

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
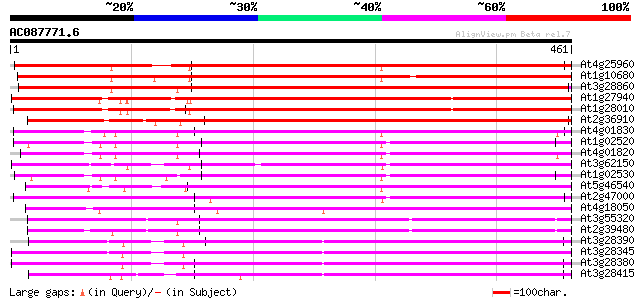
Query= AC087771.6 + phase: 0 /pseudo/partial
(461 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At4g25960 P-glycoprotein-2 (pgp2) 604 e-173
At1g10680 putative P-glycoprotein-2 emb|CAA71277 563 e-161
At3g28860 P-glycoprotein, putative 393 e-109
At1g27940 hypothetical protein 380 e-105
At1g28010 hypothetical protein 371 e-103
At2g36910 putative ABC transporter 355 4e-98
At4g01830 putative P-glycoprotein-like protein 313 1e-85
At1g02520 P-glycoprotein, putative 311 5e-85
At4g01820 P-glycoprotein-like protein pgp3 304 8e-83
At3g62150 P-glycoprotein-like proetin 304 8e-83
At1g02530 hypothetical protein 300 1e-81
At5g46540 multidrug resistance p-glycoprotein 292 3e-79
At2g47000 putative ABC transporter 290 1e-78
At4g18050 multidrug resistance protein/P-glycoprotein - like 285 4e-77
At3g55320 P-glycoprotein - like 283 2e-76
At2g39480 putative ABC transporter 282 3e-76
At3g28390 P-glycoprotein, putative 280 9e-76
At3g28345 P-glycoprotein, putative, 3'partial 280 9e-76
At3g28380 P-glycoprotein, putative 273 1e-73
At3g28415 putative protein 271 7e-73
>At4g25960 P-glycoprotein-2 (pgp2)
Length = 1233
Score = 604 bits (1558), Expect = e-173
Identities = 323/480 (67%), Positives = 367/480 (76%), Gaps = 38/480 (7%)
Query: 5 EGDERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAY 64
E ++ + KVS+LKLFSFAD YD VLM +GS+GA +HGASVPIFFIFFGKLIN+IGLAY
Sbjct: 10 EKEKEMTQPKVSLLKLFSFADFYDCVLMTLGSVGACIHGASVPIFFIFFGKLINIIGLAY 69
Query: 65 LFPKEASHKVAKSKCSNI---IFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYKFI*Y 121
LFPK+ASH+VAK + + IL L A W T + + + Y++S
Sbjct: 70 LFPKQASHRVAKYSLDFVYLSVAILFSSWLEVACWMHTGERQAAKMRRAYLRSMLS---- 125
Query: 122 *GIHWRGHFCYYK*HYYCSRCSIRE--------------------GNFLHYISRFIAGFT 161
+ + S E GNFLHYISRFIAGF
Sbjct: 126 -----------QDISLFDTEASTGEVISAITSDILVVQDALSEKVGNFLHYISRFIAGFA 174
Query: 162 IGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQ 221
IGF VWQISLVTLSIVP IALAGG YA+V IGLIA+VRK+Y++AGEIAEEVIGNVRTVQ
Sbjct: 175 IGFTSVWQISLVTLSIVPLIALAGGIYAFVAIGLIARVRKSYIKAGEIAEEVIGNVRTVQ 234
Query: 222 AFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNI 281
AF GEERAVR Y+ AL TY GRKAGL KGLGLGSMHCVLFLSWALLVW+TSVVVHK+I
Sbjct: 235 AFTGEERAVRLYREALENTYKYGRKAGLTKGLGLGSMHCVLFLSWALLVWFTSVVVHKDI 294
Query: 282 ANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKL 341
A+GG+SFTTMLNVVI+GLSLGQAAPDISAF+RAKAAAYPIF+MIER+TV+K S+K+GRKL
Sbjct: 295 ADGGKSFTTMLNVVIAGLSLGQAAPDISAFVRAKAAAYPIFKMIERNTVTKTSAKSGRKL 354
Query: 342 SKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYE 401
K+DGHIQF D FSYPSRPDV IF LNL IPAGKIVALVGGSGSGKSTV+SLIERFYE
Sbjct: 355 GKVDGHIQFKDATFSYPSRPDVVIFDRLNLAIPAGKIVALVGGSGSGKSTVISLIERFYE 414
Query: 402 PISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
PISG +LLD N+I ELD+KWLR QIGLVNQEPALFAT+I+ENILYGKDDAT EE+ RA K
Sbjct: 415 PISGAVLLDGNNISELDIKWLRGQIGLVNQEPALFATTIRENILYGKDDATAEEITRAAK 474
Score = 209 bits (533), Expect = 2e-54
Identities = 116/310 (37%), Positives = 184/310 (58%), Gaps = 8/310 (2%)
Query: 150 LHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEI 209
L + + F I F+ W+++LV L+ P + G + KAY++A +
Sbjct: 794 LQNLGLVVTSFIIAFILNWRLTLVVLATYPLVISGHISEKLFMQGYGGDLNKAYLKANML 853
Query: 210 AEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALL 269
A E + N+RTV AF EE+ + Y L++ + + G GL G +F S+ L
Sbjct: 854 AGESVSNIRTVAAFCAEEKILELYSRELLEPSKSSFRRGQIAGLFYGVSQFFIFSSYGLA 913
Query: 270 VWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQA---APDISAFIRAKAAAYPIFEMIE 326
+WY S ++ K +A T + ++++ L++G+ APD+ ++ +FE+++
Sbjct: 914 LWYGSTLMDKGLAGFKSVMKTFMVLIVTALAMGETLALAPDL---LKGNQMVASVFEILD 970
Query: 327 RDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSG 386
R T + +T +L+ ++G I+ V FSYPSRPDV IF + +L + AGK +ALVG SG
Sbjct: 971 RKT--QIVGETSEELNNVEGTIELKGVHFSYPSRPDVVIFRDFDLIVRAGKSMALVGQSG 1028
Query: 387 SGKSTVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILY 446
SGKS+V+SLI RFY+P +G+++++ DI++LDLK LR+ IGLV QEPALFAT+I ENILY
Sbjct: 1029 SGKSSVISLILRFYDPTAGKVMIEGKDIKKLDLKALRKHIGLVQQEPALFATTIYENILY 1088
Query: 447 GKDDATLEEL 456
G + A+ E+
Sbjct: 1089 GNEGASQSEV 1098
>At1g10680 putative P-glycoprotein-2 emb|CAA71277
Length = 1227
Score = 563 bits (1450), Expect = e-161
Identities = 304/463 (65%), Positives = 349/463 (74%), Gaps = 12/463 (2%)
Query: 7 DERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLF 66
++ KK VS LKLFSFAD YD VLM +GSIGA +HGASVP+FFIFFGKLIN+IGLAYLF
Sbjct: 16 EKEKKRPSVSFLKLFSFADFYDCVLMALGSIGACIHGASVPVFFIFFGKLINIIGLAYLF 75
Query: 67 PKEASHKVAKSKCSNI---IFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYK---FI* 120
P+EASHKVAK + + IL L A W T + + K Y++S +
Sbjct: 76 PQEASHKVAKYSLDFVYLSVVILFSSWLEVACWMHTGERQAAKIRKAYLRSMLSQDISLF 135
Query: 121 Y*GIHWRGHFCYYK*HYYCSRCSIRE--GNFLHYISRFIAGFTIGFVRVWQISLVTLSIV 178
I + +I E GNF+H+ISRFIAGF IGF VWQISLVTLSIV
Sbjct: 136 DTEISTGEVISAITSEILVVQDAISEKVGNFMHFISRFIAGFAIGFASVWQISLVTLSIV 195
Query: 179 PAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALM 238
P IALAGG YA+V+ GLI +VRK+YV+A EIAEEVIGNVRTVQAF GEE+AV SY+ AL
Sbjct: 196 PFIALAGGIYAFVSSGLIVRVRKSYVKANEIAEEVIGNVRTVQAFTGEEKAVSSYQGALR 255
Query: 239 KTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVISG 298
TY GRKAGLAKGLGLGS+H VLFLSWALL+W+TS+VVHK IANGGESFTTMLNVVI+G
Sbjct: 256 NTYNYGRKAGLAKGLGLGSLHFVLFLSWALLIWFTSIVVHKGIANGGESFTTMLNVVIAG 315
Query: 299 LSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYP 358
LSLGQAAPDIS F+RA AAAYPIF+MIER+T KTGRKL ++G I F DV F+YP
Sbjct: 316 LSLGQAAPDISTFMRASAAAYPIFQMIERNT----EDKTGRKLGNVNGDILFKDVTFTYP 371
Query: 359 SRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELD 418
SRPDV IF LN IPAGK+VALVGGSGSGKST++SLIERFYEP G ++LD NDIR LD
Sbjct: 372 SRPDVVIFDKLNFVIPAGKVVALVGGSGSGKSTMISLIERFYEPTDGAVMLDGNDIRYLD 431
Query: 419 LKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
LKWLR IGLVNQEP LFAT+I+ENI+YGKDDAT EE+ A K
Sbjct: 432 LKWLRGHIGLVNQEPVLFATTIRENIMYGKDDATSEEITNAAK 474
Score = 217 bits (553), Expect = 9e-57
Identities = 122/315 (38%), Positives = 187/315 (58%), Gaps = 8/315 (2%)
Query: 150 LHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEI 209
L + + F I F+ W+++LV L+ P I G + KAY++A +
Sbjct: 786 LENLGLVVTAFIISFILNWRLTLVVLATYPLIISGHISEKIFMQGYGGNLSKAYLKANML 845
Query: 210 AEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALL 269
A E I N+RTV AF EE+ + Y L++ + G G+ G +F S+ L
Sbjct: 846 AGESISNIRTVVAFCAEEKVLDLYSKELLEPSERSFRRGQMAGILYGVSQFFIFSSYGLA 905
Query: 270 VWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQA---APDISAFIRAKAAAYPIFEMIE 326
+WY S+++ K +++ T + ++++ L +G+ APD+ ++ +FE+++
Sbjct: 906 LWYGSILMEKGLSSFESVMKTFMVLIVTALVMGEVLALAPDL---LKGNQMVVSVFELLD 962
Query: 327 RDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSG 386
R T + TG +LS ++G I+ V FSYPSRPDV IF++ NL +P+GK +ALVG SG
Sbjct: 963 RRT--QVVGDTGEELSNVEGTIELKGVHFSYPSRPDVTIFSDFNLLVPSGKSMALVGQSG 1020
Query: 387 SGKSTVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILY 446
SGKS+V+SL+ RFY+P +G I++D DI++L LK LR+ IGLV QEPALFAT+I ENILY
Sbjct: 1021 SGKSSVLSLVLRFYDPTAGIIMIDGQDIKKLKLKSLRRHIGLVQQEPALFATTIYENILY 1080
Query: 447 GKDDATLEELKRAVK 461
GK+ A+ E+ A K
Sbjct: 1081 GKEGASESEVMEAAK 1095
>At3g28860 P-glycoprotein, putative
Length = 1252
Score = 393 bits (1009), Expect = e-109
Identities = 205/460 (44%), Positives = 298/460 (64%), Gaps = 8/460 (1%)
Query: 8 ERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLFP 67
E+KKE + KLFSFAD +DY+LMF+GS+GAIVHG+S+P+FF+ FG+++N G +
