASTPPSAPPSNVVPALPPAKTNASSHPQ 212
PPP+ P P PP + AP PPPPP+ A PP P P PP + P
Sbjct: 1041 PPPMHGGAPPPPPPPPMHGGAPPPPPPPMFGGAQPPPPPPMRGGAPPPPPPPMRGGAPPP 1100
Query: 213 LKCPMAGTFYRCPGPGEPP 231
PM G P P PP
Sbjct: 1101 PPPPMRG---GAPPPPPPP 1116
Score = 43.1 bits (100), Expect = 2e-04
Identities = 27/79 (34%), Positives = 30/79 (37%), Gaps = 6/79 (7%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVV---ASTPPSAPPSNVVPALPPAKTNASSHPQ 212
PPP P P PP + PPPPP + A PP P P PP + P
Sbjct: 1053 PPPPMHGGAPPPPPPPMFGGAQPPPPPPMRGGAPPPPPPPMRGGAPPPPPPPMRGGAPPP 1112
Query: 213 LKCPMAGTFYRCPGPGEPP 231
PM G P P PP
Sbjct: 1113 PPPPMHG---GAPPPPPPP 1128
Score = 42.7 bits (99), Expect = 2e-04
Identities = 31/93 (33%), Positives = 35/93 (37%), Gaps = 17/93 (18%)
Query: 156 PPPVSQQPF-------PYPAYPPSYQA--PLPPPPPVVASTPPSAPP----SNVVPALPP 202
PPP PF P P PP + P PPPPP+ PP PP P PP
Sbjct: 995 PPPPPPPPFSHVSSIPPPPPPPPMHGGAPPPPPPPPMHGGAPPPPPPPPMHGGAPPPPPP 1054
Query: 203 AKTNASSHPQLKCPMAGTFYRCPGP----GEPP 231
+ + P PM G P P G PP
Sbjct: 1055 PPMHGGAPPPPPPPMFGGAQPPPPPPMRGGAPP 1087
Score = 42.7 bits (99), Expect = 2e-04
Identities = 32/86 (37%), Positives = 36/86 (41%), Gaps = 14/86 (16%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVA--------STPPS--APPSNVVPALPPAKT 205
PPP P P P PPSY +P PPPPP PPS +PP P PP +
Sbjct: 949 PPPSYGSPPPPPPPPPSYGSPPPPPPPPPGYGSPPPPPPPPPSYGSPP----PPPPPPFS 1004
Query: 206 NASSHPQLKCPMAGTFYRCPGPGEPP 231
+ SS P P P P PP
Sbjct: 1005 HVSSIPPPPPPPPMHGGAPPPPPPPP 1030
Score = 40.8 bits (94), Expect = 0.001
Identities = 30/79 (37%), Positives = 32/79 (39%), Gaps = 6/79 (7%)
Query: 157 PPVSQQPFPYPAYPPSY-QAPLPPPPPVVASTPPSAPPS---NVVPALPPAKTNASSHPQ 212
PPV P P P P S + L PPPP S PP PP P PP + S P
Sbjct: 912 PPVKTAPPPPPPPPFSNAHSVLSPPPPSYGSPPPPPPPPPSYGSPPPPPPPPPSYGSPPP 971
Query: 213 LKCPMAGTFYRCPGPGEPP 231
P G Y P P PP
Sbjct: 972 PPPPPPG--YGSPPPPPPP 988
Score = 40.4 bits (93), Expect = 0.001
Identities = 28/87 (32%), Positives = 36/87 (41%), Gaps = 5/87 (5%)
Query: 147 MIRKKEALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPS-NVVPALPPAKT 205
++ +A+ PP P P P Y + LPPPPP P S V+P PP
Sbjct: 637 VLHSSQAVASPPPPPPPPPLPTYSHYQTSQLPPPPPPPPPFSSERPNSGTVLPPPPPPPP 696
Query: 206 NASSHPQLKCPMAGTFYRCPGPGEPPF 232
SS + P +GT P P PF
Sbjct: 697 PFSS----ERPNSGTVLPPPPPPPLPF 719
Score = 40.0 bits (92), Expect = 0.002
Identities = 29/84 (34%), Positives = 34/84 (39%), Gaps = 11/84 (13%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVV-----ASTPPSAPPSN---VVPALPPAKTNA 207
PPP P P P P S+ + +PPPPP A PP PP + P PP
Sbjct: 988 PPPSYGSPPPPPPPPFSHVSSIPPPPPPPPMHGGAPPPPPPPPMHGGAPPPPPPPPMHGG 1047
Query: 208 SSHPQLKCPMAGTFYRCPGPGEPP 231
+ P PM G P P PP
Sbjct: 1048 APPPPPPPPMHG---GAPPPPPPP 1068
Score = 37.7 bits (86), Expect = 0.008
Identities = 28/80 (35%), Positives = 31/80 (38%), Gaps = 7/80 (8%)
Query: 156 PPPVSQQPFPYPAYPPSYQ-APLPPPPPV---VASTPPSAPPSNVVPALPPAKTNASSHP 211
PPP+ P P PP + AP PPPPP A PP P PP + P
Sbjct: 1028 PPPMHGGAPPPPPPPPMHGGAPPPPPPPPMHGGAPPPPPPPMFGGAQPPPPPPMRGGAPP 1087
Query: 212 QLKCPMAGTFYRCPGPGEPP 231
PM G P P PP
Sbjct: 1088 PPPPPMRG---GAPPPPPPP 1104
Score = 35.8 bits (81), Expect = 0.029
Identities = 28/93 (30%), Positives = 35/93 (37%), Gaps = 16/93 (17%)
