

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119408.6 - phase: 0
(535 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
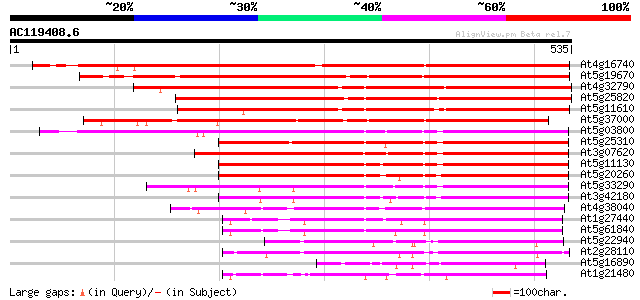
Score E
Sequences producing significant alignments: (bits) Value
At4g16740 limonene cyclase like protein 524 e-149
At5g19670 putative protein 491 e-139
At4g32790 unknown protein 473 e-133
At5g25820 putative protein 459 e-129
At5g11610 unknown protein 415 e-116
At5g37000 putative protein 365 e-101
At5g03800 putative protein 343 2e-94
At5g25310 putative protein 306 2e-83
At3g07620 hypothetical protein 299 3e-81
At5g11130 putative protein 271 9e-73
At5g20260 putative protein 269 3e-72
At5g33290 unknown protein 260 2e-69
At3g42180 unknown protein 260 2e-69
At4g38040 unknown protein 243 2e-64
At1g27440 unknown protein 125 6e-29
At5g61840 unknown protein 123 2e-28
At5g22940 putative protein 108 1e-23
At2g28110 unknown protein 106 4e-23
At5g16890 unknown protein (At5g16890) 72 6e-13
At1g21480 unknown protein 72 6e-13
>At4g16740 limonene cyclase like protein
Length = 1024
Score = 524 bits (1350), Expect = e-149
Identities = 281/521 (53%), Positives = 346/521 (65%), Gaps = 29/521 (5%)
Query: 22 RHKIDWRRLFLLVSIFTASGFVFQMIVHTSVISPNKHFHFPPGAADSYKSSNSTIQSNEF 81
R +I WR FLL SI T +++HT S + + S+ S +
Sbjct: 513 RQRIKWRNAFLLGSIMTT----IVVLLHTPTFS----------VFSDEEETESSSSSPIY 558
Query: 82 IKETILQNVHLTLPNSAVSP----KLSNNFVQSVSVEADS--INTEARRKKDRNLANTST 135
+ ++ N+H+ + V + VQ + EA ++ + R++K R
Sbjct: 559 LNGSLHLNIHIVSSEAKVENFHTLRTRTPIVQLNASEASEAVLSRKRRKRKKRKKTKDDL 618
Query: 136 TVTTSFPRGR-VPSGKQTDIRLITPTEALVYARKEIDHVTSVNEDPDLYAPLFRNVSVFK 194
+T P R V S + + P +AL YA+ EI V D DL+APLFRN+SVFK
Sbjct: 619 ILTDPPPAPRHVLSSSERRALSLPPKKALTYAKLEIQRAPEVINDTDLFAPLFRNLSVFK 678
Query: 195 RSYELMETVLKVYIYRDGSRPIFHNPSLKGIYASEGWFMKLMQENKQFVTKDPERAHLFY 254
RSYELME +LKVYIY DG +PIFH P L GIYASEGWFMKLM+ NKQFVTK+PERAHLFY
Sbjct: 679 RSYELMELILKVYIYPDGDKPIFHEPHLNGIYASEGWFMKLMESNKQFVTKNPERAHLFY 738
Query: 255 LPYSARQMEVTLYVPGSHDLKPLSIFLRDYVNKIAAKYPFWNRTHGSDHFLVACHEWGPY 314
+PYS +Q++ S P L + ++ T G +WGPY
Sbjct: 739 MPYSVKQLQKKTTSTCSPSNTPSGTALMGQIISLSLA------TIGYRKCFYVKDQWGPY 792
Query: 315 TVTEHEELARNTLKALCNADLSERIFIEGRDVSLPETTIRAPRRPLRYLG-GNRASLRPI 373
TV EH EL RN +KALCNADLS+ IF+ G+DVSLPET+IR RPLR +G GNR S RPI
Sbjct: 793 TVNEHPELKRNAIKALCNADLSDGIFVPGKDVSLPETSIRNAGRPLRNIGNGNRVSQRPI 852
Query: 374 LAFFAGSMHGRVRPTLLKYWGGEKYEDMKIYKPLPLRVSKKMTYIQHMKSSKYCLCPMGF 433
LAFFAG++HGRVRP LLK+W K EDMKIY PLP V++KMTY+QHMKSSKYCLCPMG+
Sbjct: 853 LAFFAGNLHGRVRPKLLKHWRN-KDEDMKIYGPLPHNVARKMTYVQHMKSSKYCLCPMGY 911
Query: 434 EVNSPRIVEAIYYECVPVIIADNFVLPLSEVLDWSAFSVVVAEKDIPRLKDILLSIPMRK 493
EVNSPRIVEAIYYECVPV+IADNF+LP S+VLDWSAFSVVV EK+IPRLK+ILL IPMR+
Sbjct: 912 EVNSPRIVEAIYYECVPVVIADNFMLPFSDVLDWSAFSVVVPEKEIPRLKEILLEIPMRR 971
Query: 494 YVAMQNNVKMVQKHFLWNPKPIRYDLFHMILHSIWLNKLNQ 534
Y+ MQ+NVKMVQ+HFLW+PKP +YD+FHMILHSIW N LNQ
Sbjct: 972 YLKMQSNVKMVQRHFLWSPKPRKYDVFHMILHSIWFNLLNQ 1012
>At5g19670 putative protein
Length = 600
Score = 491 bits (1263), Expect = e-139
Identities = 249/468 (53%), Positives = 326/468 (69%), Gaps = 22/468 (4%)
Query: 67 DSYKSSNSTIQSNEFIKETILQNVHLTLPNSAVSPKLSNNFVQSVSVEADSINTEARRKK 126
+ Y+ N T+QS + +K +IL +S SP N+ + ++ + +KK
Sbjct: 151 NGYQVQNVTVQSQKNVKSSILSG-----GSSIASPASGNSSLL--------VSKKVSKKK 197
Query: 127 DRNLANTSTTVTTSFPRGRVPSGKQTDIRLITPTEALVYARKEIDHVTSVNEDPDLYAPL 186
+VTT R+ + + R + ++ ARKEI++ + +LY P+
Sbjct: 198 KMRCDLPPKSVTTIDEMNRILARHRRTSRAME----ILTARKEIENAPVAKLERELYPPI 253
Query: 187 FRNVSVFKRSYELMETVLKVYIYRDGSRPIFHNPSLKGIYASEGWFMKLMQENKQFVTKD 246
FRNVS+FKRSYELME +LKVY+Y++G+RPIFH P LKG+YASEGWFMKLM+ NKQ+ KD
Sbjct: 254 FRNVSLFKRSYELMERILKVYVYKEGNRPIFHTPILKGLYASEGWFMKLMEGNKQYTVKD 313
Query: 247 PERAHLFYLPYSARQMEVTLYVPGSHDLKPLSIFLRDYVNKIAAKYPFWNRTHGSDHFLV 306
P +AHL+Y+P+SAR +E TLYV SH+ L FL++Y I++KYPF+NRT G+DHFLV