Sbjct: 17 EKKKEQSLPFFKLFSFADKFDYLLMFVGSLGAIVHGSSMPVFFLLFGQMVNGFGKNQMDL 76
Query: 68 KEASHKVAKSKCSNI---IFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYKF-I*Y*G 123
+ H+V++ + + + A W + + L K Y+++ K + +
Sbjct: 77 HQMVHEVSRYSLYFVYLGLVVCFSSYAEIACWMYSGERQVAALRKKYLEAVLKQDVGFFD 136
Query: 124 IHWRGHFCYYK*H----YYCSRCSIREGNFLHYISRFIAGFTIGFVRVWQISLVTLSIVP 179
R + S + GNF+HY+S F+AG +GFV W+++L++++++P
Sbjct: 137 TDARTGDIVFSVSTDTLLVQDAISEKVGNFIHYLSTFLAGLVVGFVSAWKLALLSVAVIP 196
Query: 180 AIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALMK 239
IA AGG YAY G+ +K R++Y AG IAE+ I VRTV ++ GE +A+ +Y A+
Sbjct: 197 GIAFAGGLYAYTLTGITSKSRESYANAGVIAEQAIAQVRTVYSYVGESKALNAYSDAIQY 256
Query: 240 TYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVISGL 299
T G KAG+AKGLGLG + + +SWAL+ WY V + +GG++FT + + ++ G+
Sbjct: 257 TLKLGYKAGMAKGLGLGCTYGIACMSWALVFWYAGVFIRNGQTDGGKAFTAIFSAIVGGM 316
Query: 300 SLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPS 359
SLGQ+ ++ AF + KAA Y + E+I + + G+ L ++ G+I+F DV FSYPS
Sbjct: 317 SLGQSFSNLGAFSKGKAAGYKLMEIINQRPTIIQDPLDGKCLDQVHGNIEFKDVTFSYPS 376
Query: 360 RPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELDL 419
RPDV IF N N+ P+GK VA+VGGSGSGKSTVVSLIERFY+P SGQILLD +I+ L L
Sbjct: 377 RPDVMIFRNFNIFFPSGKTVAVVGGSGSGKSTVVSLIERFYDPNSGQILLDGVEIKTLQL 436
Query: 420 KWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRA 459
K+LR+QIGLVNQEPALFAT+I ENILYGK DAT+ E++ A
Sbjct: 437 KFLREQIGLVNQEPALFATTILENILYGKPDATMVEVEAA 476
Score = 192 bits (488), Expect = 3e-49
Identities = 109/312 (34%), Positives = 166/312 (52%)
Query: 150 LHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEI 209
L ++ + F + F+ W++SL+ L P + LA G KA+ + I
Sbjct: 812 LQNMTSLLTSFIVAFIVEWRVSLLILGTFPLLVLANFAQQLSLKGFAGDTAKAHAKTSMI 871
Query: 210 AEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALL 269
A E + N+RTV AF + + + + L G G L+ S AL+
Sbjct: 872 AGEGVSNIRTVAAFNAQSKILSLFCHELRVPQKRSLYRSQTSGFLFGLSQLALYGSEALI 931
Query: 270 VWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDT 329
+WY + +V K ++ + + +VI+ S+ + IR A +F +++R T
Sbjct: 932 LWYGAHLVSKGVSTFSKVIKVFVVLVITANSVAETVSLAPEIIRGGEAVGSVFSVLDRQT 991
Query: 330 VSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGK 389
+ + G I+F V F+YPSRPDV +F + NL I AG ALVG SGSGK
Sbjct: 992 RIDPDDADADPVETIRGDIEFRHVDFAYPSRPDVMVFRDFNLRIRAGHSQALVGASGSGK 1051
Query: 390 STVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKD 449
S+V+++IERFY+P++G++++D DIR L+LK LR +IGLV QEPALFA +I +NI YGKD
Sbjct: 1052 SSVIAMIERFYDPLAGKVMIDGKDIRRLNLKSLRLKIGLVQQEPALFAATIFDNIAYGKD 1111
Query: 450 DATLEELKRAVK 461
AT E+ A +
Sbjct: 1112 GATESEVIDAAR 1123
>At1g27940 hypothetical protein
Length = 1245
Score = 380 bits (975), Expect = e-105
Identities = 210/475 (44%), Positives = 305/475 (64%), Gaps = 22/475 (4%)
Query: 2 ESKEGDERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIG 61
E+KE + K+ VS++ LFS AD DY LM +G +GA +HGA++P+FF+FFGK+++ +G
Sbjct: 17 EAKEEKKNIKKESVSLMGLFSAADKLDYFLMLLGGLGACIHGATLPLFFVFFGKMLDSLG 76
Query: 62 LAYLFPKEASHKVAKSKCSNIIFILDRG--GLLDAY-----WRKTSSKDENGLFKVYVKS 114
PK S +V++ N ++++ G + A+ W +T + L Y+KS
Sbjct: 77 NLSTDPKAISSRVSQ----NALYLVYLGLVNFVSAWIGVSCWMQTGERQTARLRINYLKS 132
Query: 115 RY-KFI*Y*GIHWRGHFCYYK*HYYCSRCSIREG------NFLHYISRFIAGFTIGFVRV 167
K I + R + H +++ + L Y+S+FIAGF IGF+ V
Sbjct: 133 ILAKDITFFDTEARDSNLIF--HISSDAILVQDAIGDKTDHVLRYLSQFIAGFVIGFLSV 190
Query: 168 WQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEE 227
WQ++L+TL +VP IA+AGG YA V + K AY AG++AEEV+ VRTV AF GEE
Sbjct: 191 WQLTLLTLGVVPLIAIAGGGYAIVMSTISEKSETAYADAGKVAEEVMSQVRTVYAFVGEE 250
Query: 228 RAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGES 287
+AV+SY +L K G+++GLAKGLG+G + +LF +WALL+WY S++V NG ++
Sbjct: 251 KAVKSYSNSLKKALKLGKRSGLAKGLGVGLTYSLLFCAWALLLWYASLLVRHGKTNGAKA 310
Query: 288 FTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMI-ERDTVSKKSSKTGRKLSKLDG 346
FTT+LNV+ SG +LGQAAP +SA + + AA IF MI ++ S + G L + G
Sbjct: 311 FTTILNVIFSGFALGQAAPSLSAIAKGRVAAANIFRMIGNNNSESSQRLDEGTTLQNVAG 370
Query: 347 HIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQ 406
I+F V F+YPSRP++ +F NL+ I +GK A VG SGSGKST++S+++RFYEP SG+
Sbjct: 371 RIEFQKVSFAYPSRPNM-VFENLSFTIRSGKTFAFVGPSGSGKSTIISMVQRFYEPNSGE 429
Query: 407 ILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
ILLD NDI+ L LKW R+Q+GLV+QEPALFAT+I NIL GK++A ++++ A K
Sbjct: 430 ILLDGNDIKSLKLKWFREQLGLVSQEPALFATTIASNILLGKENANMDQIIEAAK 484
Score = 207 bits (528), Expect = 7e-54
Identities = 136/476 (28%), Positives = 227/476 (47%), Gaps = 25/476 (5%)
Query: 2 ESKEGDERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIG 61
++K D +K SM+ +S ++ +GSIGA++ GA P+F + G
Sbjct: 651 KTKNDDSKKDFSSSSMIWELIKLNSPEWPYALLGSIGAVLAGAQTPLFSM---------G 701
Query: 62 LAYLFPKEASH--KVAKSKCSNIIFILDRGGLLDA------------YWRKTSSKDENGL 107
+AY+ S V K + I G++ A + +S+ L
Sbjct: 702 IAYVLTAFYSPFPNVIKRDVEKVAIIFAGAGIVTAPIYLLQHYFYTLMGERLTSRVRLSL 761
Query: 108 FKVYVKSRYKFI*Y*GIHWRGHFCYYK*HYYCSRCSI--REGNFLHYISRFIAGFTIGFV 165
F + + + + R ++ R + +S + + F
Sbjct: 762 FSAILSNEIGWFDLDENNTGSLTSILAADATLVRSALADRLSTIVQNLSLTVTALALAFF 821
Query: 166 RVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAG 225
W+++ V + P + A G +AY RA +A E I N+RTV A+
Sbjct: 822 YSWRVAAVVTACFPLLIAASLTEQLFLKGFGGDYTRAYSRATSVAREAIANIRTVAAYGA 881
Query: 226 EERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGG 285
E++ + L K N G G G G + F S+AL +WY SV+++ N G
Sbjct: 882 EKQISEQFTCELSKPTKNAFVRGHISGFGYGLSQFLAFCSYALGLWYVSVLINHKETNFG 941
Query: 286 ESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLD 345
+S + + ++++ S+ + ++ A +F ++ R+T R +S++
Sbjct: 942 DSIKSFMVLIVTAFSVSETLALTPDIVKGTQALGSVFRVLHRETKISPDQPNSRMVSQVK 1001
Query: 346 GHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISG 405
G I+F +V F YP+RP++ IF NLNL + AGK +A+VG SGSGKSTV+ LI RFY+P +G
Sbjct: 1002 GDIEFRNVSFVYPTRPEIDIFKNLNLRVSAGKSLAVVGPSGSGKSTVIGLIMRFYDPSNG 1061
Query: 406 QILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
+ +D DI+ L+L+ LR+++ LV QEPALF+T+I ENI YG ++A+ E+ A K
Sbjct: 1062 NLCIDGQDIKTLNLRSLRKKLALVQQEPALFSTTIYENIKYGNENASEAEIMEAAK 1117
>At1g28010 hypothetical protein
Length = 1247
Score = 371 bits (952), Expect = e-103
Identities = 207/474 (43%), Positives = 304/474 (63%), Gaps = 24/474 (5%)
Query: 4 KEGDERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLA 63
KE ++ K+ VS++ LFS AD+ DY LMF+G +G +HG ++P+FF+FFG +++ +G
Sbjct: 20 KEEKKKMKKESVSLMGLFSAADNVDYFLMFLGGLGTCIHGGTLPLFFVFFGGMLDSLGKL 79
Query: 64 YLFPKEASHKVAKSKCSNIIFILDRG--GLLDAY-----WRKTSSKDENGLFKVYVKSRY 116
P S +V++ N ++++ G L+ A+ W +T + L Y+KS
Sbjct: 80 STDPNAISSRVSQ----NALYLVYLGLVNLVSAWIGVACWMQTGERQTARLRINYLKSIL 135
Query: 117 -KFI*Y*GIHWR-GHFCYYK*HYYCSRCSIRE------GNFLHYISRFIAGFTIGFVRVW 168
K I + R +F + H +++ G+ L Y+ +FIAGF IGF+ VW
Sbjct: 136 AKDITFFDTEARDSNFIF---HISSDAILVQDAIGDKTGHVLRYLCQFIAGFVIGFLSVW 192
Query: 169 QISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEER 228
Q++L+TL +VP IA+AGG YA V + K AY AG++AEEV+ VRTV AF GEE+
Sbjct: 193 QLTLLTLGVVPLIAIAGGGYAIVMSTISEKSEAAYADAGKVAEEVMSQVRTVYAFVGEEK 252
Query: 229 AVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESF 288
AV+SY +L K +++GLAKGLG+G + +LF +WALL WY S++V NG ++F
Sbjct: 253 AVKSYSNSLKKALKLSKRSGLAKGLGVGLTYSLLFCAWALLFWYASLLVRHGKTNGAKAF 312
Query: 289 TTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTV-SKKSSKTGRKLSKLDGH 347
TT+LNV+ SG +LGQA P +SA + + AA IF+MI + + S + + G L + G
Sbjct: 313 TTILNVIYSGFALGQAVPSLSAISKGRVAAANIFKMIGNNNLESSERLENGTTLQNVVGK 372
Query: 348 IQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQI 407
I+F V F+YPSRP++ +F NL+ I +GK A VG SGSGKST++S+++RFYEP SG+I