Query: 149 RKKEALQPPPVSQQPFPYPAYPPSYQAPL--------PPPPPVVASTPPSAPP--SNVVP 198
+ + + PPP P PY + PS + P PPV + PP PP SN
Sbjct: 876 KANDYILPPP----PLPYTSIAPSPSVKILPLHGISSAPSPPVKTAPPPPPPPPFSNAHS 931
Query: 199 ALPPAKTNASSHPQLKCPMAGTFYRCPGPGEPP 231
L P + S P P Y P P PP
Sbjct: 932 VLSPPPPSYGSPPPPPPPPPS--YGSPPPPPPP 962
Score = 35.0 bits (79), Expect = 0.050
Identities = 22/76 (28%), Positives = 28/76 (35%), Gaps = 15/76 (19%)
Query: 149 RKKEALQPPPVSQQPFPYPAYPPSYQAPLPPPPP---------------VVASTPPSAPP 193
R E L PPP P P+ + + + LPPPPP ST S PP
Sbjct: 803 RNSETLLPPPPPPPPPPFASVRRNSETLLPPPPPPPPWKSLYASTFETHEACSTSSSPPP 862
Query: 194 SNVVPALPPAKTNASS 209
P P T ++
Sbjct: 863 PPPPPPFSPLNTTKAN 878
Score = 34.3 bits (77), Expect = 0.085
Identities = 27/89 (30%), Positives = 36/89 (40%), Gaps = 7/89 (7%)
Query: 157 PPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVV---PALPPAKT---NASSH 210
PP P P+ + P+ LPPPPP P S V P PP K+ +A +
Sbjct: 689 PPPPPPPPPFSSERPNSGTVLPPPPPPPLPFSSERPNSGTVLPPPPSPPWKSVYASALAI 748
Query: 211 PQL-KCPMAGTFYRCPGPGEPPFVKVGDQ 238
P + A T P P P + VG +
Sbjct: 749 PAICSTSQAPTSSPTPPPPPPAYYSVGQK 777
Score = 33.5 bits (75), Expect = 0.15
Identities = 18/39 (46%), Positives = 20/39 (51%), Gaps = 3/39 (7%)
Query: 156 PPPVSQQP-FPYPAYPPSYQAPLPPPPPVVASTPPSAPP 193
PPP + P P P PP +AP PPPPP PP P
Sbjct: 1138 PPPGGRGPGAPPPPPPPGGRAPGPPPPP--GPRPPGGGP 1174
Score = 32.3 bits (72), Expect = 0.32
Identities = 32/124 (25%), Positives = 44/124 (34%), Gaps = 17/124 (13%)
Query: 153 ALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPP----------SNVVPALPP 202
+L P Q P P P+ + L V + PP PP S + P PP
Sbjct: 614 SLPPASPHQAPPPLPSLTSEAKTVLHSSQAVASPPPPPPPPPLPTYSHYQTSQLPPPPPP 673
Query: 203 AKTNASSHPQLKCPMAGTFYRCPGPGEPPFVKVGDQVQKGQVVCIIEAMKLMNEIEADRS 262
+S P +GT P P PPF ++ G V+ L E S
Sbjct: 674 PPPFSSERPN-----SGTVLPPPPPPPPPF--SSERPNSGTVLPPPPPPPLPFSSERPNS 726
Query: 263 GTIV 266
GT++
Sbjct: 727 GTVL 730
Score = 32.3 bits (72), Expect = 0.32
Identities = 26/102 (25%), Positives = 35/102 (33%), Gaps = 18/102 (17%)
Query: 149 RKKEALQPPPVSQQPFP---------YPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPA 199
R E L PPP P+ + A S P PPPPP + + ++P
Sbjct: 825 RNSETLLPPPPPPPPWKSLYASTFETHEACSTSSSPPPPPPPPPFSPLNTTKANDYILPP 884
Query: 200 LPPAKTNASSHPQLK---------CPMAGTFYRCPGPGEPPF 232
P T+ + P +K P P P PPF
Sbjct: 885 PPLPYTSIAPSPSVKILPLHGISSAPSPPVKTAPPPPPPPPF 926
Score = 32.3 bits (72), Expect = 0.32
Identities = 19/50 (38%), Positives = 22/50 (44%), Gaps = 3/50 (6%)
Query: 156 PPPVSQQPFPYPAYPPSYQ---APLPPPPPVVASTPPSAPPSNVVPALPP 202
PPP + P P PP + AP PPPPP + P PP P P
Sbjct: 1125 PPPPMRGGAPPPPPPPGGRGPGAPPPPPPPGGRAPGPPPPPGPRPPGGGP 1174
Score = 32.0 bits (71), Expect = 0.42
Identities = 26/82 (31%), Positives = 34/82 (40%), Gaps = 7/82 (8%)
Query: 155 QPPPVSQQPFPYPAYPPSYQAPL-PPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQL 213
Q PP P P+ + P+ L PPPPP + V+P PP SS
Sbjct: 666 QLPPPPPPPPPFSSERPNSGTVLPPPPPPPPPFSSERPNSGTVLPPPPPPPLPFSS---- 721
Query: 214 KCPMAGTFYRCPGPGEPPFVKV 235
+ P +GT P P PP+ V
Sbjct: 722 ERPNSGTV--LPPPPSPPWKSV 741
Score = 30.0 bits (66), Expect = 1.6
Identities = 25/89 (28%), Positives = 36/89 (40%), Gaps = 15/89 (16%)
Query: 154 LQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNV----------VPALPPA 203