Sbjct: 314 PRKAHLYYMPFSARMLEYTLYVRNSHNRTNLRQFLKEYTEHISSKYPFFNRTDGADHFLV 373
Query: 307 ACHEWGPYTVTEHEELARNTLKALCNADLSERIFIEGRDVSLPETTIRAPRRPLRYLGGN 366
ACH+W PY H E + +KALCNAD++ I GRD+SLPET +RA + PLR LGG
Sbjct: 374 ACHDWAPYETRHHME---HCIKALCNADVTAGFKI-GRDISLPETYVRAAKNPLRDLGGK 429
Query: 367 RASLRPILAFFAGSMHGRVRPTLLKYWGGEKYEDMKIYKPLPLRVSKKMTYIQHMKSSKY 426
S R LAF+AGSMHG +R LL++W +K DMKI+ +P V+ KM YI+ MKSSKY
Sbjct: 430 PPSQRRTLAFYAGSMHGYLRQILLQHW-KDKDPDMKIFGRMPFGVASKMNYIEQMKSSKY 488
Query: 427 CLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLSEVLDWSAFSVVVAEKDIPRLKDIL 486
C+CP G+EVNSPR+VE+I+YECVPVII+DNFV P EVLDWSAFSV+VAEKDIPRLKDIL
Sbjct: 489 CICPKGYEVNSPRVVESIFYECVPVIISDNFVPPFFEVLDWSAFSVIVAEKDIPRLKDIL 548
Query: 487 LSIPMRKYVAMQNNVKMVQKHFLWNPKPIRYDLFHMILHSIWLNKLNQ 534
LSIP KYV MQ V+ Q+HFLW+ KP +YDLFHM+LHSIW N++ Q
Sbjct: 549 LSIPEDKYVKMQMAVRKAQRHFLWHAKPEKYDLFHMVLHSIWYNRVFQ 596
>At4g32790 unknown protein
Length = 593
Score = 473 bits (1217), Expect = e-133
Identities = 234/421 (55%), Positives = 302/421 (71%), Gaps = 9/421 (2%)
Query: 119 NTEARRKKDRNLANTSTTVTTSFP----RGRVPSGKQTDIRLITPTEALVYARKEIDHVT 174
N + KK N++N+ T + R R T L+YAR +I++
Sbjct: 178 NLSSEVKKFMNVSNSGVVSITEMMNLLHQSRTSHVSLKVKRSSTIDHELLYARTQIENPP 237
Query: 175 SVNEDPDLYAPLFRNVSVFKRSYELMETVLKVYIYRDGSRPIFHNPSLKGIYASEGWFMK 234
+ DP L+ PL+ N+S+FKRSYELME LKVY+YR+G RP+ H P LKGIYASEGWFMK
Sbjct: 238 LIENDPLLHTPLYWNLSMFKRSYELMEKKLKVYVYREGKRPVLHKPVLKGIYASEGWFMK 297
Query: 235 LMQENKQFVTKDPERAHLFYLPYSARQMEVTLYVPGSHDLKPLSIFLRDYVNKIAAKYPF 294
++ ++ FVTKDP +AHLFYLP+S++ +E TLYVPGSH K L FL++Y++ I++KY F
Sbjct: 298 QLKSSRTFVTKDPRKAHLFYLPFSSKMLEETLYVPGSHSDKNLIQFLKNYLDMISSKYSF 357
Query: 295 WNRTHGSDHFLVACHEWGPYTVTEHEELARNTLKALCNADLSERIFIEGRDVSLPETTIR 354
WN+T GSDHFLVACH+W P +E + ++ALCN+D+SE F+ G+DV+LPETTI
Sbjct: 358 WNKTGGSDHFLVACHDWAP---SETRQYMAKCIRALCNSDVSEG-FVFGKDVALPETTIL 413
Query: 355 APRRPLRYLGGNRASLRPILAFFAGSMHGRVRPTLLKYWGGEKYEDMKIYKPLPLRVSKK 414
PRRPLR LGG S R ILAFFAG MHG +RP LL+ WGG + DMKI+ +P KK
Sbjct: 414 VPRRPLRALGGKPVSQRQILAFFAGGMHGYLRPLLLQNWGGNRDPDMKIFSEIPKSKGKK 473
Query: 415 MTYIQHMKSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLSEVLDWSAFSVVV 474
+Y+++MKSSKYC+CP G EVNSPR+VEA++YECVPVII+DNFV P EVL+W +F+V V
Sbjct: 474 -SYMEYMKSSKYCICPKGHEVNSPRVVEALFYECVPVIISDNFVPPFFEVLNWESFAVFV 532
Query: 475 AEKDIPRLKDILLSIPMRKYVAMQNNVKMVQKHFLWNPKPIRYDLFHMILHSIWLNKLNQ 534
EKDIP LK+IL+SI +Y MQ VKMVQKHFLW+ KP R+D+FHMILHSIW N++ Q
Sbjct: 533 LEKDIPDLKNILVSITEERYREMQMRVKMVQKHFLWHSKPERFDIFHMILHSIWYNRVFQ 592
Query: 535 I 535
I
Sbjct: 593 I 593
>At5g25820 putative protein
Length = 654
Score = 459 bits (1180), Expect = e-129
Identities = 218/379 (57%), Positives = 286/379 (74%), Gaps = 7/379 (1%)
Query: 159 PTEALVYARKEIDHVTSVNEDPDLYAPLFRNVSVFKRSYELMETVLKVYIYRDGSRPIFH 218
P L+ A+ +I++ ++DP LYAPL+RNVS+FKRSYELME +LKVY Y++G++PI H
Sbjct: 279 PDLELLQAKYDIENAPIDDKDPFLYAPLYRNVSMFKRSYELMEKILKVYAYKEGNKPIMH 338
Query: 219 NPSLKGIYASEGWFMKLMQENK-QFVTKDPERAHLFYLPYSARQMEVTLYVPGSHDLKPL 277
+P L+GIYASEGWFM +++ N +FVTKDP +AHLFYLP+S+R +EVTLYV SH + L
Sbjct: 339 SPILRGIYASEGWFMNIIESNNNKFVTKDPAKAHLFYLPFSSRMLEVTLYVQDSHSHRNL 398
Query: 278 SIFLRDYVNKIAAKYPFWNRTHGSDHFLVACHEWGPYTVTEHEELARNTLKALCNADLSE 337
+L+DY++ I+AKYPFWNRT G+DHFL ACH+W P +H +++ALCN+D+ E
Sbjct: 399 IKYLKDYIDFISAKYPFWNRTSGADHFLAACHDWAPSETRKH---MAKSIRALCNSDVKE 455
Query: 338 RIFIEGRDVSLPETTIRAPRRPLRYLGGNRASLRPILAFFAGSM-HGRVRPTLLKYWGGE 396
F+ G+D SLPET +R P++PL +GG A+ RPILAFFAG HG +RP LL YWG
Sbjct: 456 G-FVFGKDTSLPETFVRDPKKPLSNMGGKSANQRPILAFFAGKPDHGYLRPILLSYWGNN 514
Query: 397 KYEDMKIYKPLPLRVSKKMTYIQHMKSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIADN 456
K D+KI+ LP R Y+Q MK+SKYC+C GFEVNSPR+VEAI+Y+CVPVII+DN
Sbjct: 515 KDPDLKIFGKLP-RTKGNKNYLQFMKTSKYCICAKGFEVNSPRVVEAIFYDCVPVIISDN 573
Query: 457 FVLPLSEVLDWSAFSVVVAEKDIPRLKDILLSIPMRKYVAMQNNVKMVQKHFLWNPKPIR 516
FV P EVL+W +F++ + EKDIP LK IL+SIP +Y +MQ VK VQKHFLW+ KP +
