

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149576.4 + phase: 0 /pseudo
(548 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
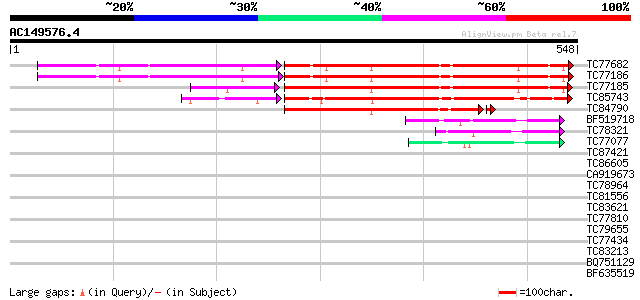
Score E
Sequences producing significant alignments: (bits) Value
TC77682 phosphate transporter 273 2e-83
TC77186 phosphate transporter 272 2e-83
TC77185 similar to GP|12697486|emb|CAC28219. phosphate transport... 270 3e-80
TC85743 phosphate transporter PT4 [Medicago truncatula] 219 1e-62
TC84790 similar to GP|13676624|gb|AAK38197.1 phosphate transport... 189 1e-48
BF519718 weakly similar to PIR|G84564|G84 probable sugar transpo... 60 2e-09
TC78321 similar to GP|21689841|gb|AAM67564.1 unknown protein {Ar... 51 1e-06
TC77077 similar to GP|9293909|dbj|BAB01812.1 sugar transporter p... 47 2e-05
TC87421 sugar transporter 37 0.015
TC86605 similar to PIR|T01853|T01853 probable hexose transport p... 37 0.020
CA919673 similar to GP|9858106|gb| putative glucose translocator... 36 0.044
TC78964 weakly similar to PIR|T00450|T00450 probable monosacchar... 36 0.044
TC81556 similar to GP|2688830|gb|AAB88879.1| putative sugar tran... 35 0.097
TC83621 similar to PIR|T01506|T01506 probable hexose transport p... 34 0.17
TC77810 similar to GP|13676624|gb|AAK38197.1 phosphate transport... 32 0.63
TC79655 similar to GP|10177966|dbj|BAB11349. uridine kinase-like... 32 0.82
TC77434 similar to GP|21554191|gb|AAM63270.1 unknown {Arabidopsi... 31 1.4
TC83213 weakly similar to GP|16945177|emb|CAC69071. STP5 protein... 30 1.8
BQ751129 weakly similar to GP|2258125|emb| AmMst-1 {Amanita musc... 30 2.4
BF635519 similar to GP|17064732|gb putative protein {Arabidopsis... 30 3.1
>TC77682 phosphate transporter
Length = 2245
Score = 273 bits (698), Expect(2) = 2e-83
Identities = 151/293 (51%), Positives = 191/293 (64%), Gaps = 13/293 (4%)
Frame = +2
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEEHPLPPATNV---AYPLLSREFLWRHGRDLF 322
YTALV +N QAA DM KVL V + EE + T+ +Y L S++F RHG LF
Sbjct: 842 YTALVAKNAKQAAADMSKVLQVELE--VEEEKVEKMTSDKRNSYGLFSKQFAARHGLALF 1015
Query: 323 ACSANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPG 380
+ WFLLDI FYSQ LFQ +I+ ++ + + E + +AR Q ++A+CST+PG
Sbjct: 1016 GTCSTWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPG 1195
Query: 381 YFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFF 440
Y+FTV FID +GR IQMMGFFFM V FAL PY HW+K EN GF+VIY L FFF
Sbjct: 1196 YWFTVAFIDHMGRFAIQMMGFFFMTVFMFALAIPY-DHWSKEENRI--GFVVIYSLTFFF 1366
Query: 441 ANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHK----EKEEGY 496