L PP P P+ + + + LPPPPP PP P ++V P PP
Sbjct: 786 LPSPPPPPPPPPFASVRRNSETLLPPPPP-----PPPPPFASVRRNSETLLPPPPPPPPW 840
Query: 204 KTNASSHPQLKCPMAGTFYRCPGPGEPPF 232
K+ +S + + + P P PPF
Sbjct: 841 KSLYASTFETHEACSTSSSPPPPPPPPPF 869
Score = 29.3 bits (64), Expect = 2.7
Identities = 13/38 (34%), Positives = 18/38 (47%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPP 193
PPP + P P P P PPPPP++ + + P
Sbjct: 1152 PPPGGRAPGPPPPPGPRPPGGGPPPPPMLGARGAAVDP 1189
Score = 28.1 bits (61), Expect = 6.1
Identities = 32/104 (30%), Positives = 34/104 (31%), Gaps = 32/104 (30%)
Query: 154 LQPPP------VSQQPFPYPAYPPSYQAPL-----PPPPPVVASTP-----------PSA 191
L PPP V PA + QAP PPPPP S PS
Sbjct: 730 LPPPPSPPWKSVYASALAIPAICSTSQAPTSSPTPPPPPPAYYSVGQKSSDLQTSQLPSP 789
Query: 192 PPSNVVPALPPAKTNASSHPQLKCPMAGTFYRCPGPGEPPFVKV 235
PP P P A +S L P P P PPF V
Sbjct: 790 PPP--PPPPPFASVRRNSETLLPPP--------PPPPPPPFASV 823
>At3g22800 hypothetical protein
Length = 470
Score = 48.5 bits (114), Expect = 4e-06
Identities = 21/46 (45%), Positives = 25/46 (53%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALP 201
PPP S P+ YP PP Y P PP PP V PP +P + P+ P
Sbjct: 415 PPPPSPPPYVYPPPPPPYVYPPPPSPPYVYPPPPPSPQPYMYPSPP 460
Score = 47.4 bits (111), Expect = 1e-05
Identities = 30/81 (37%), Positives = 32/81 (39%), Gaps = 21/81 (25%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPS---NVVPALPPAKTNASSHPQ 212
PPP P P P PP P PPPPP V +PP PPS V P PP
Sbjct: 381 PPPPPPPPPPPPPPPPPPPPPPPPPPPYVYPSPPPPPPSPPPYVYPPPPPP--------- 431
Query: 213 LKCPMAGTFYRCPGPGEPPFV 233
Y P P PP+V
Sbjct: 432 ---------YVYPPPPSPPYV 443
Score = 43.9 bits (102), Expect = 1e-04
Identities = 23/64 (35%), Positives = 28/64 (42%), Gaps = 6/64 (9%)
Query: 156 PPPVSQQPFPYPAYPPSYQAP----LPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHP 211
PPP P+ YP+ PP +P PPPPP PP +PP P PP +P
Sbjct: 400 PPPPPPPPYVYPSPPPPPPSPPPYVYPPPPPPYVYPPPPSPPYVYPP--PPPSPQPYMYP 457
Query: 212 QLKC 215
C
Sbjct: 458 SPPC 461
>At2g14890 arabinogalactan-protein AGP9
Length = 191
Score = 48.5 bits (114), Expect = 4e-06
Identities = 30/72 (41%), Positives = 34/72 (46%), Gaps = 5/72 (6%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLKC 215
PPPVS P P PP+ P+ PPP VAS PP+ PP A PP AS Q+
Sbjct: 71 PPPVSSPPPASP--PPATPPPVASPPPPVASPPPATPPP---VATPPPAPLASPPAQVPA 125
Query: 216 PMAGTFYRCPGP 227
P T P P
Sbjct: 126 PAPTTKPDSPSP 137
Score = 46.6 bits (109), Expect = 2e-05
Identities = 28/68 (41%), Positives = 32/68 (46%), Gaps = 2/68 (2%)
Query: 153 ALQPPPVSQQPFPYPAYPPSYQAPLP--PPPPVVASTPPSAPPSNVVPALPPAKTNASSH 210
A PPPVS P + PP AP P PPPPV + P S PP+ P P AS
Sbjct: 42 AATPPPVSAPPPVTTSPPPVTTAPPPANPPPPVSSPPPASPPPATPPPVASPPPPVASPP 101
Query: 211 PQLKCPMA 218
P P+A
Sbjct: 102 PATPPPVA 109
Score = 41.6 bits (96), Expect = 5e-04
Identities = 24/76 (31%), Positives = 35/76 (45%), Gaps = 4/76 (5%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPS---APPSNVVPALP-PAKTNASSHP 211
PPPV+ P P + PP+ P+ PPP ++PP+ AP P P P+ +++ P
Sbjct: 87 PPPVASPPPPVASPPPATPPPVATPPPAPLASPPAQVPAPAPTTKPDSPSPSPSSSPPLP 146
Query: 212 QLKCPMAGTFYRCPGP 227
P T P P
Sbjct: 147 SSDAPGPSTDSISPAP 162
Score = 40.0 bits (92), Expect = 0.002
Identities = 28/88 (31%), Positives = 35/88 (38%), Gaps = 10/88 (11%)