Sbjct: 574 FVPPFFEVLNWESFAIFIPEKDIPNLKKILMSIPESRYRSMQMRVKKVQKHFLWHAKPEK 633
Query: 517 YDLFHMILHSIWLNKLNQI 535
YD+FHMILHSIW N++ QI
Sbjct: 634 YDMFHMILHSIWYNRVFQI 652
>At5g11610 unknown protein
Length = 546
Score = 415 bits (1066), Expect = e-116
Identities = 203/377 (53%), Positives = 270/377 (70%), Gaps = 12/377 (3%)
Query: 161 EALVYARKEIDHVTSVNEDPDLYAPLFRNVSVFKRSYELMETVLKVYIYRDGSRPIFHNP 220
+ L AR +I V +D LYAPL+ N+S+FKRSYELME LKVY+Y +G RPIFH P
Sbjct: 177 QELKTARDKIKKAALVKKDDTLYAPLYHNISIFKRSYELMEQTLKVYVYSEGDRPIFHQP 236
Query: 221 S--LKGIYASEGWFMKLMQENKQFVTKDPERAHLFYLPYSARQMEVTLYVPGSHDLKPLS 278
++GIYASEGWFMKLM+ + +F+TKDP +AHLFY+P+S+R ++ LYV SH L
Sbjct: 237 EAIMEGIYASEGWFMKLMESSHRFLTKDPTKAHLFYIPFSSRILQQKLYVHDSHSRNNLV 296
Query: 279 IFLRDYVNKIAAKYPFWNRTHGSDHFLVACHEWGPYTVTEHEELARNTLKALCNADLSER 338
+L +Y++ IA+ YP WNRT GSDHF ACH+W P TE N ++ALCNAD+
Sbjct: 297 KYLGNYIDLIASNYPSWNRTCGSDHFFTACHDWAP---TETRGPYINCIRALCNADVGID 353
Query: 339 IFIEGRDVSLPETTIRAPRRPLRYLGGNRASLRPILAFFAGSMHGRVRPTLLKYWGGEKY 398
F+ G+DVSLPET + + + P +GG+R S R ILAFFAGS+HG VRP LL W
Sbjct: 354 -FVVGKDVSLPETKVSSLQNPNGKIGGSRPSKRTILAFFAGSLHGYVRPILLNQWSSRPE 412
Query: 399 EDMKIYKPLPLRVSKKMTYIQHMKSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFV 458
+DMKI+ R+ K +YI++MK S++C+C G+EVNSPR+VE+I Y CVPVII+DNFV
Sbjct: 413 QDMKIFN----RIDHK-SYIRYMKRSRFCVCAKGYEVNSPRVVESILYGCVPVIISDNFV 467
Query: 459 LPLSEVLDWSAFSVVVAEKDIPRLKDILLSIPMRKYVAMQNNVKMVQKHFLWNP-KPIRY 517
P E+L+W +F+V V EK+IP L+ IL+SIP+R+YV MQ V VQKHF+W+ +P+RY
Sbjct: 468 PPFLEILNWESFAVFVPEKEIPNLRKILISIPVRRYVEMQKRVLKVQKHFMWHDGEPVRY 527
Query: 518 DLFHMILHSIWLNKLNQ 534
D+FHMILHS+W N++ Q
Sbjct: 528 DIFHMILHSVWYNRVFQ 544
>At5g37000 putative protein
Length = 547
Score = 365 bits (937), Expect = e-101
Identities = 203/469 (43%), Positives = 285/469 (60%), Gaps = 35/469 (7%)
Query: 71 SSNSTIQSNEFIKETI----LQNVHLTLPNSAVSPKLSNNFVQSVSVEADSINT----EA 122
S N + S E ++ + + N+ L P V S + SV +
Sbjct: 86 SENDVVISKEKVEMNVSFIAIGNISLRNPKMVVVSSESESDPNSVMIRVKDSRKGNVLSL 145
Query: 123 RRKKDRN---LANTSTTVTTSFPRGRVPSGKQTDIRLITPTEALVYARKEIDHVTSVNED 179
RR K + ++ ++ + S + P + + R ++ AR EI+ V+ V++
Sbjct: 146 RRHKQGSAISISQMNSLLIQSLSSFKSPKPRWSSAR----DSEMLSARSEIEKVSLVHDF 201
Query: 180 PDLYAPLFRNVSVFKRS--------------YELMETVLKVYIYRDGSRPIFHNPSLKGI 225
L ++RN+S F RS Y+LME LK+Y+Y++G +PIFH P +GI
Sbjct: 202 LGLNPLVYRNISKFLRSGDMSRFSMCCLFRSYDLMERKLKIYVYKEGGKPIFHTPMPRGI 261
Query: 226 YASEGWFMKLMQENKQFVTKDPERAHLFYLPYSARQMEVTLYVPGSHDLKPLSIFLRDYV 285
YASEGWFMKLM+ NK+FV KDP +AHLFY+P S + + +L + K L+ L++YV
Sbjct: 262 YASEGWFMKLMESNKKFVVKDPRKAHLFYIPISIKALRSSLGLDFQTP-KSLADHLKEYV 320
Query: 286 NKIAAKYPFWNRTHGSDHFLVACHEWGPYTVTEHEELARNTLKALCNADLSERIFIEGRD 345
+ IA KY FWNRT G+DHFLVACH+WG T+ +N++++LCN+++++ I G D
Sbjct: 321 DLIAGKYKFWNRTGGADHFLVACHDWGNKLTTK---TMKNSVRSLCNSNVAQGFRI-GTD 376
Query: 346 VSLPETTIRAPRRPLRYLGGNRASLRPILAFFAGSMHGRVRPTLLKYWGGEKYEDMKIYK 405
+LP T IR+ PL YLGG +S R ILAFFAGSMHG +RP L+K W K DMKI+
Sbjct: 377 TALPVTYIRSSEAPLEYLGGKTSSERKILAFFAGSMHGYLRPILVKLWEN-KEPDMKIFG 435
Query: 406 PLPLRVSKKMTYIQHMKSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLSEVL 465
P+P K Y ++MKSS+YC+C G+EV++PR+VEAI ECVPVIIADN+V P EVL
Sbjct: 436 PMPRDPKSKKQYREYMKSSRYCICARGYEVHTPRVVEAIINECVPVIIADNYVPPFFEVL 495
Query: 466 DWSAFSVVVAEKDIPRLKDILLSIPMRKYVAMQNNVKMVQKHFLWNPKP 514
+W F+V V EKDIP L++ILLSIP +Y+ MQ VK VQ+HFLW+ KP
Sbjct: 496 NWEEFAVFVEEKDIPNLRNILLSIPEDRYIGMQARVKAVQQHFLWHKKP 544
>At5g03800 putative protein
Length = 1388
Score = 343 bits (879), Expect = 2e-94
Identities = 197/511 (38%), Positives = 289/511 (56%), Gaps = 29/511 (5%)
Query: 29 RLFL-LVSIFTASGFVFQMIVHTSVISPNKHFHFPPGAADSYKSSNSTIQSNEFIKETIL 87
+LFL +V + SGFVF I G DS S + + L
Sbjct: 895 KLFLFMVPLVVISGFVFVNI----------------GPKDSTSLLTSLSTTTSHLPPPFL 938
Query: 88 QNVHLTLPNSAVSPKLSNNFVQSVSVEADSINTEARRKKDRNLAN-TSTTVTTSFPRGRV 146