Sbjct: 373 IEFCGVSFAYPSRPNM-VFENLSFTIHSGKTFAFVGPSGSGKSTIISMVQRFYEPRSGEI 431
Query: 408 LLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
LLD NDI+ L LKWLR+Q+GLV+QEPALFAT+I NIL GK+ A ++++ A K
Sbjct: 432 LLDGNDIKNLKLKWLREQMGLVSQEPALFATTIASNILLGKEKANMDQIIEAAK 485
Score = 207 bits (526), Expect = 1e-53
Identities = 109/317 (34%), Positives = 176/317 (55%)
Query: 145 REGNFLHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYV 204
R + +S I + F W+++ V + P + A G +AY
Sbjct: 803 RLSTIVQNLSLTITALALAFFYSWRVAAVVTACFPLLIAASLTEQLFLKGFGGDYTRAYS 862
Query: 205 RAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFL 264
RA +A E I N+RTV AF+ E++ + L K + G G G G C+ F
Sbjct: 863 RATSLAREAISNIRTVAAFSAEKQISEQFTCELSKPTKSALLRGHISGFGYGLSQCLAFC 922
Query: 265 SWALLVWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEM 324
S+AL +WY SV++ +N N +S + + ++++ S+ + ++ A +F +
Sbjct: 923 SYALGLWYISVLIKRNETNFEDSIKSFMVLLVTAYSVAETLALTPDIVKGTQALGSVFRV 982
Query: 325 IERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGG 384
+ R+T R ++ + G I+F +V F+YP+RP++ IF NLNL + AGK +A+VG
Sbjct: 983 LHRETEIPPDQPNSRLVTHIKGDIEFRNVSFAYPTRPEIAIFKNLNLRVSAGKSLAVVGP 1042
Query: 385 SGSGKSTVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENI 444
SGSGKSTV+ LI RFY+P +G + +D +DI+ ++L+ LR+++ LV QEPALF+TSI ENI
Sbjct: 1043 SGSGKSTVIGLIMRFYDPSNGNLCIDGHDIKSVNLRSLRKKLALVQQEPALFSTSIHENI 1102
Query: 445 LYGKDDATLEELKRAVK 461
YG ++A+ E+ A K
Sbjct: 1103 KYGNENASEAEIIEAAK 1119
>At2g36910 putative ABC transporter
Length = 1286
Score = 355 bits (910), Expect = 4e-98
Identities = 200/459 (43%), Positives = 284/459 (61%), Gaps = 16/459 (3%)
Query: 15 VSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLFPKEASHKV 74
V+ +LF FAD DYVLM IGS+GA VHG S+P+F FF L+N G ++ +V
Sbjct: 27 VAFKELFRFADGLDYVLMGIGSVGAFVHGCSLPLFLRFFADLVNSFGSNSNNVEKMMEEV 86
Query: 75 AKSKCSNIIFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYKF--------I*Y*GIHW 126
K + F++ + + W + S +G + K R K+ I +
Sbjct: 87 LKYA---LYFLVVGAAIWASSWAEISCWMWSGERQT-TKMRIKYLEAALNQDIQFFDTEV 142
Query: 127 RGHFCYYK*HYYC----SRCSIREGNFLHYISRFIAGFTIGFVRVWQISLVTLSIVPAIA 182
R + + S + GNF+HY++ F++GF +GF VWQ++LVTL++VP IA
Sbjct: 143 RTSDVVFAINTDAVMVQDAISEKLGNFIHYMATFVSGFIVGFTAVWQLALVTLAVVPLIA 202
Query: 183 LAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALMKTYV 242
+ GG + L K +++ +AG I E+ + +R V AF GE RA ++Y +AL
Sbjct: 203 VIGGIHTTTLSKLSNKSQESLSQAGNIVEQTVVQIRVVMAFVGESRASQAYSSALKIAQK 262
Query: 243 NGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVISGLSLG 302
G K GLAKG+GLG+ + V+F +ALL+WY +V ++ NGG + TM V+I GL+LG
Sbjct: 263 LGYKTGLAKGMGLGATYFVVFCCYALLLWYGGYLVRHHLTNGGLAIATMFAVMIGGLALG 322
Query: 303 QAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPD 362
Q+AP ++AF +AK AA IF +I+ +++S++G +L + G ++ +V FSYPSRPD
Sbjct: 323 QSAPSMAAFAKAKVAAAKIFRIIDHKPTIERNSESGVELDSVTGLVELKNVDFSYPSRPD 382
Query: 363 VGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELDLKWL 422
V I N L +PAGK +ALVG SGSGKSTVVSLIERFY+P SGQ+LLD D++ L L+WL
Sbjct: 383 VKILNNFCLSVPAGKTIALVGSSGSGKSTVVSLIERFYDPNSGQVLLDGQDLKTLKLRWL 442
Query: 423 RQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
RQQIGLV+QEPALFATSIKENIL G+ DA E++ A +
Sbjct: 443 RQQIGLVSQEPALFATSIKENILLGRPDADQVEIEEAAR 481
Score = 193 bits (490), Expect = 2e-49
Identities = 111/300 (37%), Positives = 167/300 (55%), Gaps = 1/300 (0%)
Query: 161 TIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTV 220
T GFV W+++LV +++ P + A G + A+ + ++A E I NVRTV
Sbjct: 836 TAGFVLQWRLALVLVAVFPVVVAATVLQKMFMTGFSGDLEAAHAKGTQLAGEAIANVRTV 895
Query: 221 QAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKN 280
AF E + VR Y A L G G G G L+ S+AL +WY S +V
Sbjct: 896 AAFNSEAKIVRLYTANLEPPLKRCFWKGQIAGSGYGVAQFCLYASYALGLWYASWLVKHG 955
Query: 281 IANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDT-VSKKSSKTGR 339
I++ ++ + +++S + FI+ A +FE+++R T + T
Sbjct: 956 ISDFSKTIRVFMVLMVSANGAAETLTLAPDFIKGGQAMRSVFELLDRKTEIEPDDPDTTP 1015
Query: 340 KLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERF 399
+L G ++ + FSYPSRPD+ IF +L+L AGK +ALVG SG GKS+V+SLI+RF
Sbjct: 1016 VPDRLRGEVELKHIDFSYPSRPDIQIFRDLSLRARAGKTLALVGPSGCGKSSVISLIQRF 1075
Query: 400 YEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRA 459
YEP SG++++D DIR+ +LK +R+ I +V QEP LF T+I ENI YG + AT E+ +A
Sbjct: 1076 YEPSSGRVMIDGKDIRKYNLKAIRKHIAIVPQEPCLFGTTIYENIAYGHECATEAEIIQA 1135
>At4g01830 putative P-glycoprotein-like protein
Length = 1230
Score = 313 bits (802), Expect = 1e-85
Identities = 177/470 (37%), Positives = 271/470 (57%), Gaps = 17/470 (3%)
Query: 4 KEGDERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLA 63
K+G+ V KLF F+DS D +LM +GSIGAI +G P+ + FG+LI+ +G
Sbjct: 2 KKGNLEANTKTVPFYKLFFFSDSTDVLLMIVGSIGAIANGVCSPLMTLLFGELIDAMG-- 59
Query: 64 YLFPKEASHKVAK---SKCSNIIFI----LDRGGLLDAYWRKTSSKDENGLFKVYVKSRY 116
P + + ++ + C +++++ L L A W T + + +Y+K+
Sbjct: 60 ---PNQNNEEIVERVSKVCLSLVYLGLGALGAAFLQVACWMITGERQAARIRSLYLKTIL 116
Query: 117 KF-I*Y*GIHWRGHFCYYK*H----YYCSRCSIREGNFLHYISRFIAGFTIGFVRVWQIS 171
+ I + + + + G F+ IS F+ GF I F+R W ++
Sbjct: 117 RQDIGFFDVEMTTGEVVGRMSGDTVLILDAMGEKVGKFIQLISTFVGGFVIAFLRGWLLT 176
Query: 172 LVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVR 231
LV L+ +P +A++G A + ++ + AY +A + E+ +G++RTV +F GE++A+
Sbjct: 177 LVMLTSIPLLAMSGAAIAIIVTRASSQEQAAYAKASNVVEQTLGSIRTVASFTGEKQAMS 236
Query: 232 SYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTM 291
SYK + Y + K G GLGLG M V F ++AL W+ ++ + GG M
Sbjct: 237 SYKELINLAYKSNVKQGFVTGLGLGVMFLVFFSTYALGTWFGGEMILRKGYTGGAVINVM 296
Query: 292 LNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFN 351
+ VV S ++LGQA+P ++AF KAAAY +FE IER+ + G+ L + G I+
Sbjct: 297 VTVVSSSIALGQASPCLTAFTAGKAAAYKMFETIEREPLIDTFDLNGKVLEDIRGEIELR 356
Query: 352 DVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDK 411
DVCFSYP+RP +F +L IP+G ALVG SGSGKSTV+SLIERFY+P SGQ+L+D
Sbjct: 357 DVCFSYPARPKEEVFGGFSLLIPSGTTTALVGESGSGKSTVISLIERFYDPNSGQVLIDG 416
Query: 412 NDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
D++E LKW+R +IGLV+QEP LF++SI ENI YGK+ AT+EE++ A K
Sbjct: 417 VDLKEFQLKWIRGKIGLVSQEPVLFSSSIMENIGYGKEGATVEEIQAASK 466
Score = 207 bits (526), Expect = 1e-53
Identities = 116/309 (37%), Positives = 177/309 (56%), Gaps = 8/309 (2%)
Query: 153 ISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEE 212
++ + G I F W+++++ L I+P I + G G A + Y A ++A +
Sbjct: 790 VASLVTGLIIAFTASWEVAIIILVIIPFIGINGYIQIKFMKGFSADAKAKYEEASQVAND 849
Query: 213 VIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWY 272
+G++RTV +F EE+ + YK T +G K GL G+G G VL+ +A +
Sbjct: 850 AVGSIRTVASFCAEEKVMEMYKKRCEDTIKSGIKQGLISGVGFGISFFVLYSVYASCFYV 909
Query: 273 TSVVVHKNIANGGESFTTMLNVVISGLSLGQA---APDISAFIRAKAAAYPIFEMIERDT 329
+ +V N + F L + ++ + + QA APD S + K AA IF +I+R +
Sbjct: 910 GARLVKAGRTNFNDVFQVFLALTLTAVGISQASSFAPDSS---KGKGAAVSIFRIIDRIS 966
Query: 330 VSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGK 389
++G L + G I+ + F+Y +RPDV +F +L L I AG+ VALVG SGSGK
Sbjct: 967 KIDSRDESGMVLENVKGDIELCHISFTYQTRPDVQVFRDLCLSIRAGQTVALVGESGSGK 1026
Query: 390 STVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGK- 448
STV+SL++RFY+P SG I LD ++++L LKWLRQQ+GLV QEP LF +I+ NI YGK
Sbjct: 1027 STVISLLQRFYDPDSGHITLDGVELKKLRLKWLRQQMGLVGQEPVLFNDTIRANIAYGKG 1086
Query: 449 -DDATLEEL 456
++AT E+
Sbjct: 1087 GEEATEAEI 1095
>At1g02520 P-glycoprotein, putative
Length = 1278
Score = 311 bits (797), Expect = 5e-85
Identities = 181/478 (37%), Positives = 270/478 (55%), Gaps = 26/478 (5%)
Query: 4 KEGDERKKEHK-------VSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKL 56