Query: 152 EALQPPPVSQQPFPYPAYPPSYQAP----LPPP----PPVVASTPPSAPPSNVVPALPPA 203
+A PP + P P PP P PPP PP V + PP A P V + PPA
Sbjct: 21 QAPTSPPTATPAPPTPTTPPPAATPPPVSAPPPVTTSPPPVTTAPPPANPPPPVSSPPPA 80
Query: 204 KTNASSHPQLKCPMAGTFYRCPGPGEPP 231
++ P + P P P PP
Sbjct: 81 SPPPATPPPVASPPPPV--ASPPPATPP 106
Score = 28.5 bits (62), Expect = 4.7
Identities = 19/60 (31%), Positives = 25/60 (41%), Gaps = 1/60 (1%)
Query: 153 ALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVAS-TPPSAPPSNVVPALPPAKTNASSHP 211
A PP + P P + P AP P P S +P S+PP A P+ + S P
Sbjct: 103 ATPPPVATPPPAPLASPPAQVPAPAPTTKPDSPSPSPSSSPPLPSSDAPGPSTDSISPAP 162
>At5g26080 extensin - like protein
Length = 141
Score = 48.1 bits (113), Expect = 6e-06
Identities = 29/84 (34%), Positives = 39/84 (45%), Gaps = 12/84 (14%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPP--PPPVVASTPPSAPPSNV-----VPALPPAKTNAS 208
PPPV +P +P PP Y P PP PPP+ + PP P + P PP K +
Sbjct: 56 PPPVYSRPVAFPPPPPIYSPPPPPIYPPPIYSPPPPPIYPPPIYSPPPTPISPPPKVH-- 113
Query: 209 SHPQLKCPMAGTFYRCPGP--GEP 230
HP + A + + P P G+P
Sbjct: 114 -HPAPQAQKAFYYRQSPPPPSGQP 136
Score = 42.4 bits (98), Expect = 3e-04
Identities = 19/49 (38%), Positives = 24/49 (48%), Gaps = 2/49 (4%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPL--PPPPPVVASTPPSAPPSNVVPALPP 202
PPP + P P PP Y P+ PPPPP+ + PP P + PP
Sbjct: 43 PPPPYRSPVTIPPPPPVYSRPVAFPPPPPIYSPPPPPIYPPPIYSPPPP 91
Score = 33.9 bits (76), Expect = 0.11
Identities = 22/67 (32%), Positives = 29/67 (42%), Gaps = 4/67 (5%)
Query: 166 YPAYPPSYQAPLP-PPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLKCPMAGTFYRC 224
Y PP Y++P+ PPPP V S P + PP + + PP P + P Y
Sbjct: 40 YSPPPPPYRSPVTIPPPPPVYSRPVAFPPPPPIYSPPPPPIYP---PPIYSPPPPPIYPP 96
Query: 225 PGPGEPP 231
P PP
Sbjct: 97 PIYSPPP 103
>At1g12040 leucine-rich repeat/extensin 1 (LRX1)
Length = 744
Score = 47.8 bits (112), Expect = 7e-06
Identities = 33/87 (37%), Positives = 36/87 (40%), Gaps = 14/87 (16%)
Query: 156 PPPVSQQPFPYPAY-----PPSYQAPLPPPPPVVASTPP------SAPPSNVVPALPPAK 204
PPP P P P Y PP Y PPPPP V S+PP S+PP V + PP
Sbjct: 457 PPPPPPSPSPPPPYVYSSPPPPYVYSSPPPPPYVYSSPPPPPYVYSSPPPPYVYSSPPPP 516
Query: 205 TNASSHPQLKCPMAGTFYRCPGPGEPP 231
SS P P CP PP
Sbjct: 517 YVYSSPPP---PPPSPPPPCPESSPPP 540
Score = 40.4 bits (93), Expect = 0.001
Identities = 29/85 (34%), Positives = 32/85 (37%), Gaps = 10/85 (11%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPP------SAPPSNVVPALPPAKTNASS 209
PPP S+ AYPP PPPP V S+PP S PP V + PP S
Sbjct: 443 PPPYSKMSPSVRAYPPPPPPSPSPPPPYVYSSPPPPYVYSSPPPPPYVYSSPPPPPYVYS 502
Query: 210 HPQ----LKCPMAGTFYRCPGPGEP 230
P P Y P P P
Sbjct: 503 SPPPPYVYSSPPPPYVYSSPPPPPP 527
Score = 38.9 bits (89), Expect = 0.003
Identities = 31/91 (34%), Positives = 37/91 (40%), Gaps = 20/91 (21%)
Query: 156 PPPVSQQ----------PFPYPAYPPSYQAPLPPPPPVVASTPP---SAPPSNVVPALPP 202
PPP S+ P PY PS +A PPPPP + PP S+PP V + PP
Sbjct: 426 PPPSSKMSPSVRAYSPPPPPYSKMSPSVRAYPPPPPPSPSPPPPYVYSSPPPPYVYSSPP 485
Query: 203 AKTNASSHPQLKCPMAGTFYRCPGPGEPPFV 233
S P P Y P PP+V
Sbjct: 486 PPPYVYSSP----PPPPYVYSSP---PPPYV 509
Score = 37.4 bits (85), Expect = 0.010
Identities = 29/93 (31%), Positives = 38/93 (40%), Gaps = 15/93 (16%)
Query: 156 PPPV----SQQPFPYPAYPPSYQAPLPPPPP-------VVASTPPSAPPSNVVPAL---- 200