ANFGPN TTF+VPAE+FPAR RSTCHGIS A GK GAI+G+ GFL+A+ + ++GY
Sbjct: 1367 ANFGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDKGY 1546
Query: 497 PKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLE----ENEDEQSHHAEE 545
P GIG+K SLI+LG + VGM T E+ G+SLE ENE E + E+
Sbjct: 1547 PTGIGIKNSLIMLGVINFVGMLCT-LLVPESKGKSLEELSGENEGEGAEATEQ 1702
Score = 55.1 bits (131), Expect(2) = 2e-83
Identities = 79/245 (32%), Positives = 103/245 (41%), Gaps = 10/245 (4%)
Frame = +3
Query: 28 KIRKWQED*RFYQPLTQQKHNTTISRPSS*QEWDFSLTLMTFFPSHSSQRC*EEYTTPTT 87
+I+ QE * L +HN S EWD SL M F SQ C* EY T
Sbjct: 108 QIQPCQES*EC*MHLMWPRHNCITSLQL*LLEWDSSLMHMIFSAFPLSQSC*GEYIT--L 281
Query: 88 KKSVKSTRSWSQPLSPL----HS*EPPSVSSYSDASET*KAGATSTVSPYY*C*QAL*LL 143
+ + + Q L+ L H E + T G S V + + LL
Sbjct: 282 NQIQQDQEHFHQVLNQL*LVLHLLEH*QANYSLVGLATNLVGKKSMV*HLF-SWWFVQLL 458
Query: 144 ASPSVQQRELVFYYLLVSLDSS*GLELVVIILYLLPLCLSLLIKGQEVLL*LRFFLCKGL 203
+ Q + V + VSL L LVVI L+ L CL++L K EV L L F CK L
Sbjct: 459 QDSLLDQVQRVLWQHFVSLGFGLVLVLVVITLFQLQSCLNMLTKKLEVHLLLLFLPCKVL 638
Query: 204 VYWLVQLLL-WWFAWYFAAVR----SRQRLLMC-RRRLTLHGG*Y**LVQFQLH*LIIGV 257
V +V+LLL W + +R + +L C L + G + V FQLH* G
Sbjct: 639 VS*VVELLL*LWLLYLITNIRFLPLKKIQLHHC*CLNLIMFGDLFLCSVLFQLH*RTTGE 818
Query: 258 **CQK 262
* CQK
Sbjct: 819 *KCQK 833
>TC77186 phosphate transporter
Length = 2314
Score = 272 bits (695), Expect(2) = 2e-83
Identities = 150/293 (51%), Positives = 191/293 (64%), Gaps = 13/293 (4%)
Frame = +1
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEEHPLPPATNV---AYPLLSREFLWRHGRDLF 322
YTALV +N QAA DM KVL V + EE + T+ +Y L S++F RHG LF
Sbjct: 1120 YTALVAKNAKQAAADMSKVLQVELE--VEEEKVQKMTSDKRNSYGLFSKQFAARHGLALF 1293
Query: 323 ACSANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPG 380
+ WFLLDI FYSQ LFQ +I+ ++ + + E + +AR Q ++A+CST+PG
Sbjct: 1294 GTCSTWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPG 1473
Query: 381 YFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFF 440
Y+FTV FID +GR IQMMGFFFM V FAL PY HW+K EN GF+V+Y L FFF
Sbjct: 1474 YWFTVAFIDHMGRFAIQMMGFFFMTVFMFALAIPY-DHWSKEENRI--GFVVMYSLTFFF 1644
Query: 441 ANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHK----EKEEGY 496
ANFGPN TTF+VPAE+FPAR RSTCHGIS A GK GAI+G+ GFL+A+ + ++GY
Sbjct: 1645 ANFGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDKGY 1824
Query: 497 PKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLE----ENEDEQSHHAEE 545
P GIG+K SLI+LG + VGM T E+ G+SLE ENE E + E+
Sbjct: 1825 PTGIGIKNSLIMLGVINFVGMLCT-LLVPESKGKSLEELSGENEGEGAEATEQ 1980
Score = 55.8 bits (133), Expect(2) = 2e-83