KEG+E KKE K V KLF+FADS D +LM GSIGAI +G S+P + FG L
Sbjct: 23 KEGEETKKEEKSEEKANTVPFYKLFAFADSSDVLLMICGSIGAIGNGMSLPFMTLLFGDL 82
Query: 57 INVIGLAYLFPKEASHK----VAKSKCSNIIFI----LDRGGLLDAYWRKTSSKDENGLF 108
I+ G K ++K V C +++ L L A W T + +
Sbjct: 83 IDSFG------KNQNNKDIVDVVSKVCLKFVYLGLGTLGAAFLQVACWMITGERQAARIR 136
Query: 109 KVYVKSRYKF-I*Y*GIHWRGHFCYYK*H----YYCSRCSIREGNFLHYISRFIAGFTIG 163
Y+K+ + I + + + + G F+ +S F+ GF +
Sbjct: 137 STYLKTILRQDIGFFDVETNTGEVVGRMSGDTVLIQDAMGEKVGKFIQLVSTFVGGFVLA 196
Query: 164 FVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAF 223
F++ W ++LV L+ +P +A+AG A + ++ + AY +A + E+ IG++RTV +F
Sbjct: 197 FIKGWLLTLVMLTSIPLLAMAGAAMALIVTRASSRGQAAYAKAATVVEQTIGSIRTVASF 256
Query: 224 AGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIAN 283
GE++A+ SYK + Y + + G + GLGLG M V F S+AL +W+ ++ +
Sbjct: 257 TGEKQAINSYKKFITSAYKSSIQQGFSTGLGLGVMFFVFFSSYALAIWFGGKMILEKGYT 316
Query: 284 GGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSK 343
GG ++ VV +SLGQ +P ++AF +AAAY +FE I+R + G+ L
Sbjct: 317 GGAVINVIIIVVAGSMSLGQTSPCVTAFAAGQAAAYKMFETIKRKPLIDAYDVNGKVLED 376
Query: 344 LDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPI 403
+ G I+ DV FSYP+RPD IF +L IP+G ALVG SGSGKSTV+SLIERFY+P
Sbjct: 377 IRGDIELKDVHFSYPARPDEEIFDGFSLFIPSGATAALVGESGSGKSTVISLIERFYDPK 436
Query: 404 SGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
SG +L+D +++E LKW+R +IGLV+QEP LF++SI ENI YGK++AT+EE+K A +
Sbjct: 437 SGAVLIDGVNLKEFQLKWIRSKIGLVSQEPVLFSSSIMENIAYGKENATVEEIKAATE 494
Score = 206 bits (525), Expect = 2e-53
Identities = 114/294 (38%), Positives = 168/294 (56%), Gaps = 6/294 (2%)
Query: 158 AGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNV 217
+G I F W+++L+ L ++P I + G G A + Y A ++A + +G++
Sbjct: 842 SGLIIAFTASWELALIILVMLPLIGINGFVQVKFMKGFSADAKSKYEEASQVANDAVGSI 901
Query: 218 RTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVV 277
RTV +F EE+ ++ YK +G K G GLG G +LF +A + + +V
Sbjct: 902 RTVASFCAEEKVMQMYKKQCEGPIKDGIKQGFISGLGFGFSFFILFCVYATSFYAGARLV 961
Query: 278 HKNIANGGESFTTMLNVVISGLSLGQA---APDISAFIRAKAAAYPIFEMIERDTVSKKS 334
F + ++ + + Q+ APD S +AK AA IF +I+R + S
Sbjct: 962 EDGKTTFNNVFQVFFALTMAAIGISQSSTFAPDSS---KAKVAAASIFAIIDRKSKIDSS 1018
Query: 335 SKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVS 394
+TG L + G I+ + F+YP+RPD+ IF +L L I AGK VALVG SGSGKSTV+S
Sbjct: 1019 DETGTVLENVKGDIELRHLSFTYPARPDIQIFRDLCLTIRAGKTVALVGESGSGKSTVIS 1078
Query: 395 LIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGK 448
L++RFY+P SG I LD ++++L LKWLRQQ+GLV QEP LF +I+ NI YGK
Sbjct: 1079 LLQRFYDPDSGHITLDGVELKKLQLKWLRQQMGLVGQEPVLFNDTIRANIAYGK 1132
>At4g01820 P-glycoprotein-like protein pgp3
Length = 1229
Score = 304 bits (778), Expect = 8e-83
Identities = 175/465 (37%), Positives = 263/465 (55%), Gaps = 19/465 (4%)
Query: 10 KKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLFPKE 69
+K V KLFSF+DS D +LM +GSIGAI +G P+ + FG LI+ IG +
Sbjct: 3 EKTKTVPFYKLFSFSDSTDVLLMIVGSIGAIGNGVGFPLMTLLFGDLIDSIG------QN 56
Query: 70 ASHK----VAKSKCSNIIFI----LDRGGLLDAYWRKTSSKDENGLFKVYVKSRYKF-I* 120
S+K + C +++ L L A W T + + +Y+K+ + I
Sbjct: 57 QSNKDIVEIVSKVCLKFVYLGLGTLGAAFLQVACWMITGERQAARIRSLYLKTILRQDIG 116
Query: 121 Y*GIHWRGHFCYYK*H----YYCSRCSIREGNFLHYISRFIAGFTIGFVRVWQISLVTLS 176
+ + + + G F+ I+ F+ GF + FV+ W ++LV L
Sbjct: 117 FFDVETSTGEVVGRMSGDTVLILEAMGEKVGKFIQLIATFVGGFVLAFVKGWLLTLVMLV 176
Query: 177 IVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAA 236
+P +A+AG + ++ + AY +A + E+ +G++RTV +F GE++A++SY+
Sbjct: 177 SIPLLAIAGAAMPIIVTRASSREQAAYAKASTVVEQTLGSIRTVASFTGEKQAMKSYREF 236
Query: 237 LMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVI 296
+ Y K G + GLGLG + V F S+AL +W+ ++ K GGE M+ VV
Sbjct: 237 INLAYRASVKQGFSMGLGLGVVFFVFFCSYALAIWFGGEMILKKGYTGGEVVNVMVTVVA 296
Query: 297 SGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFS 356
S +SLGQ P ++AF KAAAY +FE IER G+ L + G I+ DVCFS
Sbjct: 297 SSMSLGQTTPCLTAFAAGKAAAYKMFETIERKPSIDAFDLNGKVLEDIRGEIELRDVCFS 356
Query: 357 YPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRE 416
YP+RP +F +L IP+G ALVG SGSGKS+V+SLIERFY+P SG +L+D +++E
Sbjct: 357 YPARPMEEVFGGFSLLIPSGATAALVGESGSGKSSVISLIERFYDPSSGSVLIDGVNLKE 416
Query: 417 LDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
LKW+R +IGLV+QEP LF++SI ENI YGK++AT+EE++ A K
Sbjct: 417 FQLKWIRGKIGLVSQEPVLFSSSIMENIGYGKENATVEEIQAAAK 461
Score = 204 bits (518), Expect = 1e-52
Identities = 115/310 (37%), Positives = 179/310 (57%), Gaps = 8/310 (2%)
Query: 157 IAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGN 216
++G I F W+++++ L ++P I + G G A + Y A ++A + +G+
Sbjct: 793 VSGLIIAFTASWKLAVIILVMIPLIGINGYLQIKFIKGFTADAKAKYEEASQVANDAVGS 852
Query: 217 VRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVV 276
+RTV +F EE+ + YK T +G K GL G+G G VL+ +A + + +
Sbjct: 853 IRTVASFCAEEKVMEMYKKRCEDTIKSGIKQGLISGVGFGISFFVLYSVYASCFYVGARL 912
Query: 277 VHKNIANGGESFTTMLNVVISGLSLGQA---APDISAFIRAKAAAYPIFEMIERDTVSKK 333
V N + F L + ++ + + QA APD S +AK AA IF +I+ ++
Sbjct: 913 VKAGRTNFNDVFQVFLALTMTAIGISQASSFAPDSS---KAKGAAASIFGIIDGKSMIDS 969
Query: 334 SSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVV 393
++G L + G I+ + F+Y +RPDV IF +L I AG+ VALVG SGSGKSTV+
Sbjct: 970 RDESGLVLENVKGDIELCHISFTYQTRPDVQIFRDLCFAIRAGQTVALVGESGSGKSTVI 1029
Query: 394 SLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGK--DDA 451
SL++RFY+P SG I LD+ ++++L LKW+RQQ+GLV QEP LF +I+ NI YGK D+A
Sbjct: 1030 SLLQRFYDPDSGHITLDRVELKKLQLKWVRQQMGLVGQEPVLFNDTIRSNIAYGKGGDEA 1089
Query: 452 TLEELKRAVK 461
+ E+ A +
Sbjct: 1090 SEAEIIAAAE 1099
>At3g62150 P-glycoprotein-like proetin
Length = 1292
Score = 304 bits (778), Expect = 8e-83
Identities = 175/480 (36%), Positives = 271/480 (56%), Gaps = 33/480 (6%)
Query: 2 ESKEGDERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIG 61
E + +E +K V KLF+FADS+D +LM +G+IGA+ +G PI I FG +I+V G
Sbjct: 50 EKNKQEEDEKTKTVPFHKLFAFADSFDIILMILGTIGAVGNGLGFPIMTILFGDVIDVFG 109
Query: 62 LAYLFPKEASHKVAKSKCSNIIFILDRGGLLDAY-----WRKTSSKDENGLFKVYVKSRY 116
+ S K+AK + L G L+ A W + + + +Y+++
Sbjct: 110 QNQN-SSDVSDKIAKVALKFVY--LGLGTLVAALLQVSGWMISGERQAGRIRSLYLQTIL 166
Query: 117 KFI*Y*GIHWRGHFCYYK*HYYCSRCSIRE---------------GNFLHYISRFIAGFT 161
R ++ R G + +S FI GF
Sbjct: 167 ----------RQDIAFFDVETNTGEVVGRMSGDTVLIQDAMGEKVGKAIQLVSTFIGGFV 216
Query: 162 IGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQ 221
I F W ++LV +S +P + ++G A V + ++ + +Y +A + E+ +G++RTV
Sbjct: 217 IAFTEGWLLTLVMVSSIPLLVMSGAALAIVISKMASRGQTSYAKAAVVVEQTVGSIRTVA 276
Query: 222 AFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNI 281
+F GE++A+ +Y L+ Y G G + GLGLG+++ V+F ++AL VWY ++ +
Sbjct: 277 SFTGEKQAISNYNKHLVSAYRAGVFEGASTGLGLGTLNIVIFCTYALAVWYGGKMILEKG 336
Query: 282 ANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKL 341
GG+ + V+ +SLGQA+P +SAF +AAAY +FE I+R S TG+ L
Sbjct: 337 YTGGQVLIIIFAVLTGSMSLGQASPCLSAFAAGQAAAYKMFEAIKRKPEIDASDTTGKVL 396
Query: 342 SKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYE 401
+ G I+ N+V FSYP+RP+ IF +L I +G VALVG SGSGKSTVVSLIERFY+
Sbjct: 397 DDIRGDIELNNVNFSYPARPEEQIFRGFSLSISSGSTVALVGQSGSGKSTVVSLIERFYD 456
Query: 402 PISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
P SG++ +D +++E LKW+R +IGLV+QEP LF +SIKENI YGK++AT+EE+++A +
Sbjct: 457 PQSGEVRIDGINLKEFQLKWIRSKIGLVSQEPVLFTSSIKENIAYGKENATVEEIRKATE 516
Score = 213 bits (541), Expect = 2e-55
Identities = 122/308 (39%), Positives = 180/308 (57%), Gaps = 11/308 (3%)
Query: 158 AGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNV 217
AG I FV WQ++ + L+++P I L G Y +G A ++A ++A + +G++
