

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149581.7 + phase: 0
(392 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
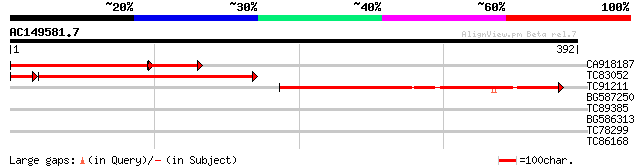
Score E
Sequences producing significant alignments: (bits) Value
CA918187 weakly similar to GP|4063748|gb| unknown protein {Arabi... 213 1e-72
TC83052 similar to GP|4063748|gb|AAC98456.1| unknown protein {Ar... 171 7e-50
TC91211 weakly similar to GP|4096756|gb|AAD10477.1| alpha-1 3/4-... 154 6e-38
BG587250 weakly similar to GP|3947735|emb| NL27 {Solanum tuberos... 32 0.55
TC89385 similar to GP|19774173|gb|AAL99063.1 NBS-LRR-Toll resist... 30 2.1
BG586313 29 3.6
TC78299 similar to GP|18086379|gb|AAL57649.1 AT3g16910/K14A17_3 ... 29 3.6
TC86168 homologue to GP|13936812|gb|AAK49947.1 TGF-beta receptor... 28 8.0
>CA918187 weakly similar to GP|4063748|gb| unknown protein {Arabidopsis
thaliana}, partial (83%)
Length = 720
Score = 213 bits (541), Expect(2) = 1e-72
Identities = 98/99 (98%), Positives = 98/99 (98%)
Frame = +2
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 60
MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH
Sbjct: 323 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 502
Query: 61 DSRYGHDLLYNEYYLAQLQELLKKYQDVREIWFDGAKDP 99
DSRYGHDLLYNEYYLAQLQELLKKYQDVREIWFDGAK P
Sbjct: 503 DSRYGHDLLYNEYYLAQLQELLKKYQDVREIWFDGAKIP 619
Score = 78.6 bits (192), Expect(2) = 1e-72
Identities = 36/38 (94%), Positives = 38/38 (99%)
Frame = +1
Query: 96 AKDPRAQNVTYYFSDWFSMVKELQSSINIFSDAGPDVR 133
+KDPRAQNVTYYFSDWFSMVKELQSSINIFS+AGPDVR
Sbjct: 607 SKDPRAQNVTYYFSDWFSMVKELQSSINIFSNAGPDVR 720
>TC83052 similar to GP|4063748|gb|AAC98456.1| unknown protein {Arabidopsis
thaliana}, partial (86%)
Length = 877
Score = 171 bits (433), Expect(2) = 7e-50
Identities = 77/152 (50%), Positives = 104/152 (67%), Gaps = 1/152 (0%)
Frame = +3
Query: 21 HSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRHDSRYGHDLLYNEYYLAQLQE 80
+SV SS W+NG GDVV + AA + G+ GIYLSPWDRH+ YG L YN++YLAQ+ E
Sbjct: 420 YSVRSSGWRNGNGDVVADVAAAAGEAGVGFGIYLSPWDRHELCYGDTLRYNQHYLAQMTE 599
Query: 81 LLKKYQDVREIWFDGAKDPRAQNVTYYFSDWFSMVKELQSSINIFSDAGPDVRWVGDETG 140
LL +Y ++++++ DGAK +N+ Y+F WFS++ +LQ IFSD+GPD RWVG+E G
Sbjct: 600 LLTRYGEIKDVFLDGAKGEGEKNMKYFFESWFSLIHQLQPGAAIFSDSGPDTRWVGNEQG 779
Query: 141 TAGDTCWSTINRTSLSI-GASNITQYLNTGDP 171
AG TCWS N++ + I G N QY GDP
Sbjct: 780 VAGSTCWSLFNQSVIEIGGVDNDPQYQKQGDP 875
Score = 44.3 bits (103), Expect(2) = 7e-50
Identities = 16/19 (84%), Positives = 19/19 (99%)
Frame = +2
Query: 1 MILTAKHHDGFCLWPSKYT 19
++LTAKHHDGFCLWPS+YT
Sbjct: 359 VLLTAKHHDGFCLWPSEYT 415
>TC91211 weakly similar to GP|4096756|gb|AAD10477.1| alpha-1 3/4-fucosidase
precursor {Streptomyces sp.}, partial (8%)
Length = 993
Score = 154 bits (389), Expect = 6e-38
Identities = 87/203 (42%), Positives = 125/203 (60%), Gaps = 6/203 (2%)
Frame = +1
Query: 187 PGWFWHKSESPKKLSDLLDIYYKSVGRNCVLLLNVPPNTTGLISENDAHRLKEFRSAIDT 246
PGWFWH SE PK LL+IYYKSVGRNC LLLNVPPN++GLIS D L+EF +
Sbjct: 1 PGWFWHASEHPKSARKLLEIYYKSVGRNCKLLLNVPPNSSGLISAEDIQVLREFSELRHS 180
Query: 247 IFHKNIAENRYVKVSSQRGG-KEGGFGPENMLDSDHLWSYWTPREDDKEKDHWIEIWGND 305