PPP+ S P P P+ +A PPPPP V A +PP P S + P++
Sbjct: 398 PPPIYVYSSPPPPPSSKMSPTVRAYSPPPPPSSKMSPSVRAYSPPPPPYSKMSPSVRAYP 457
Query: 201 PPAKTNASSHPQLKCPMAGTFYRCPGPGEPPFV 233
PP + S P Y P PP+V
Sbjct: 458 PPPPPSPSPPPPYVYSSPPPPYVYSSPPPPPYV 490
Score = 37.4 bits (85), Expect = 0.010
Identities = 25/79 (31%), Positives = 34/79 (42%), Gaps = 10/79 (12%)
Query: 156 PPPVSQQPFPYPAYPPSY--QAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQL 213
PPP P+ Y + PP Y +P PPPP P S+PP VV P ++ P
Sbjct: 504 PPP----PYVYSSPPPPYVYSSPPPPPPSPPPPCPESSPPPPVVYYAPVTQSPPPPSPVY 559
Query: 214 KCPMAGTFYRCPGPGEPPF 232
P+ + P P P +
Sbjct: 560 YPPVT----QSPPPPSPVY 574
Score = 37.0 bits (84), Expect = 0.013
Identities = 30/95 (31%), Positives = 38/95 (39%), Gaps = 15/95 (15%)
Query: 152 EALQPPPV-------SQQPFPYPA-YPPSYQAPLPPP----PPVVASTPPSAPPSNVVPA 199
E+ PPPV P P P YPP Q+P PP PPV S PP +P
Sbjct: 535 ESSPPPPVVYYAPVTQSPPPPSPVYYPPVTQSPPPPSPVYYPPVTNSPPPPSPVYYPPVT 594
Query: 200 LPPAKTNASSHPQL---KCPMAGTFYRCPGPGEPP 231
P + +PQ+ P + +Y P PP
Sbjct: 595 YSPPPPSPVYYPQVTPSPPPPSPLYYPPVTPSPPP 629
Score = 35.8 bits (81), Expect = 0.029
Identities = 27/79 (34%), Positives = 30/79 (37%), Gaps = 5/79 (6%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPP-PPPVVASTPPSAPP--SNVVPALPPAKTNASSHPQ 212
PPP S P YP PS P P PPV S PP +P V P+ PP
Sbjct: 612 PPPPS--PLYYPPVTPSPPPPSPVYYPPVTPSPPPPSPVYYPPVTPSPPPPSPVYYPSET 669
Query: 213 LKCPMAGTFYRCPGPGEPP 231
P +Y P PP
Sbjct: 670 QSPPPPTEYYYSPSQSPPP 688
Score = 35.0 bits (79), Expect = 0.050
Identities = 21/54 (38%), Positives = 23/54 (41%), Gaps = 7/54 (12%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPP-----PPVVASTPPSAPP--SNVVPALPP 202
PP P P P Y P PPP PPV S PP +P V P+ PP
Sbjct: 591 PPVTYSPPPPSPVYYPQVTPSPPPPSPLYYPPVTPSPPPPSPVYYPPVTPSPPP 644
Score = 34.3 bits (77), Expect = 0.085
Identities = 22/61 (36%), Positives = 26/61 (42%), Gaps = 5/61 (8%)
Query: 156 PPPVSQQPFPYPAYPPS--YQAPL---PPPPPVVASTPPSAPPSNVVPALPPAKTNASSH 210
PPP P P + PP Y AP+ PPPP V P + P P P TN+
Sbjct: 525 PPPSPPPPCPESSPPPPVVYYAPVTQSPPPPSPVYYPPVTQSPPPPSPVYYPPVTNSPPP 584
Query: 211 P 211
P
Sbjct: 585 P 585
Score = 34.3 bits (77), Expect = 0.085
Identities = 20/67 (29%), Positives = 24/67 (34%), Gaps = 6/67 (8%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPS------NVVPALPPAKTNASS 209
PP P P P Y P PPP PV + +PP + + PP K
Sbjct: 636 PPVTPSPPPPSPVYYPPVTPSPPPPSPVYYPSETQSPPPPTEYYYSPSQSPPPTKACKEG 695
Query: 210 HPQLKCP 216
HP P
Sbjct: 696 HPPQATP 702
Score = 33.9 bits (76), Expect = 0.11
Identities = 22/80 (27%), Positives = 33/80 (40%), Gaps = 13/80 (16%)
Query: 165 PYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPAL----PPAKTNASSHPQLKCPMAGT 220
P P + S + PPP V S+PP P S + P + PP ++ P +
Sbjct: 384 PPPTFKMSPEVRTLPPPIYVYSSPPPPPSSKMSPTVRAYSPPPPPSSKMSPSV------- 436
Query: 221 FYRCPGPGEPPFVKVGDQVQ 240
R P PP+ K+ V+
Sbjct: 437 --RAYSPPPPPYSKMSPSVR 454
Score = 33.5 bits (75), Expect = 0.15
Identities = 31/97 (31%), Positives = 35/97 (35%), Gaps = 24/97 (24%)
Query: 156 PPPVSQQPFPYPAYPPSY-QAPLPP----------PPPVVAST---PPSAPPSNVVP--- 198
PP P P P Y PS Q+P PP PPP A PP A PS P
Sbjct: 651 PPVTPSPPPPSPVYYPSETQSPPPPTEYYYSPSQSPPPTKACKEGHPPQATPSYEPPPEY 710
Query: 199 ---ALPPAKTNASSHPQLKCPMAGTFYRCPGPGEPPF 232