Sbjct: 862 AGLVIAFVASWQLAFIVLAMLPLIGLNGYIYMKFMVGFSADAKEA----SQVANDAVGSI 917
Query: 218 RTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVV 277
RTV +F EE+ ++ YK G + G+ G+G G VLF S+A + + +V
Sbjct: 918 RTVASFCAEEKVMKMYKKKCEGPMRTGIRQGIVSGIGFGVSFFVLFSSYAASFYAGARLV 977
Query: 278 HKNIANGGESFTTMLNVVISGLSLGQAA---PDISAFIRAKAAAYPIFEMIERDTVSKKS 334
F + ++ +++ Q++ PD S +A AA IF +I+R++ S
Sbjct: 978 DDGKTTFDSVFRVFFALTMAAVAISQSSSLSPDSS---KASNAAASIFAVIDRESKIDPS 1034
Query: 335 SKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVS 394
++GR L + G I+ + F YPSRPDV IF +L L I AGK +ALVG SGSGKSTV++
Sbjct: 1035 DESGRVLDNVKGDIELRHISFKYPSRPDVQIFQDLCLSIRAGKTIALVGESGSGKSTVIA 1094
Query: 395 LIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGK-DDATL 453
L++RFY+P SGQI LD +I+ L LKWLRQQ GLV+QEP LF +I+ NI YGK DAT
Sbjct: 1095 LLQRFYDPDSGQITLDGVEIKTLQLKWLRQQTGLVSQEPVLFNETIRANIAYGKGGDATE 1154
Query: 454 EELKRAVK 461
E+ A +
Sbjct: 1155 TEIVSAAE 1162
Score = 30.4 bits (67), Expect = 2.1
Identities = 19/59 (32%), Positives = 34/59 (57%), Gaps = 5/59 (8%)
Query: 11 KEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLFPKE 69
KE KVS ++ + + + ++ +GSI A+++G +PI FG LI+ + A+ P E
Sbjct: 710 KEKKVSFFRVAAL-NKPEIPMLILGSIAAVLNGVILPI----FGILISSVIKAFFKPPE 763
>At1g02530 hypothetical protein
Length = 1273
Score = 300 bits (768), Expect = 1e-81
Identities = 176/481 (36%), Positives = 267/481 (54%), Gaps = 31/481 (6%)
Query: 5 EGDERKKEHKVS----------MLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFG 54
EGD EH S + KLF+FADS+D LM GS+GAI +G +P+ + FG
Sbjct: 8 EGDSVSHEHSTSKTDEKAKTVPLYKLFAFADSFDVFLMICGSLGAIGNGVCLPLMTLLFG 67
Query: 55 KLINVIGLAYLFPKEASHK----VAKSKCSNIIFI----LDRGGLLDAYWRKTSSKDENG 106
LI+ G K ++K V C +++ L L A W T +
Sbjct: 68 DLIDSFG------KNQNNKDIVDVVSKVCLKFVYLGLGRLGAAFLQVACWMITGERQAAK 121
Query: 107 LFKVYVKSRYKF-I*Y*GIHWR-----GHFCYYK*HYYCSRCSIREGNFLHYISRFIAGF 160
+ Y+K+ + I + + G H + G F+ +S F+ GF
Sbjct: 122 IRSNYLKTILRQDIGFFDVETNTGEVVGRMSGDTVHIQ-DAMGEKVGKFIQLVSTFVGGF 180
Query: 161 TIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTV 220
+ F + W ++LV L+ +P +A+AG A + ++ + AY +A + E+ IG++RTV
Sbjct: 181 ALAFAKGWLLTLVMLTSIPFLAMAGAAMALLVTRASSRGQAAYAKAATVVEQTIGSIRTV 240
Query: 221 QAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKN 280
+F GE++A+ SYK + Y + + G + GLGLG M V F S+AL +W+ ++ +
Sbjct: 241 ASFTGEKQAINSYKKYITSAYKSSIQQGFSTGLGLGVMIYVFFSSYALAIWFGGKMILEK 300
Query: 281 IANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRK 340
GG ++ VV +SLGQ +P ++AF +AAAY +FE I+R + G+
Sbjct: 301 GYTGGSVINVIIIVVAGSMSLGQTSPCVTAFAAGQAAAYKMFETIKRKPLIDAYDVNGKV 360
Query: 341 LSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFY 400
L + G I+ DV FSYP+RPD IF +L IP+G ALVG SGSGKSTV++LIERFY
Sbjct: 361 LGDIRGDIELKDVHFSYPARPDEEIFDGFSLFIPSGATAALVGESGSGKSTVINLIERFY 420
Query: 401 EPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAV 460
+P +G++L+D +++E LKW+R +IGLV QEP LF++SI ENI YGK++ATL+E+K A
Sbjct: 421 DPKAGEVLIDGINLKEFQLKWIRSKIGLVCQEPVLFSSSIMENIAYGKENATLQEIKVAT 480
Query: 461 K 461
+
Sbjct: 481 E 481
Score = 204 bits (518), Expect = 1e-52
Identities = 113/294 (38%), Positives = 168/294 (56%), Gaps = 6/294 (2%)
Query: 158 AGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNV 217
+G I F W+++L+ L ++P I + G G A + Y A ++A + +G++
Sbjct: 837 SGLIIAFTASWELALIILVMLPLIGINGFLQVKFMKGFSADAKSKYEEASQVANDAVGSI 896
Query: 218 RTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVV 277
RTV +F EE+ ++ Y +G K G GLG G +LF +A + + +V
Sbjct: 897 RTVASFCAEEKVMQMYNKQCEGPIKDGVKQGFISGLGFGFSFFILFCVYATSFYAAARLV 956
Query: 278 HKNIANGGESFTTMLNVVISGLSLGQA---APDISAFIRAKAAAYPIFEMIERDTVSKKS 334
+ F + ++ + + Q+ APD S +AK AA IF +I+R + S
Sbjct: 957 EDGKTTFIDVFQVFFALTMAAIGISQSSTFAPDSS---KAKVAAASIFAIIDRKSKIDSS 1013
Query: 335 SKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVS 394
+TG L + G I+ + F+YP+RP + IF +L L I AGK VALVG SGSGKSTV+S
Sbjct: 1014 DETGTVLENVKGDIELRHLSFTYPARPGIQIFRDLCLTIRAGKTVALVGESGSGKSTVIS 1073
Query: 395 LIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGK 448
L++RFY+P SGQI LD ++++L LKWLRQQ+GLV QEP LF +I+ NI YGK
Sbjct: 1074 LLQRFYDPDSGQITLDGVELKKLQLKWLRQQMGLVGQEPVLFNDTIRANIAYGK 1127
>At5g46540 multidrug resistance p-glycoprotein
Length = 1248
Score = 292 bits (747), Expect = 3e-79
Identities = 166/467 (35%), Positives = 263/467 (55%), Gaps = 32/467 (6%)
Query: 14 KVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLA---YLFPKEA 70
+++ KLF+FAD YD VLM IG++ A+ +G + P I G+LINV G + ++F KE
Sbjct: 17 RIAFYKLFTFADRYDIVLMVIGTLSAMANGLTQPFMSILMGQLINVFGFSDHDHVF-KEV 75
Query: 71 SHKVAKSKCSNIIFILDRGGLLD----AYWRKTSSKDENGLFKVYVKSRYKFI*Y*GIHW 126
S K +++ G++ + W T + + ++Y+K+ +
Sbjct: 76 SKVAVK-----FLYLAAYAGVVSFLQVSCWMVTGERQSTRIRRLYLKTILR-------QD 123
Query: 127 RGHF-CYYK*HYYCSRCS-----------IREGNFLHYISRFIAGFTIGFVRVWQISLVT 174
G F R S + G F +S F+ GFT+ F+ +++L
Sbjct: 124 IGFFDTETNTGEVIGRMSGDTILIQDSMGEKVGKFTQLVSSFVGGFTVAFIVGMKLTLAL 183
Query: 175 LSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYK 234
L VP I GG Y+ +V+ AY AG + ++ +G++RTV AF GE++++ Y+
Sbjct: 184 LPCVPLIVGTGGAMTYIMSKKAQRVQLAYTEAGNVVQQAVGSIRTVVAFTGEKQSMGKYE 243
Query: 235 AALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNV 294
L Y + K GL GLG+G M V++ ++ +WY + + + GG+ + ++
Sbjct: 244 KKLEIAYKSMVKQGLYSGLGIGIMMVVVYCTYGFAIWYGARQIIEKGYTGGQVMNVITSI 303
Query: 295 VISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVC 354
+ G++LGQ P +++F AAAY +FE I+R +G L ++ G I+ DV
Sbjct: 304 LTGGMALGQTLPSLNSFAAGTAAAYKMFETIKRKPKIDAYDMSGEVLEEIKGDIELRDVY 363
Query: 355 FSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDI 414
F YP+RPDV IF +L +P G VALVG SGSGKSTV+SLIERFY+P SG++L+D D+
Sbjct: 364 FRYPARPDVQIFVGFSLTVPNGMTVALVGQSGSGKSTVISLIERFYDPESGEVLIDGIDL 423
Query: 415 RELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
++ +KW+R +IGLV+QEP LFAT+I+ENI+YGK DA+ +E++ A+K
Sbjct: 424 KKFQVKWIRSKIGLVSQEPILFATTIRENIVYGKKDASDQEIRTALK 470
Score = 197 bits (500), Expect = 1e-50
Identities = 118/319 (36%), Positives = 177/319 (54%), Gaps = 7/319 (2%)
Query: 147 GNFLHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRA 206
G + ++ I F I F W ++L+ L + P + G G AK R Y A
Sbjct: 804 GLIMQNMATIIGAFIIAFTANWLLALMALLVAPVMFFQGYYQIKFITGFGAKARGKYEEA 863
Query: 207 GEIAEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSW 266
++A + + ++RTV +F E++ + Y+ + G K GL GL G + L++
Sbjct: 864 SQVASDAVSSIRTVASFCAEDKVMDLYQEKCDEPKQQGFKLGLVSGLCYGGSYLALYVIE 923
Query: 267 ALLVWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQA---APDISAFIRAKAAAYPIFE 323
++ S ++ A GE F + ++ + + Q APDI+ +AK +A IF+
Sbjct: 924 SVCFLGGSWLIQNRRATFGEFFQVFFALTLTAVGVTQTSTMAPDIN---KAKDSAASIFD 980
Query: 324 MIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVG 383
+++ SS+ G L + G I+ V F YP RPD+ IF++L L I +G+ VALVG
Sbjct: 981 ILDSKPKIDSSSEKGTILPIVHGDIELQHVSFRYPMRPDIQIFSDLCLTISSGQTVALVG 1040
Query: 384 GSGSGKSTVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKEN 443
SGSGKSTV+SL+ERFY+P SG+ILLD+ +I+ L L WLR+Q+GLV+QEP LF +I N
Sbjct: 1041 ESGSGKSTVISLLERFYDPDSGKILLDQVEIQSLKLSWLREQMGLVSQEPVLFNETIGSN 1100
Query: 444 ILYGK-DDATLEELKRAVK 461
I YGK AT EE+ A K
Sbjct: 1101 IAYGKIGGATEEEIITAAK 1119
>At2g47000 putative ABC transporter
Length = 1286
Score = 290 bits (742), Expect = 1e-78
Identities = 174/463 (37%), Positives = 259/463 (55%), Gaps = 5/463 (1%)
Query: 4 KEGDERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLA 63
K+ +E +K V KLF+FADS+D++LM +G++G+I +G P+ + FG LI+ G
Sbjct: 35 KKDEEHEKTKTVPFYKLFAFADSFDFLLMILGTLGSIGNGLGFPLMTLLFGDLIDAFGEN 94
Query: 64 YLFPKEASHKVAKSKCSNIIFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYK-FI*Y* 122
