

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149576.6 - phase: 0
(597 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
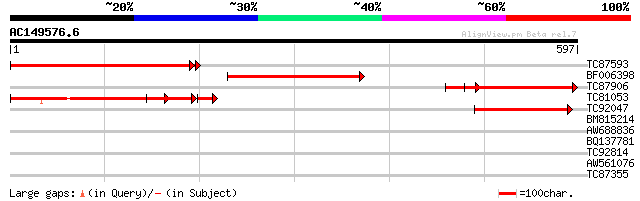
Sequences producing significant alignments: (bits) Value
TC87593 similar to GP|16604659|gb|AAL24122.1 unknown protein {Ar... 382 e-106
BF006398 similar to GP|10177495|db gb|AAC61821.1~gene_id:K21C13.... 283 1e-76
TC87906 243 2e-64
TC81053 similar to GP|16604659|gb|AAL24122.1 unknown protein {Ar... 209 2e-60
TC92047 69 4e-12
BM815214 similar to GP|22136992|gb unknown protein {Arabidopsis ... 32 0.90
AW688836 similar to GP|13872920|db hypothetical protein {Oryza s... 30 2.0
BQ137781 GP|14165182|gb zinc family member 5 protein {Homo sapie... 30 2.6
TC92814 weakly similar to PIR|G86333|G86333 hypothetical protein... 28 10.0
AW561076 weakly similar to GP|2262099|gb|A thaumatin isolog {Ara... 28 10.0
TC87355 similar to GP|2576411|gb|AAC61784.1| similar to dynamin-... 28 10.0
>TC87593 similar to GP|16604659|gb|AAL24122.1 unknown protein {Arabidopsis
thaliana}, partial (24%)
Length = 870
Score = 382 bits (982), Expect(2) = e-106
Identities = 186/194 (95%), Positives = 186/194 (95%)
Frame = +2
Query: 1 MNRNRLGLSAHHSGSTQSEESALDLERNYYGHPSSSPLHMQTFAVGVQHSEGNAAYFSWP 60
MNRNRLGLSAHHSGSTQSEESALDLERNYYGHPSSSPLHMQTFAVGVQHSEGNAAYFSWP
Sbjct: 260 MNRNRLGLSAHHSGSTQSEESALDLERNYYGHPSSSPLHMQTFAVGVQHSEGNAAYFSWP 439
Query: 61 TLNRWNDAAEDRANYFGNLQKGVLPETLGRLPSGQQATTLLELMTIRAFHSKILRRFSLG 120
TLNRWNDAAEDRANYFGNLQKGVLPETLGRLPSGQQATTLLELMTIRAFHSKILRRFSLG
Sbjct: 440 TLNRWNDAAEDRANYFGNLQKGVLPETLGRLPSGQQATTLLELMTIRAFHSKILRRFSLG 619
Query: 121 TAIGFRIRGGVLTDIPAILVFVAHKVHRQWLNHVQCLPAALEGPGGVWCDVDVVEFSYYG 180
TAIGFRIRG VLTDIPAILVFVAHKVHRQWLNHVQCLPAALEGPG VWCDVDVVEFS YG
Sbjct: 620 TAIGFRIRGSVLTDIPAILVFVAHKVHRQWLNHVQCLPAALEGPGSVWCDVDVVEFS*YG 799
Query: 181 APAPTPKEQLYTEL 194
APAPTPKE L
Sbjct: 800 APAPTPKEHYIRSL 841
Score = 21.2 bits (43), Expect(2) = e-106
Identities = 9/12 (75%), Positives = 10/12 (83%)
Frame = +1
Query: 190 LYTELADGLRGS 201
LYTELAD RG+
Sbjct: 826 LYTELADA*RGA 861
>BF006398 similar to GP|10177495|db gb|AAC61821.1~gene_id:K21C13.22~strong
similarity to unknown protein {Arabidopsis thaliana},
partial (22%)
Length = 434
Score = 283 bits (724), Expect = 1e-76
Identities = 138/144 (95%), Positives = 141/144 (97%)
Frame = +1