P+ + L + S+S + +SI + R N+ N T+T+ S
Sbjct: 939 STAPAPAPSPLLPEILPSLPASSLSTKVESIQGDYNRTIQLNMINVTATSNNVSSTASLE 998
Query: 147 PSGKQTDIRLITPTEALVYARKEIDHVTSVN--EDPDLY--APLFRNVSVFKRSYELMET 202
P ++ L L AR I + + +DPD P++ N VF RSY ME
Sbjct: 999 PKKRRVLSNLEKIEFKLQKARASIKAASMDDPVDDPDYVPLGPMYWNAKVFHRSYLEMEK 1058
Query: 203 VLKVYIYRDGSRPIFHNPSLKGIYASEGWFMKLMQENKQFVTKDPERAHLFYLPYSARQM 262
K+Y+Y++G P+FH+ K IY+ EG F+ ++ + +F T +P++AH+FYLP+S +M
Sbjct: 1059 QFKIYVYKEGEPPLFHDGPCKSIYSMEGSFIYEIETDTRFRTNNPDKAHVFYLPFSVVKM 1118
Query: 263 EVTLYVPGSHDLKPLSIFLRDYVNKIAAKYPFWNRTHGSDHFLVACHEWGPYTVTEHEEL 322
+Y S D P+ ++DY+N + KYP+WNR+ G+DHF+++CH+WGP H L
Sbjct: 1119 VRYVYERNSRDFSPIRNTVKDYINLVGDKYPYWNRSIGADHFILSCHDWGPEASFSHPHL 1178
Query: 323 ARNTLKALCNADLSERIFIEGRDVSLPETTIRAPRRPLRYLGGNRASLRPILAFFAGSMH 382
N+++ALCNA+ SER F +DVS+PE +R +GG S RPILAFFAG +H
Sbjct: 1179 GHNSIRALCNANTSER-FKPRKDVSIPEINLRTGSL-TGLVGGPSPSSRPILAFFAGGVH 1236
Query: 383 GRVRPTLLKYWGGEKYEDMKIYKPLPLRVSKKMTYIQHMKSSKYCLCPMGFEVNSPRIVE 442
G VRP LL++W K D++++K LP + +Y M++SK+C+CP G+EV SPRIVE
Sbjct: 1237 GPVRPVLLQHW-ENKDNDIRVHKYLP----RGTSYSDMMRNSKFCICPSGYEVASPRIVE 1291
Query: 443 AIYYECVPVIIADNFVLPLSEVLDWSAFSVVVAEKDIPRLKDILLSIPMRKYVAMQNNVK 502
A+Y CVPV+I +V P S+VL+W +FSV+V+ +DIP LK IL SI R+Y+ M V
Sbjct: 1292 ALYSGCVPVLINSGYVPPFSDVLNWRSFSVIVSVEDIPNLKTILTSISPRQYLRMYRRVL 1351
Query: 503 MVQKHFLWNPKPIRYDLFHMILHSIWLNKLN 533
V++HF N R+D+FHMILHSIW+ +LN
Sbjct: 1352 KVRRHFEVNSPAKRFDVFHMILHSIWVRRLN 1382
>At5g25310 putative protein
Length = 334
Score = 306 bits (783), Expect = 2e-83
Identities = 152/337 (45%), Positives = 221/337 (65%), Gaps = 10/337 (2%)
Query: 200 METVLKVYIYRDGSRPIFHNPSLKGIYASEGWFMKLMQENK-QFVTKDPERAHLFYLPYS 258
ME KVY+Y +G P+ H+ K +YA EG F+ M++ + +F T DP +A++++LP+S
Sbjct: 1 MEKRFKVYVYEEGEPPLVHDGPCKSVYAVEGRFITEMEKRRTKFRTYDPNQAYVYFLPFS 60
Query: 259 ARQMEVTLYVPGSHDLKPLSIFLRDYVNKIAAKYPFWNRTHGSDHFLVACHEWGPYTVTE 318
+ LY G+ D KPL F+ DY+ ++ +PFWNRT+G+DHF++ CH+WGP T
Sbjct: 61 VTWLVRYLY-EGNSDAKPLKTFVSDYIRLVSTNHPFWNRTNGADHFMLTCHDWGPLTSQA 119
Query: 319 HEELARNTLKALCNADLSERIFIEGRDVSLPETTIRAPR--RPLRYLGGNRASLRPILAF 376
+ +L +++ +CNA+ SE F +DV+LPE + LR AS RP L F
Sbjct: 120 NRDLFNTSIRVMCNANSSEG-FNPTKDVTLPEIKLYGGEVDHKLRLSKTLSASPRPYLGF 178
Query: 377 FAGSMHGRVRPTLLKYWGGEKYEDMKIYKPLPLRVSKKMTYIQHMKSSKYCLCPMGFEVN 436
FAG +HG VRP LLK+W ++ DM +Y+ LP K + Y M+SSK+C CP G+EV
Sbjct: 179 FAGGVHGPVRPILLKHWK-QRDLDMPVYEYLP----KHLNYYDFMRSSKFCFCPSGYEVA 233
Query: 437 SPRIVEAIYYECVPVIIADNFVLPLSEVLDWSAFSVVVAEKDIPRLKDILLSIPMRKYVA 496
SPR++EAIY EC+PVI++ NFVLP ++VL W FSV+V +IPRLK+IL+SI KY
Sbjct: 234 SPRVIEAIYSECIPVILSVNFVLPFTDVLRWETFSVLVDVSEIPRLKEILMSISNEKYEW 293
Query: 497 MQNNVKMVQKHFLWNPKPIRYDLFHMILHSIWLNKLN 533
+++N++ V++HF N P R+D FH+ LHSIWL +LN
Sbjct: 294 LKSNLRYVRRHFELNDPPQRFDAFHLTLHSIWLRRLN 330
>At3g07620 hypothetical protein
Length = 470
Score = 299 bits (765), Expect = 3e-81
Identities = 152/358 (42%), Positives = 224/358 (62%), Gaps = 8/358 (2%)
Query: 177 NEDPDLYAPLFRNVSVFKRSYELMETVLKVYIYRDGSRPIFHNPSLKGIYASEGWFMKLM 236
+ED + ++RN F RSY LME + K+Y+Y +G PIFH K IY+ EG F+ M
Sbjct: 116 DEDYVPHGDIYRNPYAFHRSYLLMEKMFKIYVYEEGDPPIFHYGLCKDIYSMEGLFLNFM 175
Query: 237 QENK-QFVTKDPERAHLFYLPYSARQMEVTLYVPGSHDLKPLSIFLRDYVNKIAAKYPFW 295
+ + ++ T+DP++AH+++LP+S + L+ P D L + DYV I+ KYP+W
Sbjct: 176 ENDVLKYRTRDPDKAHVYFLPFSVVMILHHLFDPVVRDKAVLERVIADYVQIISKKYPYW 235
Query: 296 NRTHGSDHFLVACHEWGPYTVTEHEELARNTLKALCNADLSERIFIEGRDVSLPETTIRA 355
N + G DHF+++CH+WG ++L N+++ LCNA++SE F +D PE +
Sbjct: 236 NTSDGFDHFMLSCHDWGHRATWYVKKLFFNSIRVLCNANISE-YFNPEKDAPFPEINLLT 294
Query: 356 PRRPLRYLGGNRASLRPILAFFAGSMHGRVRPTLLKYWGGEKYEDMKIYKPLPLRVSKKM 415
GG R LAFFAG HG++RP LL +W EK +D+ +Y+ LP +
Sbjct: 295 GDIN-NLTGGLDPISRTTLAFFAGKSHGKIRPVLLNHWK-EKDKDILVYENLP----DGL 348
Query: 416 TYIQHMKSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLSEVLDWSAFSVVVA 475
Y + M+ S++C+CP G EV SPR+ EAIY CVPV+I++N+VLP S+VL+W FSV V+
Sbjct: 349 DYTEMMRKSRFCICPSGHEVASPRVPEAIYSGCVPVLISENYVLPFSDVLNWEKFSVSVS 408
Query: 476 EKDIPRLKDILLSIPMRKYVAMQNNVKMVQKHFLWNPKPIRYDLFHMILHSIWLNKLN 533
K+IP LK IL+ IP +Y+ + VK V++H L N P RYD+F+MI+HSIWL +LN
Sbjct: 409 VKEIPELKRILMDIPEERYMRLYEGVKKVKRHILVNDPPKRYDVFNMIIHSIWLRRLN 466
>At5g11130 putative protein
Length = 336
Score = 271 bits (692), Expect = 9e-73
Identities = 148/337 (43%), Positives = 206/337 (60%), Gaps = 11/337 (3%)
Query: 200 METVLKVYIYRDGSRPIFHNPSLKGIYASEGWFMKLMQE-NKQFVTKDPERAHLFYLPYS 258
ME K++ YR+G P+FH L IYA EG FM ++ N +F PE A +FY+P
Sbjct: 1 MEKRFKIWTYREGEAPLFHKGPLNNIYAIEGQFMDEIENGNSRFKAASPEEATVFYIPVG 60
Query: 259 ARQMEVTLYVP-GSHDLKPLSIFLRDYVNKIAAKYPFWNRTHGSDHFLVACHEWGPYTVT 317
+ +Y P S+ L ++DY++ I+ +YP+WNR+ G+DHF ++CH+W P
Sbjct: 61 IVNIIRFVYRPYTSYARDRLQNIVKDYISLISNRYPYWNRSRGADHFFLSCHDWAPDVSA 120
Query: 318 EHEELARNTLKALCNADLSERIFIEGRDVSLPETTIRAPRRPLRYLG-GNRASLRPILAF 376
EL ++ ++ALCNA+ SE F RDVSLPE I P L ++ G R +LAF
Sbjct: 121 VDPELYKHFIRALCNANSSEG-FTPMRDVSLPEINI--PHSQLGFVHTGEPPQNRKLLAF 177
Query: 377 FAGSMHGRVRPTLLKYWGGEKYEDMKIYKPLPLRVSKKMTYIQHMKSSKYCLCPMGFEVN 436
FAG HG VR L ++W EK +D+ +Y+ LP K M Y + M +K+CLCP G+EV
Sbjct: 178 FAGGSHGDVRKILFQHWK-EKDKDVLVYENLP----KTMNYTKMMDKAKFCLCPSGWEVA 232
Query: 437 SPRIVEAIYYECVPVIIADNFVLPLSEVLDWSAFSVVVAEKDIPRLKDILLSIPMRKYVA 496
SPRIVE++Y CVPVIIAD +VLP S+VL+W FSV + +P +K IL +I +Y+
Sbjct: 233 SPRIVESLYSGCVPVIIADYYVLPFSDVLNWKTFSVHIPISKMPDIKKILEAITEEEYLN 292
Query: 497 MQNNVKMVQKHFLWNPKPIRYDLFHMILHSIWLNKLN 533
MQ V V+KHF+ N YD+ HMI+HSIWL +LN
Sbjct: 293 MQRRVLEVRKHFVINRPSKPYDMLHMIMHSIWLRRLN 329
>At5g20260 putative protein
Length = 334
Score = 269 bits (688), Expect = 3e-72
Identities = 145/341 (42%), Positives = 208/341 (60%), Gaps = 18/341 (5%)
Query: 200 METVLKVYIYRDGSRPIFHNPSLKGIYASEGWFMKLMQENKQ-FVTKDPERAHLFYLPYS 258
ME KV++YR+G P+ H + IY+ EG FM ++ F +PE AH F LP S
Sbjct: 1 MEKKFKVWVYREGETPLVHMGPMNNIYSIEGQFMDEIETGMSPFAANNPEEAHAFLLPVS 60
Query: 259 ARQMEVTLYVP-GSHDLKPLSIFLRDYVNKIAAKYPFWNRTHGSDHFLVACHEWGPYTVT 317
+ LY P ++ + L DYV+ +A KYP+WNR+ G+DHF V+CH+W P
Sbjct: 61 VANIVHYLYRPLVTYSREQLHKVFLDYVDVVAHKYPYWNRSLGADHFYVSCHDWAPDVSG 120
Query: 318 EHEELARNTLKALCNADLSERIFIEGRDVSLPETTIRAPRRPLRYLGGNRASL-----RP 372
+ EL +N ++ LCNA+ SE F+ RDVS+PE I P +LG R S RP
Sbjct: 121 SNPELMKNLIRVLCNANTSEG-FMPQRDVSIPEINI-----PGGHLGPPRLSRSSGHDRP 174
Query: 373 ILAFFAGSMHGRVRPTLLKYWGGEKYEDMKIYKPLPLRVSKKMTYIQHMKSSKYCLCPMG 432
ILAFFAG HG +R LL++W +K E++++++ L +K Y + M ++++CLCP G
Sbjct: 175 ILAFFAGGSHGYIRRILLQHWK-DKDEEVQVHEYL----AKNKDYFKLMATARFCLCPSG 229
Query: 433 FEVNSPRIVEAIYYECVPVIIADNFVLPLSEVLDWSAFSVVVAEKDIPRLKDILLSIPMR 492
+EV SPR+V AI CVPVII+D++ LP S+VLDW+ F++ V K IP +K IL SI R
Sbjct: 230 YEVASPRVVAAINLGCVPVIISDHYALPFSDVLDWTKFTIHVPSKKIPEIKTILKSISWR 289
Query: 493 KYVAMQNNVKMVQKHFLWNPKPIRYDLFHMILHSIWLNKLN 533
+Y +Q V VQ+HF+ N +D+ M+LHS+WL +LN
Sbjct: 290 RYRVLQRRVLQVQRHFVINRPSQPFDMLRMLLHSVWLRRLN 330
>At5g33290 unknown protein
Length = 500
Score = 260 bits (664), Expect = 2e-69
Identities = 151/425 (35%), Positives = 229/425 (53%), Gaps = 29/425 (6%)
Query: 131 ANTSTTVTTSFPRGRVPSGKQTDIRLITPTEALVYARKE--IDHVTS-----------VN 177
+NT+ T+ S S Q + +PT + RK +D + S
Sbjct: 78 SNTTNTILASSSSSSSFSDHQNQNKSPSPTSKKIVIRKRSGLDKIESDLAKARAAIKKAA 137
Query: 178 EDPDLYAPLFRNVSVFKRSYELMETVLKVYIYRDGSRPIFHNPSLKGIYASEGWFMKLM- 236
+ + L++N + F +S+ M KV+ Y +G P+FH+ + IY EG FM M
Sbjct: 138 STQNYVSSLYKNPAAFHQSHTEMMNRFKVWTYTEGEVPLFHDGPVNDIYGIEGQFMDEMC 197
Query: 237 ----QENKQFVTKDPERAHLFYLPYSARQMEVTLYVP----GSHDLKPLSIFLRDYVNKI 288
+ +F PE AH+F++P+S ++ +Y P L + DYV+ +
Sbjct: 198 VDGPKSRSRFRADRPENAHVFFIPFSVAKVIHFVYKPITSVEGFSRARLHRLIEDYVDVV 257
Query: 289 AAKYPFWNRTHGSDHFLVACHEWGPYTVTEHEELARNTLKALCNADLSERIFIEGRDVSL 348
A K+P+WNR+ G DHF+V+CH+W P + + +L ++ LCNA+ SE F DVS+
Sbjct: 258 ATKHPYWNRSQGGDHFMVSCHDWAPDVIDGNPKLFEKFIRGLCNANTSEG-FRPNVDVSI 316
