

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
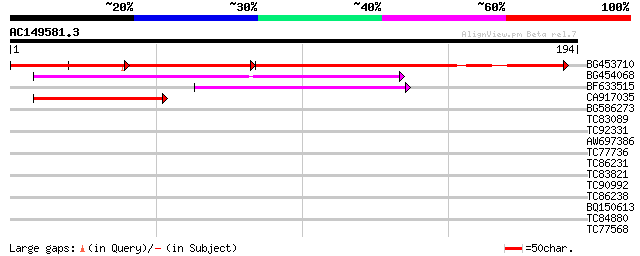
Query= AC149581.3 + phase: 0
(194 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG453710 weakly similar to GP|21537057|gb| unknown {Arabidopsis ... 102 7e-44
BG454068 weakly similar to GP|22202804|dbj hypothetical protein~... 109 8e-25
BF633515 52 2e-07
CA917035 similar to GP|7303305|gb|A CG6061-PA {Drosophila melano... 46 8e-06
BG586273 weakly similar to PIR|F86470|F8 probable retroelement p... 29 1.3
TC83089 28 2.2
TC92331 similar to PIR|B86447|B86447 hypothetical protein AAF813... 28 2.9
AW697386 similar to SP|P53535|PHS2 Alpha-1 4 glucan phosphorylas... 28 2.9
TC77736 similar to GP|12324651|gb|AAG52287.1 putative GTP-bindin... 27 3.8
TC86231 similar to GP|21593313|gb|AAM65262.1 protein disulfide i... 27 5.0
TC83821 similar to GP|8953400|emb|CAB96673.1 1-D-deoxyxylulose 5... 27 5.0
TC90992 similar to PIR|F86242|F86242 unknown protein 98896-9585... 27 6.5
TC86238 similar to GP|21536871|gb|AAM61203.1 unknown {Arabidopsi... 26 8.5
BQ150613 similar to GP|10728365|gb CG18830 gene product {Drosoph... 26 8.5
TC84880 similar to PIR|T01491|T01491 ubiquitin--protein ligase h... 26 8.5
TC77568 weakly similar to GP|9857516|gb|AAG00871.1| Unknown prot... 26 8.5
>BG453710 weakly similar to GP|21537057|gb| unknown {Arabidopsis thaliana},
partial (6%)
Length = 651
Score = 102 bits (255), Expect(2) = 7e-44
Identities = 57/107 (53%), Positives = 72/107 (67%)
Frame = -3
Query: 85 AMNQDCAIDLMIRLLGMSEVDVRVEVRSESAGHISYPTLKRVYEHHLTEARRLEEPQTRE 144
AM Q AIDLM RLLGMS+ + E+R+ESAGHISYPTLKRVYE HL EARRLE+P TRE
Sbjct: 325 AMGQ*HAIDLMTRLLGMSDANAMAEIRTESAGHISYPTLKRVYEDHLIEARRLEDPLTRE 146
Query: 145 KLQERGRKRAIGKWSWGGMALAYLYDYLDDSVILNNRTMAGSTILFM 191
+LQER R+R +W + L YL D V+ +T ++++
Sbjct: 145 ELQERARRR---QWCVRSLLL-----YLVDCVLFTYKTNRHIDLIYL 29
Score = 91.3 bits (225), Expect(2) = 7e-44
Identities = 44/64 (68%), Positives = 50/64 (77%)
Frame = -1
Query: 21 FDLHDLIYTGYSTVTHAMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGRIL 80
FD+H + Y S MI AMCERWHTETSSFHL +GEMTI LDD++NLLHIPIHG +L
Sbjct: 501 FDIHGVFYRDPS-----MIRAMCERWHTETSSFHLPMGEMTITLDDVYNLLHIPIHGCML 337
Query: 81 DHDE 84
DHDE
Sbjct: 336 DHDE 325
Score = 63.9 bits (154), Expect = 4e-11
Identities = 32/43 (74%), Positives = 34/43 (78%), Gaps = 2/43 (4%)
Frame = -2
Query: 1 MKELGKPEAGLRWFWEPVEGFDLHDLIYTGYSTVTHA--MICA 41
MKELGKPEA LRWF E VEG LHDLIYTGY TVTH ++CA
Sbjct: 575 MKELGKPEACLRWFLELVEGSGLHDLIYTGYFTVTHP*YVLCA 447
>BG454068 weakly similar to GP|22202804|dbj hypothetical protein~predicted by
GeneMark.hmm etc. {Oryza sativa (japonica
cultivar-group)}, partial (5%)
Length = 673
Score = 109 bits (272), Expect = 8e-25
Identities = 52/127 (40%), Positives = 77/127 (59%)
Frame = +3
Query: 9 AGLRWFWEPVEGFDLHDLIYTGYSTVTHAMICAMCERWHTETSSFHLLVGEMTIPLDDMH 68
A + WFW + L+ L+ T Y V H ++ A RWH+ETSSFHL VGEMTI LDD+
Sbjct: 249 AAMGWFWNVIRASGLYPLLETNYGQVDHGLLIAFSLRWHSETSSFHLPVGEMTIALDDIS 428
Query: 69 NLLHIPIHGRILDHDEAMNQDCAIDLMIRLLGMSEVDVRVEVRSESAGHISYPTLKRVYE 128
LLHIP+ G +L H E+++ + ++ LG+ ++ E + + HI+Y TL +Y
Sbjct: 429 CLLHIPVGGNLLFH-ESLSIHQGTEYLVNYLGLEFEEIAAETKRLKSAHITYDTLLSIYT 605