Query: 230 NREVGFLTNRHVAVDLDYPNQKMFHPLPPSLGPGVYLGAVERATSFITDDLWYGIFAGTN 289
+R + F +NRHVAVDLDYPNQKMFHPLPPSLGPGVYLGAVERATSFITDDLWYGIFAGTN
Sbjct: 1 SRMLVFYSNRHVAVDLDYPNQKMFHPLPPSLGPGVYLGAVERATSFITDDLWYGIFAGTN 180
Query: 290 PETFVRADGAFIPFAEDFNMNNVITSIRGVGDIGEVHRIDLQSPINSLIGRQVIKVGRSS 349
PETFVRADGAFIPFAEDFNMNNVITSIRGVGDIGEVHRIDLQSPINSLIGRQVIKVGRSS
Sbjct: 181 PETFVRADGAFIPFAEDFNMNNVITSIRGVGDIGEVHRIDLQSPINSLIGRQVIKVGRSS 360
Query: 350 GLTTGTIMAYALEYNDEKGICFLT 373
GLTTGTIMAYALEYNDEKGICFLT
Sbjct: 361 GLTTGTIMAYALEYNDEKGICFLT 432
>TC87906
Length = 817
Score = 243 bits (620), Expect = 2e-64
Identities = 118/118 (100%), Positives = 118/118 (100%)
Frame = +3
Query: 480 PTVPIKEKLEESFEPFCLNMEHVPVEEPSTIVKPSLRPCEFHIRNEIETVPNVEHQFIRT 539
PTVPIKEKLEESFEPFCLNMEHVPVEEPSTIVKPSLRPCEFHIRNEIETVPNVEHQFIRT
Sbjct: 63 PTVPIKEKLEESFEPFCLNMEHVPVEEPSTIVKPSLRPCEFHIRNEIETVPNVEHQFIRT 242
Query: 540 SFAGKSPVHQSFLKEDMQFKSLSELRNEPDEDNFVSLHLGEPEAKRRKHSNSSLSLKN 597
SFAGKSPVHQSFLKEDMQFKSLSELRNEPDEDNFVSLHLGEPEAKRRKHSNSSLSLKN
Sbjct: 243 SFAGKSPVHQSFLKEDMQFKSLSELRNEPDEDNFVSLHLGEPEAKRRKHSNSSLSLKN 416
Score = 43.9 bits (102), Expect = 2e-04
Identities = 24/37 (64%), Positives = 27/37 (72%), Gaps = 1/37 (2%)
Frame = +1
Query: 460 GQEQMNGSTAGIGSTVGE-SSPTVPIKEKLEESFEPF 495
GQEQMNGSTAGIGSTVGE SSP K+ L+ + F
Sbjct: 1 GQEQMNGSTAGIGSTVGESSSPLCQ*KKSLKRALSLF 111
>TC81053 similar to GP|16604659|gb|AAL24122.1 unknown protein {Arabidopsis
thaliana}, partial (33%)
Length = 879
Score = 209 bits (533), Expect(2) = 2e-60
Identities = 119/171 (69%), Positives = 126/171 (73%), Gaps = 3/171 (1%)
Frame = +3
Query: 1 MNRNRLGLSAHHSGSTQSEESALDLERNYYGH---PSSSPLHMQTFAVGVQHSEGNAAYF 57
M R RL SGST SEESALDLERN YGH PS SP +Q FA QH E NAAYF
Sbjct: 219 MERPRLNSRVRCSGSTPSEESALDLERNCYGHSNLPSLSPPTLQPFASAGQHGESNAAYF 398
Query: 58 SWPTLNRWNDAAEDRANYFGNLQKGVLPETLGRLPSGQQATTLLELMTIRAFHSKILRRF 117
SWP+ R DAAE+RANYF NLQKGVLPETLGRLP GQQATTLLELMTIRAFHSKILR +
Sbjct: 399 SWPS--RLPDAAEERANYFLNLQKGVLPETLGRLPKGQQATTLLELMTIRAFHSKILRCY 572
Query: 118 SLGTAIGFRIRGGVLTDIPAILVFVAHKVHRQWLNHVQCLPAALEGPGGVW 168
SLGTAIGFRIR GVLTDIPAILVFV+ KVH+ Q P + P G W
Sbjct: 573 SLGTAIGFRIRRGVLTDIPAILVFVSRKVHKAM---AQPNPVSSYCP*GAW 716
Score = 92.8 bits (229), Expect = 3e-19
Identities = 39/52 (75%), Positives = 44/52 (84%)
Frame = +1
Query: 145 KVHRQWLNHVQCLPAALEGPGGVWCDVDVVEFSYYGAPAPTPKEQLYTELAD 196