Query: 349 PETTIRAPRRPLRYLGGNRASLRPILAFFAGSMHGRVRPTLLKYWGGEKYEDMKIYKPLP 408
PE + + +LG + +R ILAFFAG HG +R L ++W E ++++Y LP
Sbjct: 317 PEIYLPKGKLGPSFLGKS-PRVRSILAFFAGRSHGEIRKILFQHWK-EMDNEVQVYDRLP 374
Query: 409 LRVSKKMTYIQHMKSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLSEVLDWS 468
Y + M SK+CLCP G+EV SPR VEAIY CVPVII+DN+ LP S+VL+W
Sbjct: 375 ----PGKDYTKTMGMSKFCLCPSGWEVASPREVEAIYAGCVPVIISDNYSLPFSDVLNWD 430
Query: 469 AFSVVVAEKDIPRLKDILLSIPMRKYVAMQNNVKMVQKHFLWNPKPIRYDLFHMILHSIW 528
+FS+ + I +K IL S+ + +Y+ M V V++HF+ N YD+ HM+LHSIW
Sbjct: 431 SFSIQIPVSRIKEIKTILQSVSLVRYLKMYKRVLEVKQHFVLNRPAKPYDVMHMMLHSIW 490
Query: 529 LNKLN 533
L +LN
Sbjct: 491 LRRLN 495
>At3g42180 unknown protein
Length = 341
Score = 260 bits (664), Expect = 2e-69
Identities = 142/345 (41%), Positives = 204/345 (58%), Gaps = 20/345 (5%)
Query: 200 METVLKVYIYRDGSRPIFHNPSLKGIYASEGWFMKLMQ-----ENKQFVTKDPERAHLFY 254
M KV+ Y++G +P+ H+ + IY EG F+ + + +F PE AH F+
Sbjct: 1 MMKTFKVWSYKEGEQPLVHDGPVNDIYGIEGQFIDELSYVMGGPSGRFRASRPEEAHAFF 60
Query: 255 LPYSARQMEVTLYVP----GSHDLKPLSIFLRDYVNKIAAKYPFWNRTHGSDHFLVACHE 310
LP+S + +Y P + L DYV+ +A K+PFWN+++G+DHF+V+CH+
Sbjct: 61 LPFSVANIVHYVYQPITSPADFNRARLHRIFNDYVDVVAHKHPFWNQSNGADHFMVSCHD 120
Query: 311 WGPYTVTEHEELARNTLKALCNADLSERIFIEGRDVSLPETTIRAPRRPLR--YLGGNRA 368
W P E +N ++ LCNA+ SE F D S+PE I P+R L+ ++G N
Sbjct: 121 WAPDVPDSKPEFFKNFMRGLCNANTSEG-FRRNIDFSIPEINI--PKRKLKPPFMGQNPE 177
Query: 369 SLRPILAFFAGSMHGRVRPTLLKYWGGEKYEDMKIYKPLPLRVSKKMTYIQHMKSSKYCL 428
+ R ILAFFAG HG +R L +W G K +D+++Y L +K Y + + SK+CL
Sbjct: 178 N-RTILAFFAGRAHGYIREVLFSHWKG-KDKDVQVYDHL----TKGQNYHELIGHSKFCL 231
Query: 429 CPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLSEVLDWSAFSVVVAEKDIPRLKDILLS 488
CP G+EV SPR VEAIY CVPV+I+DN+ LP ++VLDWS FSV + IP +K IL
Sbjct: 232 CPSGYEVASPREVEAIYSGCVPVVISDNYSLPFNDVLDWSKFSVEIPVDKIPDIKKILQE 291
Query: 489 IPMRKYVAMQNNVKMVQKHFLWNPKPIRYDLFHMILHSIWLNKLN 533
IP KY+ M NV V++HF+ N +D+ HMILHS+WL +LN
Sbjct: 292 IPHDKYLRMYRNVMKVRRHFVVNRPAQPFDVIHMILHSVWLRRLN 336
>At4g38040 unknown protein
Length = 425
Score = 243 bits (620), Expect = 2e-64
Identities = 134/389 (34%), Positives = 219/389 (55%), Gaps = 28/389 (7%)
Query: 154 IRLITPTEALVYARKEIDHVTSVNE----------DPDLYAPLFRNVSVFKRSYELMETV 203
+R P+ +V + H T VNE + + Y+ ++ + F+ +Y ME
Sbjct: 45 LRTSPPSPVIVVTPIHVPH-TFVNEYKTDNETPTMEEETYSDVYHSPEAFRLNYAEMEKR 103
Query: 204 LKVYIYRDGSRPIFHNPSLK--GIYASEGWFMKLMQENKQFVTKDPERAHLFYLPYSARQ 261
KVYIY DG F+ K G YASEG+F + ++E++ F T DP+ A LF++P S +
Sbjct: 104 FKVYIYPDGDPNTFYQTPRKVTGKYASEGYFFQNIRESR-FRTLDPDEADLFFIPISCHK 162
Query: 262 MEVTLYVPGSHDLKPLSIFLRDYVNKIAAKYPFWNRTHGSDHFLVACHEWGPYTVTEHEE 321
M + +++ +++YV+ + AKYP+WNRT G+DHF V CH+ G
Sbjct: 163 MRGK-----GTSYENMTVIVQNYVDGLIAKYPYWNRTLGADHFFVTCHDVGVRAFEGSPL 217
Query: 322 LARNTLKALCNADLSERIFIEGRDVSLPETTIRAPRRPLRY-LGGNRASLRPILAFFAGS 380
L +NT++ +C+ + FI +DV+LP+ +P GGN R L F+AG
Sbjct: 218 LIKNTIRVVCSPSYNVG-FIPHKDVALPQVL-----QPFALPAGGNDVENRTTLGFWAGH 271
Query: 381 MHGRVRPTLLKYWGGEKYEDMKIYKPLPLRVSKKMTYIQHMKSSKYCLCPMGFEVNSPRI 440
+ ++R L W E ++ I R + + Y + +K+C+CP G +VNS RI
Sbjct: 272 RNSKIRVILAHVW--ENDTELDISNNRINRATGHLVYQKRFYRTKFCICPGGSQVNSARI 329
Query: 441 VEAIYYECVPVIIADNFVLPLSEVLDWSAFSVVVAEKDIPRLKDILLSIPMRKYVAMQNN 500
++I+Y C+PVI++D + LP +++L+W F+VV+ E+D+ LK IL +IP ++V++ NN
Sbjct: 330 TDSIHYGCIPVILSDYYDLPFNDILNWRKFAVVLREQDVYNLKQILKNIPHSEFVSLHNN 389
Query: 501 VKMVQKHFLWNPKPIRYDLFHMILHSIWL 529
+ VQKHF WN P+++D FHMI++ +WL
Sbjct: 390 LVKVQKHFQWNSPPVKFDAFHMIMYELWL 418
>At1g27440 unknown protein
Length = 412
Score = 125 bits (314), Expect = 6e-29
Identities = 102/350 (29%), Positives = 161/350 (45%), Gaps = 41/350 (11%)
Query: 204 LKVYIY----RDGSRPIFHNPS-LKGIYASEGWFMKLMQENKQFVTKDPERAHLFYLPYS 258
LKVY+Y + + + +P L ++A+E FM + T++P+ A FY P
Sbjct: 46 LKVYVYELPSKYNKKLLQKDPRCLTHMFAAE-IFMHRFLLSSPVRTRNPDEADWFYTP-- 102