Identities = 79/247 (31%), Positives = 104/247 (41%), Gaps = 10/247 (4%)
Frame = +2
Query: 28 KIRKWQED*RFYQPLTQQKHNTTISRPSS*QEWDFSLTLMTFFPSHSSQRC*EEYTTPTT 87
+I+ E+* Q+ N TIS * EWD SL M F SQ C* EY T
Sbjct: 386 QIQSCLEN*ECLMHSMSQRRNCTISLQL**LEWDSSLMHMIFSAFPLSQSC*GEYIT--L 559
Query: 88 KKSVKSTRSWSQPLSPL----HS*EPPSVSSYSDASET*KAGATSTVSPYY*C*QAL*LL 143
+ + + Q L+ L H E + T G S V + + LL
Sbjct: 560 NQIQQDQEHFHQVLNQL*LVLHLLEH*QANYSLVGLATNLVGKKSMV*HLF-SWWFVQLL 736
Query: 144 ASPSVQQRELVFYYLLVSLDSS*GLELVVIILYLLPLCLSLLIKGQEVLL*LRFFLCKGL 203
+ Q + V + VSL L LVVI L+ L CL++L K EV L L F CK L
Sbjct: 737 QDSLLDQVQRVLWQHFVSLGFGLVLVLVVITLFQLQSCLNMLTKKLEVHLLLLFLPCKVL 916
Query: 204 VYWLVQLLL-WWFAWYFAAVR----SRQRLLMC-RRRLTLHGG*Y**LVQFQLH*LIIGV 257
V +V+LLL W + +R + +L C L + G + F LH* G
Sbjct: 917 VS*VVELLL*LWLLYLITNIRFLPLKKIQLHHC*CLNLIMFGDLFLCSALFLLH*RTTGE 1096
Query: 258 **CQKLP 264
* CQK P
Sbjct: 1097*KCQKQP 1117
>TC77185 similar to GP|12697486|emb|CAC28219. phosphate transporter
{Sesbania rostrata}, partial (71%)
Length = 1467
Score = 270 bits (689), Expect(2) = 3e-80
Identities = 149/290 (51%), Positives = 186/290 (63%), Gaps = 11/290 (3%)
Frame = +2
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEE-HPLPPATNVAYPLLSREFLWRHGRDLFAC 324
YTALV +N QAA DM KVL V + E+ L N + L +R+F RHG L
Sbjct: 293 YTALVAKNGKQAASDMSKVLQVEIEAEEEKVQNLAENQNQKFGLFTRQFAKRHGLHLLGT 472
Query: 325 SANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYF 382
+ WFLLDI FYSQ LFQ +I+ ++ + + E + +AR Q ++A+CST+PGY+
Sbjct: 473 TTTWFLLDIAFYSQNLFQKDIFTAIGWIPPAKEMNAINELYRIARAQTLIALCSTVPGYW 652
Query: 383 FTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFAN 442
FTV FID +GR IQ+MGFFFM V FAL PY HWTK EN GF+V+Y L FFFAN
Sbjct: 653 FTVAFIDYMGRFAIQLMGFFFMTVFMFALAIPY-DHWTKKENRI--GFVVMYSLTFFFAN 823
Query: 443 FGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHK----EKEEGYPK 498
FGPN TTF+VPAE+FPAR RSTCHGIS A GK GAI+G+ GFL+A+ + + GYP
Sbjct: 824 FGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDHGYPT 1003
Query: 499 GIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLE----ENEDEQSHHAE 544
GIG+K SLI+LG V GM T F E G+SLE ENED+ + E
Sbjct: 1004GIGIKNSLIVLGVVNFFGMVFT-FLVPEPNGKSLEEMSGENEDDDAEAIE 1150
Score = 47.8 bits (112), Expect(2) = 3e-80
Identities = 36/92 (39%), Positives = 49/92 (53%), Gaps = 6/92 (6%)
Frame = +3
Query: 175 LYLLPLCLSLLIKGQEVLL*LRFFLCKGLVYWLVQ------LLLWWFAWYFAAVRSRQRL 228
L+LL LCLS+ + +E LL L+ LCK L WLV+ LLL F ++ ++
Sbjct: 3 LFLLQLCLSMRTRRREELLSLQCLLCKDLESWLVEYLH*L*LLLLITNIKFLLMKKTRKR 182
Query: 229 LMCRRRLTLHGG*Y**LVQFQLH*LIIGV**C 260
+C R L G +* LV F+ H I GV* C