+ PP + S P PM Y P P +
Sbjct: 711 SYSSSPPPPSPTSYFP----PMPSVSYDASPPPPPSY 743
Score = 33.5 bits (75), Expect = 0.15
Identities = 20/58 (34%), Positives = 29/58 (49%), Gaps = 6/58 (10%)
Query: 156 PPPVS--QQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHP 211
PPP ++ P P PSY+ PPP +S+PP P++ P +P +AS P
Sbjct: 686 PPPTKACKEGHP-PQATPSYE---PPPEYSYSSSPPPPSPTSYFPPMPSVSYDASPPP 739
Score = 31.6 bits (70), Expect = 0.55
Identities = 24/86 (27%), Positives = 31/86 (35%), Gaps = 17/86 (19%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVV-------ASTPPSAPPSNVVPAL----PPAK 204
PP P PP Y PPPPP A +PP P S + P++ PP
Sbjct: 385 PPTFKMSPEVRTLPPPIYVYSSPPPPPSSKMSPTVRAYSPPPPPSSKMSPSVRAYSPPPP 444
Query: 205 TNASSHPQLKCPMAGTFYRCPGPGEP 230
+ P ++ Y P P P
Sbjct: 445 PYSKMSPSVRA------YPPPPPPSP 464
>At4g33970 extensin-like protein
Length = 699
Score = 47.0 bits (110), Expect = 1e-05
Identities = 26/66 (39%), Positives = 32/66 (48%), Gaps = 5/66 (7%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPP---SAPPSNVVPAL--PPAKTNASSH 210
PPPV P P P + P PPPPP V S PP S PPS P + PP + +
Sbjct: 590 PPPVHSPPPPAPVHSPPPPVHSPPPPPPVYSPPPPVFSPPPSQSPPVVYSPPPRPPKINS 649
Query: 211 PQLKCP 216
P ++ P
Sbjct: 650 PPVQSP 655
Score = 42.0 bits (97), Expect = 4e-04
Identities = 29/86 (33%), Positives = 36/86 (41%), Gaps = 9/86 (10%)
Query: 148 IRKKEALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPP--SAPPSNVVPALPPAKT 205
+ K+ + P PV+ P PP Y P PPPPPV + PP S PP V PP
Sbjct: 512 VTKRRSPPPAPVNSPP------PPVYSPP-PPPPPVHSPPPPVHSPPPPPVYSPPPPPPP 564
Query: 206 NASSHPQLKCPMAGTFYRCPGPGEPP 231
S P + P + P PP
Sbjct: 565 VHSPPPPVFSPPPPVYSPPPPVHSPP 590
Score = 41.2 bits (95), Expect = 7e-04
Identities = 29/79 (36%), Positives = 34/79 (42%), Gaps = 6/79 (7%)
Query: 156 PPPVSQQPFPY--PAYPPSYQAPLPPPPPVVASTPP-SAPPSNVVPALPPAKTNASSHPQ 212
PPPV P P P PP Y P PPPPPV + PP +PP V PP ++ P
Sbjct: 537 PPPVHSPPPPVHSPPPPPVYSPP-PPPPPVHSPPPPVFSPPPPVYS--PPPPVHSPPPPV 593
Query: 213 LKCPMAGTFYRCPGPGEPP 231
P + P P P
Sbjct: 594 HSPPPPAPVHSPPPPVHSP 612
Score = 41.2 bits (95), Expect = 7e-04
Identities = 29/79 (36%), Positives = 35/79 (43%), Gaps = 7/79 (8%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPL----PPPPPVVASTPPSAPPSNVV--PALPPAKTNASS 209
PPPV P P + PPS P+ PP PP + S P +PP V PPA A S
Sbjct: 615 PPPVYSPPPPVFSPPPSQSPPVVYSPPPRPPKINSPPVQSPPPAPVEKKETPPAHAPAPS 674
Query: 210 HPQ-LKCPMAGTFYRCPGP 227
+ + P G Y P P
Sbjct: 675 DDEFIIPPFIGHQYASPPP 693
Score = 38.5 bits (88), Expect = 0.005
Identities = 21/51 (41%), Positives = 25/51 (48%), Gaps = 6/51 (11%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLP----PPPPVVASTPPSAPPSNVVPALPP 202
PPPV P P + PP +P P PPPP +PP PP + P PP
Sbjct: 569 PPPVFSPPPPVYSPPPPVHSPPPPVHSPPPPAPVHSPP--PPVHSPPPPPP 617
Score = 38.1 bits (87), Expect = 0.006
Identities = 25/73 (34%), Positives = 29/73 (39%), Gaps = 1/73 (1%)
Query: 156 PPPV-SQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLK 214
PPPV S P P P + P PPPP P +PP V PPA ++ P
Sbjct: 552 PPPVYSPPPPPPPVHSPPPPVFSPPPPVYSPPPPVHSPPPPVHSPPPPAPVHSPPPPVHS 611
Query: 215 CPMAGTFYRCPGP 227
P Y P P
Sbjct: 612 PPPPPPVYSPPPP 624
Score = 37.4 bits (85), Expect = 0.010
Identities = 29/81 (35%), Positives = 35/81 (42%), Gaps = 12/81 (14%)