+ KVA I L + W + + + +Y+K+ + I +
Sbjct: 95 QTNTTDKVSKVALKFVWLGIGTFAAAFLQLSGWMISGERQAARIRSLYLKTILRQDIAFF 154
Query: 123 GIHWRGHFCYYK*HYYCSRCSIREGNFLHYISRFIAGFTIG----FVRVWQISLVTLSIV 178
I + G + + +A F G FVR W ++LV LS +
Sbjct: 155 DIDTNTGEVVGRMSGDTVLIQDAMGEKVGKAIQLLATFVGGFVIAFVRGWLLTLVMLSSI 214
Query: 179 PAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALM 238
P + +AG A V ++ + AY +A + E+ IG++RTV +F GE++A+ +Y L+
Sbjct: 215 PLLVMAGALLAIVIAKTASRGQTAYAKAATVVEQTIGSIRTVASFTGEKQAISNYNKHLV 274
Query: 239 KTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVISG 298
Y G G + GLGLG++ V+F S+AL VWY ++ GG+ ++ V+
Sbjct: 275 TAYKAGVIEGGSTGLGLGTLFLVVFCSYALAVWYGGKLILDKGYTGGQVLNIIIAVLTGS 334
Query: 299 LSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYP 358
+SLGQ +P +SAF +AAAY +FE IER S G+ L + G I+ DV F+YP
Sbjct: 335 MSLGQTSPCLSAFAAGQAAAYKMFETIERRPNIDSYSTNGKVLDDIKGDIELKDVYFTYP 394
Query: 359 SRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELD 418
+RPD IF +L I +G VALVG SGSGKSTVVSLIERFY+P +G +L+D +++E
Sbjct: 395 ARPDEQIFRGFSLFISSGTTVALVGQSGSGKSTVVSLIERFYDPQAGDVLIDGINLKEFQ 454
Query: 419 LKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
LKW+R +IGLV+QEP LF SIK+NI YGK+DAT EE+K A +
Sbjct: 455 LKWIRSKIGLVSQEPVLFTASIKDNIAYGKEDATTEEIKAAAE 497
Score = 216 bits (550), Expect = 2e-56
Identities = 122/308 (39%), Positives = 181/308 (58%), Gaps = 7/308 (2%)
Query: 153 ISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEE 212
+S +AG I F+ WQ++ V L+++P IAL G Y G A +K Y A ++A +
Sbjct: 847 LSSILAGLIIAFLACWQLAFVVLAMLPLIALNGFLYMKFMKGFSADAKKMYGEASQVAND 906
Query: 213 VIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWY 272
+G++RTV +F E++ + Y NG + G+ G+G G VLF S+A +
Sbjct: 907 AVGSIRTVASFCAEDKVMNMYSKKCEGPMKNGIRQGIVSGIGFGFSFFVLFSSYAASFYV 966
Query: 273 TSVVVHKNIANGGESFTTMLNVVISGLSLGQAA---PDISAFIRAKAAAYPIFEMIERDT 329
+ +V F + ++ +++ Q++ PD S +A AA IF +++R++
Sbjct: 967 GARLVDDGKTTFDSVFRVFFALTMAAMAISQSSSLSPDSS---KADVAAASIFAIMDRES 1023
Query: 330 VSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGK 389
S ++GR L + G I+ V F YP+RPDV IF +L L I AGK VALVG SGSGK
Sbjct: 1024 KIDPSVESGRVLDNVKGDIELRHVSFKYPARPDVQIFQDLCLSIRAGKTVALVGESGSGK 1083
Query: 390 STVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGK- 448
STV++L++RFY+P SG+I LD +I+ L LKWLRQQ GLV+QEP LF +I+ NI YGK
Sbjct: 1084 STVIALLQRFYDPDSGEITLDGVEIKSLRLKWLRQQTGLVSQEPILFNETIRANIAYGKG 1143
Query: 449 DDATLEEL 456
DA+ E+
Sbjct: 1144 GDASESEI 1151
>At4g18050 multidrug resistance protein/P-glycoprotein - like
Length = 1323
Score = 285 bits (729), Expect = 4e-77
Identities = 171/459 (37%), Positives = 252/459 (54%), Gaps = 16/459 (3%)
Query: 14 KVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLFPKEASH- 72
KVS KLFSFAD D VLM +G+I A +G + P + FG+LIN G + H
Sbjct: 15 KVSFFKLFSFADKTDVVLMTVGTIAAAGNGLTQPFMTLIFGQLINAFGTT-----DPDHM 69
Query: 73 -----KVAKSKCSNIIFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYK-FI*Y*GIHW 126
KVA ++ L + W T + + +Y+K+ + I Y
Sbjct: 70 VREVWKVAVKFIYLAVYSCVVAFLQVSCWMVTGERQSATIRGLYLKTILRQDIGYFDTET 129
Query: 127 RGHFCYYK*HYYCSRCSIREGNFLHYISRFIAGFTIGFVRVWQ----ISLVTLSIVPAIA 182
+ G + ++ + F GF + ++ V S +P I
Sbjct: 130 NTGEVIGRMSGDTILIQDAMGEKVGKFTQLLCTFLGGFAIAFYKGPLLAGVLCSCIPLIV 189
Query: 183 LAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALMKTYV 242
+AG + + + + + AY AG + E+ +G +RTV AF GE++A Y++ L Y
Sbjct: 190 IAGAAMSLIMSKMAGRGQVAYAEAGNVVEQTVGAIRTVVAFTGEKQATEKYESKLEIAYK 249
Query: 243 NGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVISGLSLG 302
+ GL G GLG+M V+F S+ L VWY + ++ + NGG+ + V+ G+SLG
Sbjct: 250 TVVQQGLISGFGLGTMLAVIFCSYGLAVWYGAKLIMEKGYNGGQVINVIFAVLTGGMSLG 309
Query: 303 QAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPD 362
Q +P ++AF +AAA+ +FE I+R +G L + G I+ DV F YP+RPD
Sbjct: 310 QTSPSLNAFAAGRAAAFKMFETIKRSPKIDAYDMSGSVLEDIRGDIELKDVYFRYPARPD 369
Query: 363 VGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELDLKWL 422
V IF +L +P GK VALVG SGSGKSTV+SLIERFY+P SGQ+L+D D+++L LKW+
Sbjct: 370 VQIFAGFSLFVPNGKTVALVGQSGSGKSTVISLIERFYDPESGQVLIDNIDLKKLQLKWI 429
Query: 423 RQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
R +IGLV+QEP LFAT+IKENI YGK+DAT +E++ A++
Sbjct: 430 RSKIGLVSQEPVLFATTIKENIAYGKEDATDQEIRTAIE 468
Score = 201 bits (511), Expect = 7e-52
Identities = 121/317 (38%), Positives = 171/317 (53%), Gaps = 15/317 (4%)
Query: 153 ISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEE 212
I+ G I F W ++L+ L++ P I + G G A + Y A ++A +
Sbjct: 840 IATVTTGLIIAFTANWILALIVLALSPFIVIQGYAQTKFLTGFSADAKAMYEEASQVAND 899
Query: 213 VIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLG-------SMHCVLFLS 265
+ ++RTV +F EE+ + Y+ NG + GL G G G ++CV F+S
Sbjct: 900 AVSSIRTVASFCAEEKVMDLYQQKCDGPKKNGVRLGLLSGAGFGFSFFFLYCINCVCFVS 959
Query: 266 WALLVWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMI 325
A L+ A GE F + I + + Q + +AK +A IF+++
Sbjct: 960 GAGLIQIGK-------ATFGEVFKVFFALTIMAIGVSQTSAMAPDSNKAKDSAASIFDIL 1012
Query: 326 ERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGS 385
+ SS G L ++G I+F V F YP RPDV IF +L L IP+GK VALVG S
Sbjct: 1013 DSTPKIDSSSDEGTTLQNVNGDIEFRHVSFRYPMRPDVQIFRDLCLTIPSGKTVALVGES 1072
Query: 386 GSGKSTVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENIL 445
GSGKSTV+S+IERFY P SG+IL+D+ +I+ L WLRQQ+GLV+QEP LF +I+ NI
Sbjct: 1073 GSGKSTVISMIERFYNPDSGKILIDQVEIQTFKLSWLRQQMGLVSQEPILFNETIRSNIA 1132
Query: 446 YGK-DDATLEELKRAVK 461
YGK AT EE+ A K
Sbjct: 1133 YGKTGGATEEEIIAAAK 1149
Score = 37.7 bits (86), Expect = 0.013
Identities = 21/71 (29%), Positives = 33/71 (45%)
Query: 2 ESKEGDERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIG 61
+ E +E HK LK + + + ++ +GSI A+VHG PIF + IN+
Sbjct: 656 DEMEDEENNVRHKKVSLKRLAHLNKPEIPVLVLGSIAAMVHGTVFPIFGLLLSSSINMFY 715
Query: 62 LAYLFPKEASH 72
K+ SH
Sbjct: 716 EPAKILKKDSH 726
>At3g55320 P-glycoprotein - like
Length = 1408
Score = 283 bits (723), Expect = 2e-76
Identities = 171/460 (37%), Positives = 259/460 (56%), Gaps = 17/460 (3%)
Query: 15 VSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLFPKEAS-HK 73
V +LF+ AD +D+VLM +GS+ A HG ++ ++ +F K+++V+ + ++ S H+
Sbjct: 71 VPFSQLFACADRFDWVLMIVGSVAAAAHGTALIVYLHYFAKIVDVLAFSNDSSQQRSEHQ 130
Query: 74 VAKSKCSNIIFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYKFI*Y*GIHWRGHFCYY 133
+ ++ + GG+ + W + S G + V R K++ F Y
Sbjct: 131 FDRLVQLSLTIVYIAGGVFISGWIEVSCWILTGERQTAV-IRSKYVQVLLNQDMSFFDTY 189
Query: 134 K*H------------YYCSRCSIREGNFLHYISRFIAGFTIGFVRVWQISLVTLSIVPAI 181
+ S S + GN++H ++ FI+G IGFV W+I+L+TL+ P I
Sbjct: 190 GNNGDIVSQVLSDVLLIQSALSEKVGNYIHNMATFISGLVIGFVNCWEIALITLATGPFI 249
Query: 182 ALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALMKTY 241
AGG L ++ AY A IAE+ I +RT+ AF E A SY +L T
Sbjct: 250 VAAGGISNIFLHRLAENIQDAYAEAAGIAEQAISYIRTLYAFTNETLAKYSYATSLQATL 309
Query: 242 VNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVISGLSL 301
G L +GLGLG + + S AL +W VH ANGGE + V++SGL L
Sbjct: 310 RYGILISLVQGLGLGFTYGLAICSCALQLWIGRFFVHNGRANGGEIIAALFAVILSGLGL 369
Query: 302 GQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRP 361
QAA + +F + + AAY +FEMI R S +++ G L+ + G+I+F +V FSY SRP
Sbjct: 370 NQAATNFYSFDQGRIAAYRLFEMITRS--SSVANQEGAVLASVQGNIEFRNVYFSYLSRP 427
Query: 362 DVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELDLKW 421
++ I + L +PA K VALVG +GSGKS+++ L+ERFY+P G++LLD +I+ L L+W
Sbjct: 428 EIPILSGFYLTVPAKKAVALVGRNGSGKSSIIPLMERFYDPTLGEVLLDGENIKNLKLEW 487
Query: 422 LRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
LR QIGLV QEPAL + SI+ENI YG+ DATL++++ A K