Sbjct: 183 RLCYRLLITFGVLF*CLVLFRQHLPITGV*RC 278
>TC85743 phosphate transporter PT4 [Medicago truncatula]
Length = 1850
Score = 219 bits (557), Expect(2) = 1e-62
Identities = 127/283 (44%), Positives = 173/283 (60%), Gaps = 4/283 (1%)
Frame = +2
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEEHPLPP--ATNVAYPLLSREFLWRHGRDLFA 323
YTA+VE N QAA DM +VLD+ + I E+ L A N YPL S EF RHGR L
Sbjct: 743 YTAIVEGNAKQAAADMARVLDIEI--IAEQDKLAEFKAAN-DYPLWSSEFFNRHGRHLIG 913
Query: 324 CSANWFLLDIVFYSQVLFQSEIYKRY--LNEKDDEDVYQEAFHLARIQAILAVCSTIPGY 381
+ WFLLDI FYSQ L Q +IY + + + + E F +R ++A+ T PGY
Sbjct: 914 TMSCWFLLDIAFYSQNLTQKDIYPAMGLIRQDKEMNAIDEVFQTSRAMFVVALFGTFPGY 1093
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFA 441
+FTV+FI+++GR KIQ++GFF M+ F +G Y + K EN + F ++YGL FFFA
Sbjct: 1094 WFTVFFIEKLGRFKIQLVGFFMMSFFMFVIGVKY--EYLKDENKNL--FALLYGLTFFFA 1261
Query: 442 NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPKGIG 501
NFGPN+TTF++PAELFP R RSTCH S A GK GA++G+ G + + +G P+ I
Sbjct: 1262 NFGPNSTTFVLPAELFPTRVRSTCHAFSAASGKAGAMVGAFGIQYYT----LDGTPRKI- 1426
Query: 502 MKASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDEQSHHAE 544
+ +++IL ++G F T+ T ET GRSLEE E +E
Sbjct: 1427 -RRAMMILAFTNLIGFFCTFLVT-ETKGRSLEEISGEDGRESE 1549
Score = 40.0 bits (92), Expect(2) = 1e-62
Identities = 38/107 (35%), Positives = 46/107 (42%), Gaps = 11/107 (10%)
Frame = +3
Query: 167 GLELVVI--ILYLLPLCLSLLIKGQEVLL*LRFFLCKGLVYWLVQLLLWWFAWYFAAVRS 224
GL+LV++ LY P CL+ KG E L * +F CK W L WF WYF
Sbjct: 426 GLDLVLVETTLYQQPSCLNTPTKGHEALS*QQFSQCKA---W-ESSLPDWFQWYFQEFLK 593
Query: 225 RQRLLMCRRRLTLH---------GG*Y**LVQFQLH*LIIGV**CQK 262
L + + G *+ * V FQL *L G * C K
Sbjct: 594 HTTRLPVSMKTQFYQHSPKGIYSGD*FS*SVLFQLR*LTTGE*KCLK 734
>TC84790 similar to GP|13676624|gb|AAK38197.1 phosphate transporter 2
{Lupinus albus}, partial (37%)
Length = 639
Score = 189 bits (481), Expect(2) = 1e-48
Identities = 99/196 (50%), Positives = 127/196 (64%), Gaps = 3/196 (1%)
Frame = +2
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEE-HPLPPATNVAYPLLSREFLWRHGRDLFAC 324
YTALV +N QAA+DM VL V + E+ + ++ L S+EFL RHG L
Sbjct: 23 YTALVAKNAKQAAQDMSSVLQVEIEAEQEKVDKIGVQDKNSFGLFSKEFLRRHGLHLLGT 202
Query: 325 SANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYF 382
++ WFLLDI +YS LFQ +IY +L D + E F +AR ++A+C T+PGY+
Sbjct: 203 TSTWFLLDIAYYSSNLFQKDIYSSIGWLPPAQDMNAIHEVFKVARATTLIALCGTVPGYW 382
Query: 383 FTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFAN 442
FTV FID +GR IQ+MGFFFM V FAL P Y HWTK EN GF+V+Y + FFFAN
Sbjct: 383 FTVAFIDVIGRFAIQLMGFFFMTVFMFALAIP-YDHWTKREN--RIGFLVMYAMTFFFAN 553