K +QWL+ +QCLP ALEGPGGVWCDVDVVEFSY+GAP P PKEQ YTE+ D
Sbjct: 655 KFTKQWLSPIQCLPTALEGPGGVWCDVDVVEFSYFGAPEPVPKEQHYTEIVD 810
Score = 42.0 bits (97), Expect(2) = 2e-60
Identities = 18/21 (85%), Positives = 19/21 (89%)
Frame = +2
Query: 198 LRGSDSCVGSGSQVASQETYG 218
LRG D C+GSGSQVASQETYG
Sbjct: 815 LRGGDPCIGSGSQVASQETYG 877
>TC92047
Length = 567
Score = 69.3 bits (168), Expect = 4e-12
Identities = 44/107 (41%), Positives = 66/107 (61%), Gaps = 4/107 (3%)
Frame = +2
Query: 490 ESFEPFCLNMEHVP--VEEPSTIVKPSLRPCEFHIRNEIETV-PNVEHQFIRTSFAGKSP 546
+ FEP L ++ +P VE S +KPS EF + + I+ P++EHQFI SF G+SP
Sbjct: 26 DKFEPLGLQIQSIPLGVEPNSQEMKPSTMEAEFKLEDGIKVGGPSIEHQFI-PSFIGRSP 202
Query: 547 VHQSFLKEDMQF-KSLSELRNEPDEDNFVSLHLGEPEAKRRKHSNSS 592
+H+ + + ++LS LRN+ +ED VSL LG+ EAKRR+ S+
Sbjct: 203 LHKHTVHDKAAAAENLSSLRNDCNEDICVSLQLGDNEAKRRRSEAST 343
>BM815214 similar to GP|22136992|gb unknown protein {Arabidopsis thaliana},
partial (24%)
Length = 596
Score = 31.6 bits (70), Expect = 0.90
Identities = 24/80 (30%), Positives = 38/80 (47%), Gaps = 3/80 (3%)
Frame = +1
Query: 204 CVGSGSQVASQETYGTLGAIVRSRTGNR-EVGFL-TNRHVAVDLDYPNQKMFHPLPPSLG 261
CV VA ++ + T + TG + + FL TN V ++ + H PPSL
Sbjct: 217 CVYEDGLVAFEDGFSTYADAIIHCTGYKYHIPFLETNGIVTIEDNRVGPLYKHVFPPSLA 396
Query: 262 PGV-YLGAVERATSFITDDL 280
PG+ ++G R T F+ +L
Sbjct: 397 PGLSFIGLTFRETIFVVIEL 456
>AW688836 similar to GP|13872920|db hypothetical protein {Oryza sativa
(japonica cultivar-group)}, partial (20%)
Length = 660
Score = 30.4 bits (67), Expect = 2.0
Identities = 23/79 (29%), Positives = 36/79 (45%), Gaps = 4/79 (5%)
Frame = +1
Query: 410 IIWGGTANRGRLKLRVGQPPENWTSGVDL----GRLLDLLELDLVTTNETLQDSGQEQMN 465
++W G +RG LR E W S + + G L+ LL L L +N + +
Sbjct: 184 LLWDGIPDRGASWLREAPIQETWKSVILVPAFGGLLVSLLNL-LSNSNSNSRPFLKAIAA 360
Query: 466 GSTAGIGSTVGESSPTVPI 484
T G G+++G P+V I
Sbjct: 361 SITLGTGNSLGPEGPSVDI 417
>BQ137781 GP|14165182|gb zinc family member 5 protein {Homo sapiens}, partial
(3%)
Length = 1213
Score = 30.0 bits (66), Expect = 2.6
Identities = 40/138 (28%), Positives = 56/138 (39%), Gaps = 19/138 (13%)
Frame = +2
Query: 85 PETLGRLPSGQQATTLLELMTIRAFHSK------ILRRFSLGTA--------IGFRIRGG 130
P L R PS AT + R S+ ++R S T I R+R
Sbjct: 809 PRALARSPSPPPATRACAISRCRVGISRAYSVTHVIRLRSACTTCSLA*SVRIDLRLR-- 982
Query: 131 VLT--DIPAILVFVAHKVHRQWLNHVQCLPAALEGPGGVWCD---VDVVEFSYYGAPAPT 185