Sbjct: 488 LRSQIGLVTQEPALLSLSIRENIAYGR-DATLDQIEEAAK 526
Score = 182 bits (462), Expect = 3e-46
Identities = 93/305 (30%), Positives = 170/305 (55%)
Query: 157 IAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGN 216
I IG + W+++LV L+ +P + L+ G +++ + +A + E+ + N
Sbjct: 968 IVALLIGLLLGWRLALVALATLPILTLSAIAQKLWLAGFSKGIQEMHRKASLVLEDAVRN 1027
Query: 217 VRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVV 276
+ TV AF + + Y+ L + G+A G G +LF ALL+W T++
Sbjct: 1028 IYTVVAFCAGNKVMELYRMQLQRILRQSYLHGMAIGFAFGFSQFLLFACNALLLWCTALS 1087
Query: 277 VHKNIANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSK 336
V++ + T + + +L + ++ + + +FE+++R +
Sbjct: 1088 VNRGYMKLSTAITEYMVFSFATFALVEPFGLAPYILKRRKSLISVFEIVDRVPTIEPDDN 1147
Query: 337 TGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLI 396
+ K + G I+ +V F YP+RP++ + +N +L I G+ VA+VG SGSGKST++SL+
Sbjct: 1148 SALKPPNVYGSIELKNVDFCYPTRPEILVLSNFSLKISGGQTVAVVGVSGSGKSTIISLV 1207
Query: 397 ERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEEL 456
ER+Y+P++GQ+LLD D++ +L+WLR +GLV QEP +F+T+I+ENI+Y + +A+ E+
Sbjct: 1208 ERYYDPVAGQVLLDGRDLKLYNLRWLRSHMGLVQQEPIIFSTTIRENIIYARHNASEAEM 1267
Query: 457 KRAVK 461
K A +
Sbjct: 1268 KEAAR 1272
>At2g39480 putative ABC transporter
Length = 1407
Score = 282 bits (721), Expect = 3e-76
Identities = 172/464 (37%), Positives = 260/464 (55%), Gaps = 25/464 (5%)
Query: 15 VSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLFPKEASHKV 74
V +LF+ AD +D+VLM GS+ A HG ++ ++ +F K++ V+ FP ++ H +
Sbjct: 69 VPFSQLFACADRFDWVLMVFGSVAAAAHGTALIVYLHYFAKIVQVLA----FPTDSDHLI 124
Query: 75 AKSKCSNII-----FILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYKFI*Y*GIHWRGH 129
+ + + ++ + GG+ + W + S G + V R K++
Sbjct: 125 SDDQFNRLLELSLTIVYIAGGVFISGWIEVSCWILTGERQTAV-IRSKYVQVLLNQDMSF 183
Query: 130 FCYYK*H------------YYCSRCSIREGNFLHYISRFIAGFTIGFVRVWQISLVTLSI 177
F Y + S S + GN++H ++ FI+G IGFV W+I+L+TL+
Sbjct: 184 FDTYGNNGDIVSQVLSDVLLIQSALSEKVGNYIHNMATFISGLIIGFVNCWEIALITLAT 243
Query: 178 VPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAAL 237
P I AGG L ++ AY A IAE+ + VRT+ AF E A SY +L
Sbjct: 244 GPFIVAAGGISNIFLHRLAENIQDAYAEAASIAEQAVSYVRTLYAFTNETLAKYSYATSL 303
Query: 238 MKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVIS 297
T G L +GLGLG + + S A+ +W V + ANGGE T + V++S
Sbjct: 304 QATLRYGILISLVQGLGLGFTYGLAICSCAMQLWIGRFFVIHHRANGGEIITALFAVILS 363
Query: 298 GLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSY 357
GL L QAA + +F + + AAY +FEMI R S +++ G LS + G+I+F +V FSY
Sbjct: 364 GLGLNQAATNFYSFDQGRIAAYRLFEMISRS--SSGTNQEGIILSAVQGNIEFRNVYFSY 421
Query: 358 PSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIREL 417
SRP++ I + L +PA K VALVG +GSGKS+++ L+ERFY+P G++LLD +I+ L
Sbjct: 422 LSRPEIPILSGFYLTVPAKKAVALVGRNGSGKSSIIPLMERFYDPTLGEVLLDGENIKNL 481
Query: 418 DLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
L+WLR QIGLV QEPAL + SI+ENI YG+ DATL++++ A K
Sbjct: 482 KLEWLRSQIGLVTQEPALLSLSIRENIAYGR-DATLDQIEEAAK 524
Score = 182 bits (463), Expect = 3e-46
Identities = 93/305 (30%), Positives = 169/305 (54%)
Query: 157 IAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGN 216
I IG + W+++LV L+ +P + L+ G +++ + +A + E+ + N
Sbjct: 967 IVAILIGLLLGWRLALVALATLPVLTLSAIAQKLWLAGFSKGIQEMHRKASLVLEDAVRN 1026
Query: 217 VRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVV 276
+ TV AF + + Y+ L + G+A G G +LF ALL+WYT++
Sbjct: 1027 IYTVVAFCAGNKVMELYRLQLQRILRQSFFHGMAIGFAFGFSQFLLFACNALLLWYTALS 1086
Query: 277 VHKNIANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSK 336
V + + T + + +L + ++ + + +FE+I+R +
Sbjct: 1087 VDRRYMKLSTALTEYMVFSFATFALVEPFGLAPYILKRRRSLASVFEIIDRVPTIEPDDT 1146
Query: 337 TGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLI 396
+ + G I+ ++ F YP+RP+V + +N +L + G+ VA+VG SGSGKST++SLI
Sbjct: 1147 SALSPPNVYGSIELKNIDFCYPTRPEVLVLSNFSLKVNGGQTVAVVGVSGSGKSTIISLI 1206
Query: 397 ERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEEL 456
ER+Y+P++GQ+LLD D++ +L+WLR +GL+ QEP +F+T+I+ENI+Y + +A+ E+
Sbjct: 1207 ERYYDPVAGQVLLDGRDLKSYNLRWLRSHMGLIQQEPIIFSTTIRENIIYARHNASEAEM 1266
Query: 457 KRAVK 461
K A +
Sbjct: 1267 KEAAR 1271
>At3g28390 P-glycoprotein, putative
Length = 1225
Score = 280 bits (717), Expect = 9e-76
Identities = 166/466 (35%), Positives = 259/466 (54%), Gaps = 32/466 (6%)
Query: 16 SMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLFPKEASHKVA 75
S+ +F AD D++LM +G IGA+ G PI F KL+N +G + + VA
Sbjct: 7 SIRSIFMHADGVDWMLMALGLIGAVGDGFITPIIFFICSKLLNNVGGSSFDDETFMQTVA 66
Query: 76 KSKCSNIIFILDRGGLL---DAY-WRKTSSKDENGLFKVYVKSRYKFI*Y*GIHWRGHFC 131
K+ + ++++ ++ + Y W +T + + + Y+K+ R
Sbjct: 67 KNAVA-LVYVACASWVICFIEGYCWTRTGERQAAKMREKYLKAVL----------RQDVG 115
Query: 132 YYK*HYYCSR----------------CSIREGNFLHYISRFIAGFTIGFVRVWQISLVTL 175
Y+ H + S + NFL S F+A + +GF+ +W++++V
Sbjct: 116 YFDLHVTSTSDVITSVSSDSLVIQDFLSEKLPNFLMNTSAFVASYIVGFLLLWRLTIVGF 175
Query: 176 SIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKA 235
+ + + G Y I + K+R+ Y AG IAE+VI +VRTV AF E++ + +
Sbjct: 176 PFIILLLIPGLMYGRALIRISMKIREEYNEAGSIAEQVISSVRTVYAFGSEKKMIEKFST 235
Query: 236 ALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVV 295
AL + G + GLAKG+ +GS + + + W L WY S +V + + GG + ++ V
Sbjct: 236 ALQGSVKLGLRQGLAKGIAIGS-NGITYAIWGFLTWYGSRMVMNHGSKGGTVSSVIVCVT 294
Query: 296 ISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCF 355
G SLGQ+ ++ F A I ++I R + G+ L K G ++FN V F
Sbjct: 295 FGGTSLGQSLSNLKYFSEAFVVGERIMKVINRVPGIDSDNLEGQILEKTRGEVEFNHVKF 354
Query: 356 SYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIR 415
+YPSRP+ IF +L L +P+GK VALVGGSGSGKSTV+SL++RFY+PI+G+IL+D I
Sbjct: 355 TYPSRPETPIFDDLCLRVPSGKTVALVGGSGSGKSTVISLLQRFYDPIAGEILIDGLPIN 414
Query: 416 ELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
+L +KWLR Q+GLV+QEP LFATSIKENIL+GK+DA+++E+ A K
Sbjct: 415 KLQVKWLRSQMGLVSQEPVLFATSIKENILFGKEDASMDEVVEAAK 460
Score = 176 bits (447), Expect = 2e-44
Identities = 98/294 (33%), Positives = 155/294 (52%)
Query: 162 IGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQ 221
IG V W+ S+V +S+ P I + + + K + ++A E + N+RT+
Sbjct: 794 IGLVISWRFSIVMMSVQPVIVVCFYTQRVLLKSMSRNAIKGQDESSKLAAEAVSNIRTIT 853
Query: 222 AFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNI 281
AF+ +ER + K + + G+ LG+ ++ AL WY ++
Sbjct: 854 AFSSQERIINLLKMVQEGPRKDSARQSWLAGIMLGTSQSLITCVSALNFWYGGKLIADGK 913
Query: 282 ANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKL 341
E L +G + +A ++ A +F +++R+T + + G
Sbjct: 914 MMSKEFLEIFLIFASTGRVIAEAGTMTKDLVKGSDAVASVFAVLDRNTTIEPENPDGYVP 973
Query: 342 SKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYE 401
K+ G I F++V F+YP+RPDV IF N ++DI GK A+VG SGSGKST++SLIERFY+
Sbjct: 974 KKVKGQISFSNVDFAYPTRPDVIIFQNFSIDIEDGKSTAIVGPSGSGKSTIISLIERFYD 1033
Query: 402 PISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEE 455
P+ G + +D DIR L+ LRQ I LV+QEP LFA +I+ENI+YG ++E
Sbjct: 1034 PLKGIVKIDGRDIRSCHLRSLRQHIALVSQEPTLFAGTIRENIMYGGASNKIDE 1087
>At3g28345 P-glycoprotein, putative, 3'partial
Length = 610
Score = 280 bits (717), Expect = 9e-76
Identities = 165/480 (34%), Positives = 261/480 (54%), Gaps = 32/480 (6%)
Query: 2 ESKEGDERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIG 61
E KE K S+ +F AD D++LM +G IGA+ G + P+ + KL+N IG
Sbjct: 5 EEKESGRNKMNCFGSVRSIFMHADGVDWLLMGLGLIGAVGDGFTTPLVLLITSKLMNNIG 64
Query: 62 LAYLFPKEASHKVAKSKCSNIIFILDRGGL---LDAY-WRKTSSKDENGLFKVYVKSRYK 117
+ ++K+ + ++++ + L+ Y W +T + + + Y+++
Sbjct: 65 GSSFNTDTFMQSISKNSVA-LLYVACGSWVVCFLEGYCWTRTGERQTARMREKYLRAVL- 122
Query: 118 FI*Y*GIHWRGHFCYYK*HYYCSR----------------CSIREGNFLHYISRFIAGFT 161
R Y+ H + S + NFL S F+ +