Query: 443 FGPNTTTFIVPAELFP 458
FGPN+TTF+VPAE+FP
Sbjct: 554 FGPNSTTFVVPAEIFP 601
Score = 21.9 bits (45), Expect(2) = 1e-48
Identities = 8/8 (100%), Positives = 8/8 (100%)
Frame = +1
Query: 462 RSTCHGIS 469
RSTCHGIS
Sbjct: 613 RSTCHGIS 636
>BF519718 weakly similar to PIR|G84564|G84 probable sugar transporter
[imported] - Arabidopsis thaliana, partial (20%)
Length = 659
Score = 60.5 bits (145), Expect = 2e-09
Identities = 48/162 (29%), Positives = 74/162 (45%), Gaps = 8/162 (4%)
Frame = +3
Query: 383 FTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIY-------- 434
F+ +DR GR + ++G MAVS F LG T N D K I
Sbjct: 39 FSALVLDRFGRRPMLLLGSSGMAVSLFGLGMGC----TLLHNSDEKPMWAIALCVVAVCA 206
Query: 435 GLAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEE 494
++FF GP TT++ +E+FP R R+ ++ +V ++ + + S+ FL S +
Sbjct: 207 AVSFFSIGLGP--TTWVYSSEIFPMRLRAQGTSLAISVNRLISGVVSMSFLSISEE---- 368
Query: 495 GYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
I +L GV ++ Y+F ET G+SLEE E
Sbjct: 369 -----ITFGGMFFVLAGVMVLATLFFYYFLPETKGKSLEEIE 479
>TC78321 similar to GP|21689841|gb|AAM67564.1 unknown protein {Arabidopsis
thaliana}, partial (80%)
Length = 2093
Score = 50.8 bits (120), Expect = 1e-06
Identities = 37/127 (29%), Positives = 57/127 (44%), Gaps = 2/127 (1%)
Frame = +2
Query: 412 GFPYYSHWTKGENHDNKGFMVIYGLAFFFANFGPN--TTTFIVPAELFPARFRSTCHGIS 469
G + + +TKG N G++ I LA + F P T ++V +E++P R+R C GI+
Sbjct: 1436 GVDHRAWYTKG-CPSNFGWIAILALALYIIFFSPGMGTVPWVVNSEIYPLRYRGICGGIA 1612
Query: 470 GAVGKVGAIIGSVGFLWASHKEKEEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMG 529
V ++ S FL IG + +I + IV +F F ET G
Sbjct: 1613 STTVWVSNLVVSQSFL---------SLTVAIGPAWTFMIFAIIAIVAIFFVIIFVPETKG 1765
Query: 530 RSLEENE 536
+EE E
Sbjct: 1766 VPMEEVE 1786
>TC77077 similar to GP|9293909|dbj|BAB01812.1 sugar transporter protein
{Arabidopsis thaliana}, partial (85%)
Length = 1752
Score = 46.6 bits (109), Expect = 2e-05
Identities = 41/158 (25%), Positives = 64/158 (39%), Gaps = 7/158 (4%)
Frame = +1
Query: 386 YFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAF-----FF 440
+ +DR GR + + M VS L +H N GL+ +
Sbjct: 1054 FMLDRYGRRPLLLTSVGGMVVSLLTLAVSLTII-----DHSNTKLNWAIGLSIATVLSYV 1218
Query: 441 ANF--GPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPK 498
A F G T++ +E+FP R R+ V +V + + S+ FL S K
Sbjct: 1219 ATFSIGAGPITWVYSSEIFPLRLRAQGAACGVVVNRVTSGVISMTFLSLS---------K 1371
Query: 499 GIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
GI + + + GG+ +G Y ET G++LEE E
Sbjct: 1372 GITIGGAFFLFGGIATIGWIFFYIMLPETQGKTLEEME 1485