Query: 129 HHLTEAR 135
+LTEA+
Sbjct: 606 SYLTEAK 626
>BF633515
Length = 249
Score = 51.6 bits (122), Expect = 2e-07
Identities = 28/74 (37%), Positives = 41/74 (54%)
Frame = +2
Query: 64 LDDMHNLLHIPIHGRILDHDEAMNQDCAIDLMIRLLGMSEVDVRVEVRSESAGHISYPTL 123
LDD+ L H+PI GR+L H + M + L++R LG+S+ + E G+ISYP L
Sbjct: 2 LDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEYGGYISYPRL 181
Query: 124 KRVYEHHLTEARRL 137
+ Y +L A L
Sbjct: 182 RDFYTTYLGRANLL 223
>CA917035 similar to GP|7303305|gb|A CG6061-PA {Drosophila melanogaster},
partial (2%)
Length = 365
Score = 46.2 bits (108), Expect = 8e-06
Identities = 19/46 (41%), Positives = 28/46 (60%)
Frame = +2
Query: 9 AGLRWFWEPVEGFDLHDLIYTGYSTVTHAMICAMCERWHTETSSFH 54
A + FW + L+ L+ T Y V H+++ A +RWH+ETSSFH
Sbjct: 227 AAMGRFWNVIMASGLYLLLETNYGQVDHSLLIAFSDRWHSETSSFH 364
>BG586273 weakly similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 705
Score = 28.9 bits (63), Expect = 1.3
Identities = 16/48 (33%), Positives = 28/48 (58%)
Frame = -2
Query: 103 EVDVRVEVRSESAGHISYPTLKRVYEHHLTEARRLEEPQTREKLQERG 150
E+ ++E RS A I Y T ++ Y+ + EARR+ + + ++ERG
Sbjct: 569 ELRNKLEARSRKAMFIGYSTTQKGYKCYDPEARRVLVSRDVKFIEERG 426
>TC83089
Length = 887
Score = 28.1 bits (61), Expect = 2.2
Identities = 11/16 (68%), Positives = 12/16 (74%)
Frame = +3
Query: 159 SWGGMALAYLYDYLDD 174
SWGGM LAYLY L +
Sbjct: 486 SWGGMTLAYLYHCLSE 533
>TC92331 similar to PIR|B86447|B86447 hypothetical protein AAF81322.1
[imported] - Arabidopsis thaliana, partial (48%)
Length = 921
Score = 27.7 bits (60), Expect = 2.9
Identities = 16/32 (50%), Positives = 17/32 (53%), Gaps = 3/32 (9%)
Frame = +2
Query: 123 LKRVYEHHLTEARRL---EEPQTREKLQERGR 151
LKR Y H L RRL PQTR+ L R R
Sbjct: 128 LKRTYRHSLRIQRRLITRNHPQTRQFLHRRTR 223
>AW697386 similar to SP|P53535|PHS2 Alpha-1 4 glucan phosphorylase L-2
isozyme chloroplast precursor (EC 2.4.1.1), partial
(14%)
Length = 556
Score = 27.7 bits (60), Expect = 2.9
Identities = 27/112 (24%), Positives = 50/112 (44%)
Frame = +3
Query: 39 ICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGRILDHDEAMNQDCAIDLMIRL 98
I A E+ T ++ L ++ + ++D H L IP RIL + ++ + A D+ R
Sbjct: 15 IIARFEKRSGMTVNWDSLPDKVVVQMNDTHPTLCIPELIRILIDVKGLSWEKAWDITKRT 194
Query: 99 LGMSEVDVRVEVRSESAGHISYPTLKRVYEHHLTEARRLEEPQTREKLQERG 150
+ + V E + S L+ + H+ +R++E T E + E G
Sbjct: 195 VAYTNHTVLPEALEK----WSLTLLQDLLPRHVEIIKRIDEEFTHEIVSEYG 338
>TC77736 similar to GP|12324651|gb|AAG52287.1 putative GTP-binding protein;
106556-109264 {Arabidopsis thaliana}, partial (76%)
Length = 2145
Score = 27.3 bits (59), Expect = 3.8
Identities = 18/53 (33%), Positives = 29/53 (53%)
Frame = +3
Query: 87 NQDCAIDLMIRLLGMSEVDVRVEVRSESAGHISYPTLKRVYEHHLTEARRLEE 139
N D D++++ + VRV ++A HI LKRV + HLT A +++E
Sbjct: 1155 NNDSETDVVLKGV------VRVTNLKDAADHIG-EVLKRVKKEHLTRAYKIKE 1292
>TC86231 similar to GP|21593313|gb|AAM65262.1 protein disulfide isomerase
precursor-like {Arabidopsis thaliana}, partial (79%)
Length = 2185
Score = 26.9 bits (58), Expect = 5.0
Identities = 10/36 (27%), Positives = 19/36 (52%)
Frame = -3
Query: 155 IGKWSWGGMALAYLYDYLDDSVILNNRTMAGSTILF 190
+G WSW G+ +L + + + + T+ GS + F