Sbjct: 123 ---------RQDVGYFDLHVTSTSDVITSVSSDSFVIQDVLSEKLPNFLMSASTFVGSYI 173
Query: 162 IGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQ 221
+GF+ +W++++V L + + + G Y I + K+R+ Y AG +AE+ I +VRTV
Sbjct: 174 VGFILLWRLAIVGLPFIVLLVIPGLMYGRALISISRKIREEYNEAGFVAEQAISSVRTVY 233
Query: 222 AFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNI 281
AF+GE + + + AL + G K GLAKG+ +GS + + F W + WY S +V +
Sbjct: 234 AFSGERKTISKFSTALQGSVKLGIKQGLAKGITIGS-NGITFAMWGFMSWYGSRMVMYHG 292
Query: 282 ANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKL 341
A GG F + I G+SLG ++ F A + I E+I R + G KL
Sbjct: 293 AQGGTVFAVAAAIAIGGVSLGGGLSNLKYFFEAASVGERIMEVINRVPKIDSDNPDGHKL 352
Query: 342 SKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYE 401
K+ G ++F +V F YPSR + IF + L +P+GK VALVGGSGSGKSTV+SL++RFY+
Sbjct: 353 EKIRGEVEFKNVKFVYPSRLETSIFDDFCLRVPSGKTVALVGGSGSGKSTVISLLQRFYD 412
Query: 402 PISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
P++G+IL+D I +L +KWLR Q+GLV+QEPALFAT+IKENIL+GK+DA+++++ A K
Sbjct: 413 PLAGEILIDGVSIDKLQVKWLRSQMGLVSQEPALFATTIKENILFGKEDASMDDVVEAAK 472
>At3g28380 P-glycoprotein, putative
Length = 1240
Score = 273 bits (699), Expect = 1e-73
Identities = 166/479 (34%), Positives = 257/479 (52%), Gaps = 30/479 (6%)
Query: 2 ESKEGDERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIG 61
+ KE K + S+ +F AD D++LM +G IGA+ G P+ F L+N +G
Sbjct: 5 DEKESGRDKMKSFGSIRSIFMHADGVDWILMALGLIGAVGDGFITPVVVFIFNTLLNNLG 64
Query: 62 LAYLFPKEASHKVAKSKCSNIIFILDRGGL--LDAY-WRKTSSKDENGLFKVYVKSRYKF 118
+ K ++K+ + + + L+ Y W +T + + + Y+++
Sbjct: 65 TSSSNNKTFMQTISKNVVALLYVACGSWVICFLEGYCWTRTGERQAARMREKYLRAVL-- 122
Query: 119 I*Y*GIHWRGHFCYYK*HYYCSR----------------CSIREGNFLHYISRFIAGFTI 162
R Y+ H + S + NFL S F+A + +
Sbjct: 123 --------RQDVGYFDLHVTSTSDVITSISSDSLVIQDFLSEKLPNFLMNASAFVASYIV 174
Query: 163 GFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQA 222
F+ +W++++V + + + G Y + + K+ + Y AG IAE+ I +VRTV A
Sbjct: 175 SFILMWRLTIVGFPFIILLLVPGLMYGRALVSISRKIHEQYNEAGSIAEQAISSVRTVYA 234
Query: 223 FAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIA 282
F E + + + AL + G + GLAKG+ +GS + V WA L WY S +V + +
Sbjct: 235 FGSENKMIGKFSTALRGSVKLGLRQGLAKGITIGS-NGVTHAIWAFLTWYGSRLVMNHGS 293
Query: 283 NGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLS 342
GG F + + G+SLGQ+ ++ F A A I E+I+R + K G+ L
Sbjct: 294 KGGTVFVVISCITYGGVSLGQSLSNLKYFSEAFVAWERILEVIKRVPDIDSNKKEGQILE 353
Query: 343 KLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEP 402
++ G ++FN V F+Y SRP+ IF +L L IPAGK VALVGGSGSGKSTV+SL++RFY+P
Sbjct: 354 RMKGEVEFNHVKFTYLSRPETTIFDDLCLKIPAGKTVALVGGSGSGKSTVISLLQRFYDP 413
Query: 403 ISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
I+G+IL+D I +L + WLR Q+GLV+QEP LFATSI ENIL+GK+DA+L+E+ A K
Sbjct: 414 IAGEILIDGVSIDKLQVNWLRSQMGLVSQEPVLFATSITENILFGKEDASLDEVVEAAK 472
Score = 176 bits (447), Expect = 2e-44
Identities = 102/303 (33%), Positives = 158/303 (51%)
Query: 153 ISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEE 212
IS I IG V W++++V +S+ P I + + L K KA + ++A E
Sbjct: 800 ISAVIIACIIGLVIAWRLAIVMISVQPLIVVCFYTQRVLLKSLSEKASKAQDESSKLAAE 859
Query: 213 VIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWY 272
+ N+RT+ AF+ +ER ++ K G+ LG+ ++ + AL WY
Sbjct: 860 AVSNIRTITAFSSQERIIKLLKKVQEGPRRESVHRSWLAGIVLGTSRSLITCTSALNFWY 919
Query: 273 TSVVVHKNIANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSK 332
++ F L V +G + A + R A +F +++R T +
Sbjct: 920 GGRLIADGKIVSKAFFEIFLIFVTTGRVIADAGTMTTDLARGLDAVGSVFAVLDRCTTIE 979
Query: 333 KSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTV 392
+ G K+ G I F +V F+YP+RPDV IF N +++I GK A+VG SGSGKST+
Sbjct: 980 PKNPDGYVAEKIKGQITFLNVDFAYPTRPDVVIFENFSIEIDEGKSTAIVGTSGSGKSTI 1039
Query: 393 VSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDAT 452
+ LIERFY+P+ G + +D DIR L+ LR+ I LV+QEP LFA +I+ENI+YG
Sbjct: 1040 IGLIERFYDPLKGTVKIDGRDIRSYHLRSLRKYISLVSQEPMLFAGTIRENIMYGGTSDK 1099
Query: 453 LEE 455
++E
Sbjct: 1100 IDE 1102
>At3g28415 putative protein
Length = 1098
Score = 271 bits (692), Expect = 7e-73
Identities = 170/457 (37%), Positives = 248/457 (54%), Gaps = 22/457 (4%)
Query: 16 SMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLFPKEASHKVA 75
S+ +F A+S D VLM +G IGA+ G PI F G L+N IG + K H +
Sbjct: 6 SVRSIFMHANSVDLVLMGLGLIGAVGDGFITPIIFFITGLLLNDIGDSSFGDKTFMHAIM 65
Query: 76 KSKCSN---------IIFILDRGG--LLDAYWRKTSSKDENGLFKVYVKSRYKFI*Y*GI 124
K+ + I F+ +R + + Y R +D G F ++V S I
Sbjct: 66 KNAVALLYVAGASLVICFVGERQASRMREKYLRAVLRQDV-GYFDLHVTSTSDVITSVSS 124
Query: 125 HWRGHFCYYK*HYYCSRCSIREGNFLHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALA 184
S + NFL S F+A + +GF+ +W++++V + +
Sbjct: 125 DTL---------VIQDVLSEKLPNFLMSASAFVASYIVGFIMLWRLTIVGFPFFILLLIP 175
Query: 185 GGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNG 244
G I + K+R+ Y AG IAE+ I VRTV AF E + + + AAL + G
Sbjct: 176 GLMCGRALINISRKIREEYNEAGSIAEQAISLVRTVYAFGSERKMISKFSAALEGSVKLG 235
Query: 245 RKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQA 304
+ G+AKG+ +GS + V + W + WY S +V + A GG F ++ + G SLG+
Sbjct: 236 LRQGIAKGIAIGS-NGVTYAIWGFMTWYGSRMVMYHGAKGGTIFAVIICITYGGTSLGRG 294
Query: 305 APDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVG 364
++ F A A I E+I+R + G+ L + G +QF V F Y SRP+
Sbjct: 295 LSNLKYFSEAVVAGERIIEVIKRVPDIDSDNPRGQVLENIKGEVQFKHVKFMYSSRPETP 354
Query: 365 IFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELDLKWLRQ 424
IF +L L IP+GK VALVGGSGSGKSTV+SL++RFY+PI G+IL+D I++L +KWLR
Sbjct: 355 IFDDLCLRIPSGKSVALVGGSGSGKSTVISLLQRFYDPIVGEILIDGVSIKKLQVKWLRS 414
Query: 425 QIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
Q+GLV+QEPALFATSI+ENIL+GK+DA+ +E+ A K
Sbjct: 415 QMGLVSQEPALFATSIEENILFGKEDASFDEVVEAAK 451
Score = 170 bits (431), Expect = 1e-42
Identities = 101/306 (33%), Positives = 159/306 (51%), Gaps = 6/306 (1%)
Query: 153 ISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCY---AYVTIGLIAKVRKAYVRAGEI 209
IS T+G W++S+V ++I P + GC+ V + K KA + ++
Sbjct: 781 ISAVSVACTLGLAISWKLSIVMIAIQPVVV---GCFYTQRIVLKSISKKAIKAQDESSKL 837
Query: 210 AEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALL 269
A E + N+RT+ AF+ +ER ++ K + G+ L + ++ + AL
Sbjct: 838 AAEAVSNIRTITAFSSQERILKLLKMVQEGPQRENIRQSWLAGIVLATSRSLMTCTSALN 897
Query: 270 VWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDT 329
WY + ++ F + V +G + A + A +F +++R T
Sbjct: 898 YWYGARLIIDGKITSKAFFELFILFVSTGRVIADAGAMTMDLAKGSDAVGSVFAVLDRYT 957
Query: 330 VSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGK 389
+ G + G I+F +V F+YP+RPDV IF N ++DI GK A+VG SGSGK
Sbjct: 958 NIEPEKPDGFVPQNIKGQIKFVNVDFAYPTRPDVIIFKNFSIDIDEGKSTAIVGPSGSGK 1017
Query: 390 STVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKD 449
ST++ LIERFY+P+ G + +D DIR L+ LRQ IGLV+QEP LFA +I+ENI+YG
Sbjct: 1018 STIIGLIERFYDPLKGIVKIDGRDIRSYHLRSLRQHIGLVSQEPILFAGTIRENIMYGGA 1077
Query: 450 DATLEE 455
++E
Sbjct: 1078 SDKIDE 1083
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.327 0.142 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,718,138
Number of Sequences: 26719
Number of extensions: 402079
Number of successful extensions: 1602
Number of sequences better than 10.0: 96
Number of HSP's better than 10.0 without gapping: 78
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 1377
Number of HSP's gapped (non-prelim): 189
length of query: 461
length of database: 11,318,596
effective HSP length: 103
effective length of query: 358
effective length of database: 8,566,539
effective search space: 3066820962
effective search space used: 3066820962
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC087771.6