>TC87421 sugar transporter
Length = 1753
Score = 37.4 bits (85), Expect = 0.015
Identities = 48/220 (21%), Positives = 88/220 (39%), Gaps = 10/220 (4%)
Frame = +2
Query: 333 IVFYSQVLFQSEIYKRYLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYFFTVYFIDRVG 392
I+FY+ VLF S + KDD + + + ++A C +I Y +D+ G
Sbjct: 932 IMFYAPVLFNS------IGFKDDASLMSAV--ITGVVNVVATCVSI-------YGVDKWG 1066
Query: 393 RVKIQMMGFFFMAVSFFA----LGFPYYSHWTKGENHDNKGFMVIYGLAFFFANF----G 444
R + + G M + A +G + + G + +V+ + + A F G
Sbjct: 1067 RRALFLEGGAQMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWG 1246
Query: 445 PNTTTFIVPAELFPARFRSTCHGISGAVGKVGA-IIGSVGFLWASHKEKEEGYPKGIGMK 503
P ++VP+E+FP RS ++ +V + ++ V + H MK
Sbjct: 1247 P--LGWLVPSEIFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCH------------MK 1384
Query: 504 ASLIILGGVCIVGMFV-TYFFTKETMGRSLEENEDEQSHH 542
L + ++ M + +F ET G +EE + H
Sbjct: 1385 FGLFLFFAFFVLVMSIYVFFLLPETKGIPIEEMDRVWKSH 1504
>TC86605 similar to PIR|T01853|T01853 probable hexose transport protein
F9D12.17 - Arabidopsis thaliana, partial (91%)
Length = 3111
Score = 37.0 bits (84), Expect = 0.020
Identities = 39/159 (24%), Positives = 70/159 (43%), Gaps = 6/159 (3%)
Frame = +3
Query: 333 IVFYSQVLFQSEIYKRYLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYFFTVYFIDRVG 392
I+FY+ VLF + L K+D +Y A I + V STI ++Y +D++G
Sbjct: 2193 IMFYAPVLFNT------LGFKNDAALYS-----AVITGAINVISTI----VSIYSVDKLG 2327
Query: 393 RVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGF------MVIYGLAFFFANFGPN 446
R K+ + M +S + +KG+ MV ++ F ++GP
Sbjct: 2328 RRKLLLEAGVQMLLSQMVIAIVLGIKVKDHSEELSKGYAALVVVMVCIFVSAFAWSWGP- 2504
Query: 447 TTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFL 485
+++P+E+FP RS ++ V + + + FL
Sbjct: 2505 -LAWLIPSEIFPLETRSAGQSVTVCVNFLFTAVIAQAFL 2618
>CA919673 similar to GP|9858106|gb| putative glucose translocator
{Mesembryanthemum crystallinum}, partial (31%)
Length = 620
Score = 35.8 bits (81), Expect = 0.044
Identities = 38/167 (22%), Positives = 64/167 (37%), Gaps = 2/167 (1%)
Frame = -3
Query: 372 LAVCSTIPGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFM 431
L S + G +DR GR + + F MA S L + W + G +
Sbjct: 576 LVGASNVFGTVIASSLMDRKGRKSLLITSFSGMAASMLLLSVSF--SWKVLAPYS--GSL 409
Query: 432 VIYGLAFFFANF--GPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASH 489
+ G + +F G ++ E+F +R R+ +S + + + FL +
Sbjct: 408 AVLGTVLYVLSFSLGAGPVPALLLPEIFASRIRAKAISLSLGTHWISNFVIGLYFLSVVN 229
Query: 490 KEKEEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
K IG+ + + VC++ + ET GRSLEE E