IF N A + + SS RGG ++ F P +L+ + + +YW P E+ + WI
Sbjct: 181 IFSHNFAASASLNASSTRGGIQDTRFSPNKVLE-EGIHTYWAPEENQSK---WILYINLK 348
Query: 306 GSLRFNVIRIQEAIGLGQRIERYEIYV---DG--KSIIQGTTIGYKRLHRLDGDVVHARV 360
+ FNV+++QE I +GQR+ ++ + DG KS++ GTTIGY+RL L + ++
Sbjct: 349 KLVSFNVLQVQEPIHMGQRVIKFHLEALNRDGLWKSVVNGTTIGYQRL--LLFPKLKSQY 522
Query: 361 VRIRFIKARGVPLISSIGLHFDP 383
+++ K+R PLIS +G++ DP
Sbjct: 523 LKLVVDKSRAEPLISYLGIYLDP 591
>BG587250 weakly similar to GP|3947735|emb| NL27 {Solanum tuberosum}, partial
(9%)
Length = 655
Score = 31.6 bits (70), Expect = 0.55
Identities = 16/54 (29%), Positives = 25/54 (45%)
Frame = -2
Query: 10 GFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRHDSR 63
GF LW S+ + ++ K + + + T +G I +SPW RH SR
Sbjct: 198 GFTLWKICVIPESINTFSFKGSKKKISEIVASIFTSEGDKHVITISPWQRHQSR 37
>TC89385 similar to GP|19774173|gb|AAL99063.1 NBS-LRR-Toll resistance gene
analog protein {Medicago ruthenica}, partial (98%)
Length = 1285
Score = 29.6 bits (65), Expect = 2.1
Identities = 15/53 (28%), Positives = 24/53 (44%)
Frame = -3
Query: 10 GFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRHDS 62
GF LW S+ + ++ K + + + T +G I +SPW RH S
Sbjct: 197 GFTLWKICVIPESINTFSFKGSKKKISEIVASIFTSEGDKHVITISPWQRHQS 39
>BG586313
Length = 553
Score = 28.9 bits (63), Expect = 3.6
Identities = 16/50 (32%), Positives = 23/50 (46%), Gaps = 3/50 (6%)
Frame = -1
Query: 168 TGDPKGTDWLPAECDVSIRPGWFWHK---SESPKKLSDLLDIYYKSVGRN 214
TG+P G W+ SIRP W + S K LSD++ + R+
Sbjct: 178 TGNPSGDSWVSCVSRSSIRPATAWKSFSDAGSTKTLSDVV*VILNRTDRD 29
>TC78299 similar to GP|18086379|gb|AAL57649.1 AT3g16910/K14A17_3
{Arabidopsis thaliana}, partial (46%)
Length = 889
Score = 28.9 bits (63), Expect = 3.6
Identities = 29/95 (30%), Positives = 46/95 (47%), Gaps = 6/95 (6%)
Frame = +2
Query: 110 DWFSMVKELQSSINIFSDA-----GPDVRWVGDETGTAGDTCWSTINRTSLSIGASNITQ 164
++FS+ +E ++ I+SD P + +GDE C R +LS GA
Sbjct: 413 EFFSLAEE---ALKIWSDKTKTFKSPILIVIGDEN------CDPKSLRYALSKGAVEYED 565
Query: 165 YLNTGDPKGTDWLPAECD-VSIRPGWFWHKSESPK 198
+L +GDP+ +W P E + SI G+ + SPK
Sbjct: 566 FLRSGDPE-YNWKPPEDEWQSIALGYTSGTTASPK 667
>TC86168 homologue to GP|13936812|gb|AAK49947.1 TGF-beta receptor-interacting
protein 1 {Phaseolus vulgaris}, complete
Length = 1322
Score = 27.7 bits (60), Expect = 8.0
Identities = 12/28 (42%), Positives = 15/28 (52%)
Frame = +2
Query: 333 DGKSIIQGTTIGYKRLHRLDGDVVHARV 360
DGKS G GY RLH D D + ++
Sbjct: 1064 DGKSFSSGGEDGYVRLHHFDPDYFNIKI 1147
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.138 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,721,039
Number of Sequences: 36976
Number of extensions: 193047
Number of successful extensions: 934
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 926
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 931
length of query: 392
length of database: 9,014,727
effective HSP length: 98
effective length of query: 294
effective length of database: 5,391,079
effective search space: 1584977226
effective search space used: 1584977226
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC149581.7