Query: 259 ARQMEVTLYVPGSHDLKPLSI--------FLRDYVNKIAAKYPFWNRTHGSDHFLVACHE 310
+ + DL P + +R + I++ +P+WNRT G+DHF V H+
Sbjct: 103 ---------IYPTCDLTPTGLPLPFKSPRMMRSSIQLISSNWPYWNRTEGADHFFVVPHD 153
Query: 311 WGP-YTVTEHEELARNTLKALCNADLSERIFIEGRDVSLPETTIRAPRRPLRYLGGNRAS 369
+G + E + + R L L A L + F + V L E +I P P +A
Sbjct: 154 FGACFHYQEEKAIERGILPLLQRATLVQT-FGQRNHVCLDEGSITIP--PFAPPQKMQAH 210
Query: 370 LRP------ILAFFAGSMHGRVRPTLLKYWG----GEKYEDMKIYKPLPLRVSKKMTYIQ 419
P I +F G + Y+ +E+ K + TY +
Sbjct: 211 FIPPDIPRSIFVYFRGLFYDVNNDPEGGYYARGARAAVWENFKNNPLFDISTDHPTTYYE 270
Query: 420 HMKSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLSEVLDWSAFSVVVAEKDI 479
M+ + +CLCP+G+ SPR+VEA+ + C+PVIIAD+ VLP ++ + W V VAEKD+
Sbjct: 271 DMQRAIFCLCPLGWAPWSPRLVEAVVFGCIPVIIADDIVLPFADAIPWEEIGVFVAEKDV 330
Query: 480 PRLKDILLSIPMRKYVAMQNNV-KMVQKHFLWNPKPIR-YDLFHMILHSI 527
P L IL SIP + Q + K + P+P + D FH IL+ +
Sbjct: 331 PELDTILTSIPTEVILRKQRLLANPSMKRAMLFPQPAQPGDAFHQILNGL 380
>At5g61840 unknown protein
Length = 415
Score = 123 bits (309), Expect = 2e-28
Identities = 99/348 (28%), Positives = 163/348 (46%), Gaps = 37/348 (10%)
Query: 204 LKVYIY----RDGSRPIFHNPS-LKGIYASEGWFMKLMQENKQFVTKDPERAHLFYLPYS 258
LKV++Y + + + +P L ++A+E + + + + T +PE A FY+P
Sbjct: 49 LKVFVYELPSKYNKKILQKDPRCLNHMFAAEIYMQRFLLSSP-VRTLNPEEADWFYVP-- 105
Query: 259 ARQMEVTLYVPGSHDLKPLSI--------FLRDYVNKIAAKYPFWNRTHGSDHFLVACHE 310
V + DL P + +R + IA+ +P+WNRT G+DHF V H+
Sbjct: 106 ---------VYTTCDLTPNGLPLPFKSPRMMRSAIQLIASNWPYWNRTEGADHFFVVPHD 156
Query: 311 WGP-YTVTEHEELARNTLKALCNADLSERIFIEGRDVSLPETTIRAPR-RPLRYLGGN-- 366
+G + E + + R L L A L + F + V L E +I P P + + +
Sbjct: 157 FGACFHYQEEKAIGRGILPLLQRATLVQT-FGQRNHVCLKEGSITVPPYAPPQKMQSHLI 215
Query: 367 -RASLRPILAFFAGSMHGRVRPTLLKYWG----GEKYEDMKIYKPLPLRVSKKMTYIQHM 421
+ R I +F G + Y+ +E+ K + TY + M
Sbjct: 216 PEKTPRSIFVYFRGLFYDVGNDPEGGYYARGARAAVWENFKDNPLFDISTEHPTTYYEDM 275
Query: 422 KSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLSEVLDWSAFSVVVAEKDIPR 481
+ + +CLCP+G+ SPR+VEA+ + C+PVIIAD+ VLP ++ + W V V EKD+P
Sbjct: 276 QRAIFCLCPLGWAPWSPRLVEAVIFGCIPVIIADDIVLPFADAIPWEDIGVFVDEKDVPY 335
Query: 482 LKDILLSIPMRKYVAMQNNV-KMVQKHFLWNPKPIR-YDLFHMILHSI 527
L IL SIP + Q + K + P+P + D FH +L+ +
Sbjct: 336 LDTILTSIPPEVILRKQRLLANPSMKQAMLFPQPAQPGDAFHQVLNGL 383
>At5g22940 putative protein
Length = 498
Score = 108 bits (269), Expect = 1e-23
Identities = 89/309 (28%), Positives = 139/309 (44%), Gaps = 37/309 (11%)
Query: 244 TKDPERAHLFYLP-YSARQMEVTLYVPGSHDLKPLSIFLRDYVNKIAAKYPFWNRTHGSD 302
T DP+ A F++P Y + + P + L L V+ ++ YPFWNR+ GSD
Sbjct: 182 TLDPDEADYFFVPVYVSCNFSTSNGFPSLSHARSL---LSSAVDFLSDHYPFWNRSQGSD 238
Query: 303 HFLVACHEWGP-YTVTEHEELARNTLKALCNADLSERIFIEGRD--------VSLPETTI 353
H VA H++G + E + K + + + + ++ + V P
Sbjct: 239 HVFVASHDFGACFHAMEDMAIEEGIPKFMKRSIILQTFGVKYKHPCQEVEHVVIPPYIPP 298
Query: 354 RAPRRPLRYLGGNRASLRPILAFFAGSMH-------GR-----VRPTLLKYWGGEKYEDM 401
+ ++ + N R I AFF G M GR VR +LK +GG +
Sbjct: 299 ESVQKAIEKAPVN--GRRDIWAFFRGKMEVNPKNISGRFYSKGVRTAILKKFGGRR---- 352
Query: 402 KIYKPLPLRVSKKMTYIQHMKSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPL 461
+ Y L + Y + S +CLCP+G+ SPR+VE+ CVPV+IAD LP
Sbjct: 353 RFY----LNRHRFAGYRSEIVRSVFCLCPLGWAPWSPRLVESAVLGCVPVVIADGIQLPF 408
Query: 462 SEVLDWSAFSVVVAEKDIPRLKDILLSIPMRKYVAMQNNV--KMVQKHFLWNPKPIRYDL 519
SE + W S+ VAEKD+ L+ +L + A+Q N+ + ++ L+N D
Sbjct: 409 SETVQWPEISLTVAEKDVRNLRKVLEHVAATNLSAIQRNLHEPVFKRALLYNVPMKEGDA 468
Query: 520 FHMILHSIW 528
IL S+W
Sbjct: 469 TWHILESLW 477
>At2g28110 unknown protein
Length = 448
Score = 106 bits (264), Expect = 4e-23
Identities = 96/360 (26%), Positives = 156/360 (42%), Gaps = 43/360 (11%)
Query: 204 LKVYIYRDGSRPIFHNPSLKGIYASEGWFMKLMQENKQFV-------TKDPERAHLFYLP 256
LK+Y+Y S+ F+ L + F + +K F+ T+DP A F++P
Sbjct: 94 LKIYVYDLPSK--FNKDWLANDRCTNHLFAAEVALHKAFLSLEGDVRTEDPYEADFFFVP 151