LT D +I+ +H + R + L L GG+WC+ V ++ FS Y P PT
Sbjct: 983 -LTSDDTRSIISRYSHHIRRHDYVCPRVLATVLAWGGGLWCETRTVVMLAFSIY-FPIPT 1156
Query: 186 PKEQLYTELADGLRGSDS 203
+ LY A G+ S S
Sbjct: 1157VADSLY---AAGISSSQS 1201
>TC92814 weakly similar to PIR|G86333|G86333 hypothetical protein AAF79910.1
[imported] - Arabidopsis thaliana, partial (38%)
Length = 622
Score = 28.1 bits (61), Expect = 10.0
Identities = 12/35 (34%), Positives = 16/35 (45%)
Frame = +3
Query: 28 NYYGHPSSSPLHMQTFAVGVQHSEGNAAYFSWPTL 62
+YY H +SP H F VG + + WP L
Sbjct: 201 SYYKHKRNSPNHRLNFKVGKDFHRHSPRHLVWPFL 305
>AW561076 weakly similar to GP|2262099|gb|A thaumatin isolog {Arabidopsis
thaliana}, partial (12%)
Length = 412
Score = 28.1 bits (61), Expect = 10.0
Identities = 12/35 (34%), Positives = 16/35 (45%)
Frame = +2
Query: 28 NYYGHPSSSPLHMQTFAVGVQHSEGNAAYFSWPTL 62
+YY H +SP H F VG + + WP L
Sbjct: 155 SYYKHKRNSPNHRLNFKVGKDFHRHSPRHLVWPFL 259
>TC87355 similar to GP|2576411|gb|AAC61784.1| similar to dynamin-like
protein encoded by GenBank Accession Number X99669
{Arabidopsis thaliana}, partial (23%)
Length = 1317
Score = 28.1 bits (61), Expect = 10.0
Identities = 35/144 (24%), Positives = 57/144 (39%), Gaps = 23/144 (15%)
Frame = +1
Query: 450 VTTNETLQDSGQEQMNGSTAGIGSTVGESSPTVPIKEKLEESFEPF-CLNMEH----VPV 504
V+ E + SG + GS+ GI S G + ++ EPF + +E + +
Sbjct: 370 VSDAEKIASSGN--IGGSSWGISSIFGGGDKSARENVASKQHAEPFHSVEVEQSFSMIHL 543
Query: 505 EEPSTIVKPSLRPCEFH-----------------IRNEIET-VPNVEHQFIRTSFAGKSP 546
EP TI++PS E +R +E VP F+ + K
Sbjct: 544 TEPPTILRPSDSNSETEAVEITVTKLLLKSYYDIVRKNVEDFVPKAIMHFLVNNT--KRE 717
Query: 547 VHQSFLKEDMQFKSLSELRNEPDE 570
+H F+K+ + E+ EPDE
Sbjct: 718 LHNVFIKKLYRDNLFEEMLQEPDE 789
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.136 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,770,146
Number of Sequences: 36976
Number of extensions: 245552
Number of successful extensions: 1191
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1181
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1189
length of query: 597
length of database: 9,014,727
effective HSP length: 102
effective length of query: 495
effective length of database: 5,243,175
effective search space: 2595371625
effective search space used: 2595371625
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC149576.6