Query: 156 PPPVSQQPFPYPAY--PPSYQAPLPP---PPPVVASTPPSAPPSNVVPALPPAKTNASSH 210
PPP P P P + PP +P PP PPP V S PP AP V + PP +
Sbjct: 561 PPPPVHSP-PPPVFSPPPPVYSPPPPVHSPPPPVHSPPPPAP----VHSPPPPVHSPPPP 615
Query: 211 PQLKCPMAGTFYRCPGPGEPP 231
P + P F P P + P
Sbjct: 616 PPVYSPPPPVF--SPPPSQSP 634
Score = 32.7 bits (73), Expect = 0.25
Identities = 26/77 (33%), Positives = 31/77 (39%), Gaps = 8/77 (10%)
Query: 155 QPPPVSQQPF-PYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQL 213
QPP S QP PY P + + PPP PV ++PP V PP S P +
Sbjct: 495 QPPKESPQPDDPYDQSPVTKRRS-PPPAPV------NSPPPPVYSPPPPPPPVHSPPPPV 547
Query: 214 KCPMAGTFYRCPGPGEP 230
P Y P P P
Sbjct: 548 HSPPPPPVYSPPPPPPP 564
Score = 29.3 bits (64), Expect = 2.7
Identities = 17/41 (41%), Positives = 19/41 (45%), Gaps = 1/41 (2%)
Query: 155 QPPPVSQQPFPYPAYPPSYQAPLPPPPPV-VASTPPSAPPS 194
Q PPV P P P S PPP PV TPP+ P+
Sbjct: 632 QSPPVVYSPPPRPPKINSPPVQSPPPAPVEKKETPPAHAPA 672
>At1g54215 putative protein
Length = 169
Score = 47.0 bits (110), Expect = 1e-05
Identities = 34/109 (31%), Positives = 45/109 (41%), Gaps = 12/109 (11%)
Query: 152 EALQPPPVSQQPFPYPAYPPSYQAPLPPPPP--------VVASTPPSAPP-SNVVPAL-- 200
++ PP Q P P P PP P PPPPP V PP PP ++++ L
Sbjct: 31 QSQDPPLFPQSPPPPPPPPPPPPPPPPPPPPPPPAVNMSVETGIPPPPPPVTDMIKPLSS 90
Query: 201 -PPAKTNASSHPQLKCPMAGTFYRCPGPGEPPFVKVGDQVQKGQVVCII 248
PP + S P K P R P P P D + KG+ V ++
Sbjct: 91 PPPPQPPPRSQPPPKPPQKNLPRRHPPPPRSPEKPKRDGLNKGKTVGLV 139
>At1g49490 hypothetical protein
Length = 847
Score = 47.0 bits (110), Expect = 1e-05
Identities = 30/80 (37%), Positives = 34/80 (42%), Gaps = 11/80 (13%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPP----PPPVVASTPPSAPPSNVVPALPPAKTNASSHP 211
PPPV P PP Y +P PP PPP VAS PP +PP V + PP + P
Sbjct: 553 PPPVHSPP------PPVYSSPPPPHVYSPPPPVASPPPPSPPP-PVHSPPPPPVFSPPPP 605
Query: 212 QLKCPMAGTFYRCPGPGEPP 231
P Y P P P
Sbjct: 606 VFSPPPPSPVYSPPPPSHSP 625
Score = 40.8 bits (94), Expect = 0.001
Identities = 28/74 (37%), Positives = 34/74 (45%), Gaps = 2/74 (2%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPS-APPSNVVPALPPAKTNASSHPQLK 214
PPPV P P PP PPP PV + PPS +PP V PP + +H +
Sbjct: 588 PPPVHSPPPPPVFSPPPPVFSPPPPSPVYSPPPPSHSPPPPVYSPPPPTFSPPPTHNTNQ 647
Query: 215 CPM-AGTFYRCPGP 227
PM A T + P P
Sbjct: 648 PPMGAPTPTQAPTP 661
Score = 39.7 bits (91), Expect = 0.002
Identities = 37/135 (27%), Positives = 49/135 (35%), Gaps = 16/135 (11%)
Query: 92 LPKEKDNHNGKP------PGPSASTDASSVSTFMDQVSELVKLVDSRDIMELELKQAGYE 145
LPK+K++ +P P P S Q+ E K V + + Y+
Sbjct: 454 LPKQKESPKPQPSKPEDSPKPEQPKPEESPKPEQPQIPEPTKPVSPPNEAQGPTPDDPYD 513
Query: 146 LMIRKKEALQPPPVSQQPFPYPAYPPSYQAPLPP-----PPPVVASTPP----SAPPSNV 196
K PPP + P PP +P PP PPP V S PP S PP +V
Sbjct: 514 ASPVKNRRSPPPPKVEDTRVPPPQPPM-PSPSPPSPIYSPPPPVHSPPPPVYSSPPPPHV 572
Query: 197 VPALPPAKTNASSHP 211
PP + P
Sbjct: 573 YSPPPPVASPPPPSP 587
Score = 36.2 bits (82), Expect = 0.022
Identities = 18/68 (26%), Positives = 30/68 (43%)
Query: 149 RKKEALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNAS 208
++KE+ +P P + P P P ++P P P + T P +PP+ P +AS
Sbjct: 456 KQKESPKPQPSKPEDSPKPEQPKPEESPKPEQPQIPEPTKPVSPPNEAQGPTPDDPYDAS 515
Query: 209 SHPQLKCP 216
+ P
Sbjct: 516 PVKNRRSP 523
Score = 35.0 bits (79), Expect = 0.050
Identities = 26/81 (32%), Positives = 28/81 (34%), Gaps = 5/81 (6%)