Sbjct: 1727 VGFWSWNGIDACFLRNL*KATTVRSTSTVIGSELFF 1620
>TC83821 similar to GP|8953400|emb|CAB96673.1 1-D-deoxyxylulose 5-phosphate
synthase-like protein {Arabidopsis thaliana}, partial
(46%)
Length = 1172
Score = 26.9 bits (58), Expect = 5.0
Identities = 14/45 (31%), Positives = 20/45 (44%)
Frame = -2
Query: 31 YSTVTHAMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPI 75
Y + A +C H + S + E+ PLD L+HIPI
Sbjct: 946 YKAASEARCQGVCRTLHQQQSPQCQDLQEVNKPLDSSEPLIHIPI 812
>TC90992 similar to PIR|F86242|F86242 unknown protein 98896-95855
[imported] - Arabidopsis thaliana, partial (14%)
Length = 763
Score = 26.6 bits (57), Expect = 6.5
Identities = 17/51 (33%), Positives = 31/51 (60%), Gaps = 2/51 (3%)
Frame = +2
Query: 65 DDMHNLLHIPIHGR--ILDHDEAMNQDCAIDLMIRLLGMSEVDVRVEVRSE 113
D++ LHI I G +L H EA +++++LLG++EVD+ ++ +E
Sbjct: 329 DEIEAGLHIRIAGEKVVLFHQEAA------EVLLQLLGVAEVDLLLQNDTE 463
>TC86238 similar to GP|21536871|gb|AAM61203.1 unknown {Arabidopsis
thaliana}, partial (58%)
Length = 698
Score = 26.2 bits (56), Expect = 8.5
Identities = 13/35 (37%), Positives = 19/35 (54%), Gaps = 1/35 (2%)
Frame = +2
Query: 23 LHDLI-YTGYSTVTHAMICAMCERWHTETSSFHLL 56
LH L+ TG+ + +IC C RW+ TS L+
Sbjct: 272 LHLLLPVTGHRLIKE*LIC*CCWRWYLRTSFIRLI 376
>BQ150613 similar to GP|10728365|gb CG18830 gene product {Drosophila
melanogaster}, partial (1%)
Length = 1151
Score = 26.2 bits (56), Expect = 8.5
Identities = 23/87 (26%), Positives = 36/87 (40%), Gaps = 6/87 (6%)
Frame = +2
Query: 73 IPIHGRILDHDEAMNQDCAIDLMIRLLGMSEVDVRVEVR------SESAGHISYPTLKRV 126
IP+ + DE ID L ++D+ V+ R S ++ P L
Sbjct: 836 IPLSRTLARRDERWEDRVVIDY---LDSFYKIDMYVDCR*YHSERQTSLASVATPALCVC 1006
Query: 127 YEHHLTEARRLEEPQTREKLQERGRKR 153
+ H+T AR + QTR+ + E G R
Sbjct: 1007AKTHITRARVVSLNQTRQYIAEDGVDR 1087
>TC84880 similar to PIR|T01491|T01491 ubiquitin--protein ligase homolog
F17O7.15 - Arabidopsis thaliana, partial (17%)
Length = 801
Score = 26.2 bits (56), Expect = 8.5
Identities = 8/15 (53%), Positives = 11/15 (73%)
Frame = +1
Query: 11 LRWFWEPVEGFDLHD 25
++WFWE V+GF D
Sbjct: 304 IQWFWEVVQGFSKED 348
>TC77568 weakly similar to GP|9857516|gb|AAG00871.1| Unknown protein
{Arabidopsis thaliana}, partial (63%)
Length = 1206
Score = 26.2 bits (56), Expect = 8.5
Identities = 14/53 (26%), Positives = 29/53 (54%)
Frame = -1
Query: 54 HLLVGEMTIPLDDMHNLLHIPIHGRILDHDEAMNQDCAIDLMIRLLGMSEVDV 106
+L++ + T+ L D H++ + R DHD + + C++D + LG+ +V
Sbjct: 660 NLIIMDSTVLLGDRHHIDRYFCNFRDCDHDTVLIR-CSLDSKLIFLGLPRREV 505
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.139 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,067,003
Number of Sequences: 36976
Number of extensions: 79838
Number of successful extensions: 460
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 459
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 459
length of query: 194
length of database: 9,014,727
effective HSP length: 91
effective length of query: 103
effective length of database: 5,649,911
effective search space: 581940833
effective search space used: 581940833
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 56 (26.2 bits)
Medicago: description of AC149581.3