Query: 257 -YSARQMEVTLYVPGSHDLKPLSIFLRDYVNKIAAKYPFWNRTHGSDHFLVACHEWGP-Y 314
Y + P + L + D + ++ +YPFWNRT GSDH A H++G +
Sbjct: 152 VYVSCNFSTINGFPAIGHARSL---INDAIKLVSTQYPFWNRTSGSDHVFTATHDFGSCF 208
Query: 315 TVTEHEELARNTLKALCNADLSERIFIE-GRDVSLPETTIRAPRRPLRYLGGNRASL--- 370
E +A L N+ + + + E + P L + ++
Sbjct: 209 HTMEDRAIADGVPIFLRNSIILQTFGVTFNHPCQEVENVVIPPYISPESLHKTQKNIPVT 268
Query: 371 --RPILAFFAGSMH------------GRVRPTLLKYWGGEKYEDMKIYKPLPLRVSKKMT 416
R I FF G M RVR + + +GG D + Y L+ +
Sbjct: 269 KERDIWVFFRGKMELHPKNISGRFYSKRVRTNIWRSYGG----DRRFY----LQRQRFAG 320
Query: 417 YIQHMKSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLSEVLDWSAFSVVVAE 476
Y + S +CLCP+G+ SPR+VE++ CVPVIIAD LP + W S+ VAE
Sbjct: 321 YQSEIARSVFCLCPLGWAPWSPRLVESVALGCVPVIIADGIRLPFPSTVRWPDISLTVAE 380
Query: 477 KDIPRLKDILLSIPMRKYVAMQNNVK--MVQKHFLWNPKPIRYDLFHMILHSIWLNKLNQ 534
+D+ +L DIL + +Q N++ V++ ++N D +L ++ KLN+
Sbjct: 381 RDVGKLGDILEHVAATNLSVIQRNLEDPSVRRALMFNVPSREGDATWQVLEAL-SKKLNR 439
>At5g16890 unknown protein (At5g16890)
Length = 511
Score = 72.4 bits (176), Expect = 6e-13
Identities = 64/231 (27%), Positives = 106/231 (45%), Gaps = 20/231 (8%)
Query: 293 PFWNRTHGSDHFLVACHEWGPYTVTEHEELARNTLKALCNADLSERIFIEGRDVSLPETT 352
P W R+ G DH H W +V + +N + L + D + + G+ VSL +
Sbjct: 228 PAWKRSEGRDHIFPIHHPWSFKSV---RKFVKNAIWLLPDMDSTGNWYKPGQ-VSLEKDL 283
Query: 353 IRAPRRPLRYLGGNR-----ASLRPILAFFAGSMH----GRVRPTLLKYWGGEKYEDMKI 403
I P P + + A +R L FF G + G++R L G K D+ I
Sbjct: 284 I-LPYVPNVDICDTKCLSESAPMRTTLLFFRGRLKRNAGGKIRAKLGAELSGIK--DIII 340
Query: 404 YKPLPLRVSKKMTYIQHMKSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLSE 463
+ K+ + M+ S +CLCP G +S R+ +AI C+PVI++D P
Sbjct: 341 SEGTAGE-GGKLAAQRGMRRSLFCLCPAGDTPSSARLFDAIVSGCIPVIVSDELEFPFEG 399
Query: 464 VLDWSAFSVVVAEKDIPR---LKDILLSIPMRKYVAMQNNVKMVQKHFLWN 511
+LD+ +V+V+ D + L + L S+ + +QNN+ +HFL++
Sbjct: 400 ILDYKKVAVLVSSSDAIQPGWLVNHLRSLTPFQVKGLQNNLAQYSRHFLYS 450
>At1g21480 unknown protein
Length = 462
Score = 72.4 bits (176), Expect = 6e-13
Identities = 83/339 (24%), Positives = 146/339 (42%), Gaps = 49/339 (14%)
Query: 204 LKVYIY--------------RDGSRPIFHNPSLKGIYASEGWFMKLMQENKQFVTKDPER 249
LK+Y+Y RDGS + LKG + S+ KL+ E+K F T +
Sbjct: 89 LKIYVYDENEIDGLKELLYGRDGS--VKTTACLKGQWGSQVKIHKLLLESK-FRTIKKDE 145
Query: 250 AHLFYLPYSARQMEVTLYVPGSHDLKPLSIFLRDYVNKIAAKYPFWNRTHGSDHFLVACH 309
A LF++P + + + G + K ++ + YV K+ ++ P++ R+ G DH V
Sbjct: 146 ADLFFVPAYVKCVRML----GGLNDKEIN---QTYV-KVLSQMPYFRRSGGRDHIFVFPS 197
Query: 310 EWGPYTVTEHEELARNTLKALCNADLSER----IFIEGRDVSLPETTIRAPRR------- 358
G + ++ AD +++ F +D+ +P A +
Sbjct: 198 GAGAHLFRSWSTFINRSIILTPEADRTDKKDTTAFNSWKDIIIPGNVDDAMTKNGQPDVQ 257
Query: 359 PLRYLGGNRASLRPILAFFAGSMHGRV-RPTLLKYWGGEKYEDMKIYKPLPLRVSKKM-- 415
PL S R LA + G G+ R L+ +++ D L ++K
Sbjct: 258 PLPL------SKRKYLANYLGRAQGKAGRLKLIDL--SKQFPDKLECPDLKFSGTEKFGR 309
Query: 416 -TYIQHMKSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLSEVLDWSAFSVVV 474
TY +H++++K+CL P G + R E+ + ECVPV+++D+ LP V+D++ S+
Sbjct: 310 TTYFEHLRNAKFCLAPRGESSWTLRFYESFFVECVPVLLSDHAELPFQNVIDYAQVSIKW 369
Query: 475 AEKDI-PRLKDILLSIPMRKYVAMQNNVKMVQKHFLWNP 512
I D L SI R M + ++ F++ P
Sbjct: 370 PSTRIGSEFLDYLASISDRDIEGMIARGRKIRCLFVYGP 408
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.137 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,899,140
Number of Sequences: 26719
Number of extensions: 503209
Number of successful extensions: 1363
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 28
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 1248
Number of HSP's gapped (non-prelim): 62
length of query: 535
length of database: 11,318,596
effective HSP length: 104
effective length of query: 431
effective length of database: 8,539,820
effective search space: 3680662420
effective search space used: 3680662420
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 62 (28.5 bits)
Medicago: description of AC119408.6