Query: 156 PPPVSQQPFPYPAYPP-----SYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSH 210
PPPV P P Y P S P PPPP PP P V + PP S
Sbjct: 560 PPPVYSSPPPPHVYSPPPPVASPPPPSPPPPVHSPPPPPVFSPPPPVFSPPPPSPVYSPP 619
Query: 211 PQLKCPMAGTFYRCPGPGEPP 231
P P + P PP
Sbjct: 620 PPSHSPPPPVYSPPPPTFSPP 640
Score = 33.9 bits (76), Expect = 0.11
Identities = 25/82 (30%), Positives = 32/82 (38%), Gaps = 20/82 (24%)
Query: 156 PPPVSQQPFPYPAY---PPSYQA-----PLPPP------------PPVVASTPPSAPPSN 195
PPPV P P P Y PPS+ PPP PP+ A TP AP +
Sbjct: 603 PPPVFSPPPPSPVYSPPPPSHSPPPPVYSPPPPTFSPPPTHNTNQPPMGAPTPTQAPTPS 662
Query: 196 VVPALPPAKTNASSHPQLKCPM 217
P ++ S Q+ P+
Sbjct: 663 SETTQVPTPSSESDQSQILSPV 684
Score = 32.7 bits (73), Expect = 0.25
Identities = 35/137 (25%), Positives = 50/137 (35%), Gaps = 14/137 (10%)
Query: 82 PVLINEPNGALPKEKDNHNGKPPGPSASTDASSVSTFMDQVSELVKLVDSRDIMELELKQ 141
PV P P N N P G T A + S+ QV D I+
Sbjct: 628 PVYSPPPPTFSPPPTHNTNQPPMGAPTPTQAPTPSSETTQVPTPSSESDQSQILS----- 682
Query: 142 AGYELMIRKKEALQPPPVSQQP--FPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPA 199
++ +Q S +P P P+ SYQAP PV A TP AP ++ +
Sbjct: 683 -----PVQAPTPVQSSTPSSEPTQVPTPSSSESYQAP--NLSPVQAPTPVQAPTTSSETS 735
Query: 200 LPPAKTNASSHPQLKCP 216
P ++ S+ + P
Sbjct: 736 QVPTPSSESNQSPSQAP 752
Score = 31.2 bits (69), Expect = 0.72
Identities = 20/65 (30%), Positives = 27/65 (40%), Gaps = 3/65 (4%)
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPP---PPPVVASTPPSAPPSNVVPALPPAKTNASSHPQ 212
PPPV P P + PP++ PP P P A TP S P+ ++ S Q
Sbjct: 626 PPPVYSPPPPTFSPPPTHNTNQPPMGAPTPTQAPTPSSETTQVPTPSSESDQSQILSPVQ 685
Query: 213 LKCPM 217
P+
Sbjct: 686 APTPV 690
Score = 29.6 bits (65), Expect = 2.1
Identities = 18/53 (33%), Positives = 26/53 (48%), Gaps = 3/53 (5%)
Query: 154 LQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTN 206
+Q P S +P P +AP P PV S+ PS+ PS ++PP + N
Sbjct: 769 VQSPTPSSEPVSSPEQSEEVEAP--EPTPVNPSSVPSSSPSTDT-SIPPPENN 818
Score = 28.5 bits (62), Expect = 4.7
Identities = 26/117 (22%), Positives = 43/117 (36%), Gaps = 11/117 (9%)
Query: 104 PGPSASTDASSVSTFMDQVSELVKLVDSRDIMELELKQAGYELMIRKKEALQPPPVSQQP 163
P PS+ +D S + + + + + S + ++ + +Q P Q P
Sbjct: 669 PTPSSESDQSQILSPVQAPTPVQSSTPSSEPTQVPTPSSSESYQAPNLSPVQAPTPVQAP 728
Query: 164 --------FPYPAYPPSY---QAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASS 209
P P+ + QAP P PV A TP S P + P+ P + S
Sbjct: 729 TTSSETSQVPTPSSESNQSPSQAPTPILEPVHAPTPNSKPVQSPTPSSEPVSSPEQS 785
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.133 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,547,246
Number of Sequences: 26719
Number of extensions: 377356
Number of successful extensions: 10879
Number of sequences better than 10.0: 421
Number of HSP's better than 10.0 without gapping: 248
Number of HSP's successfully gapped in prelim test: 185
Number of HSP's that attempted gapping in prelim test: 1945
Number of HSP's gapped (non-prelim): 2621
length of query: 302
length of database: 11,318,596
effective HSP length: 99
effective length of query: 203
effective length of database: 8,673,415
effective search space: 1760703245
effective search space used: 1760703245
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC119411.2