Sbjct: 228 K---------IGISSVYLGFSTVCLLAVLYIAANVVETKGRSLEEIE 115
>TC78964 weakly similar to PIR|T00450|T00450 probable monosaccharide
transport protein T14N5.7 - Arabidopsis thaliana,
partial (98%)
Length = 1771
Score = 35.8 bits (81), Expect = 0.044
Identities = 66/303 (21%), Positives = 120/303 (38%), Gaps = 21/303 (6%)
Frame = +1
Query: 253 LIIGV**CQKLPGYTALVEQNVL-QAAKDMEKV-----LDVSMSQITEEHPLPPATNVAY 306
+++G C + P +LVEQ L +A K +EKV +D + + L A +
Sbjct: 679 MLLGGIFCAETPN--SLVEQGRLDEARKVLEKVRGTKNVDAEFEDLKDASELAQAVKSPF 852
Query: 307 PLLSREFLWRHGRDLFACSANWFLL-----DIVFYSQVLFQSEIYKRYLNEKDDEDVYQE 361
+L + +R + A F I+FY+ V+FQS +
Sbjct: 853 KVLLKR-KYRPQLIIGALGTPAFQQLTGNNSILFYAPVIFQSSGF--------------- 984
Query: 362 AFHLARIQAILAVCSTIPGYFFTVYFIDRVGRVKI------QMMGFFFMAVSFFALGFPY 415
+ A + + + + +++ +D+ G K +M+ + A+ F +
Sbjct: 985 GSNAALFSSFITNGALLVATVISMFLVDKFGTRKFFLEAGFEMICCMIITAVVLAVEFGH 1164
Query: 416 YSHWTKGENHDNKGFMVIYGLAFFFA---NFGPNTTTFIVPAELFPARFRSTCHGISGAV 472
+KG + F+VI F A ++GP ++VP+ELFP RS I+ V
Sbjct: 1165GKELSKGIS----AFLVIMIFWFVLAYGRSWGP--LGWLVPSELFPLEIRSAAQSIAVCV 1326
Query: 473 GKV-GAIIGSVGFLWASHKEKEEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRS 531
+ A++ + L H + GI ++ GG+ +V +F ET
Sbjct: 1327NMIFTALVAQLFLLSLCHLK------YGI-----FLLFGGLIVVMSVFVFFLLPETKQVP 1473
Query: 532 LEE 534
+EE
Sbjct: 1474IEE 1482
>TC81556 similar to GP|2688830|gb|AAB88879.1| putative sugar transporter
{Prunus armeniaca}, partial (59%)
Length = 1074
Score = 34.7 bits (78), Expect = 0.097
Identities = 39/176 (22%), Positives = 66/176 (37%), Gaps = 2/176 (1%)
Frame = +1
Query: 372 LAVCSTIPGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFM 431
L S + G +D+ GR + + F MA S L + W + G +
Sbjct: 388 LVGASNVIGTAIASSLMDKQGRKSLLITSFSGMAASMLLLSLSFT--WKVLAPYS--GTL 555
Query: 432 VIYGLAFFFANF--GPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASH 489
+ G + +F G ++ E+F +R R+ +S + + + FL +
Sbjct: 556 AVLGTVLYVLSFSLGAGPVPALLLPEIFASRIRAKAVSLSLGTHWISNFVIGLYFLSVVN 735
Query: 490 KEKEEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDEQSHHAEE 545
K G+ + + VC++ + ET GRSLEE E S A E
Sbjct: 736 K---------FGISSVYLGFSAVCVLAVLYIAGNVVETKGRSLEEIERALSSSA*E 876
>TC83621 similar to PIR|T01506|T01506 probable hexose transport protein
T10M13.6 - Arabidopsis thaliana, partial (32%)
Length = 636
Score = 33.9 bits (76), Expect = 0.17
Identities = 42/162 (25%), Positives = 67/162 (40%), Gaps = 9/162 (5%)
Frame = +2
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVS----FFALGFPYYSHWTKGENHD-NKGFM--VIY 434
F ++ +DR+GR + + G M + LG + G+N + +KG+ V+
Sbjct: 20 FISIAIVDRLGRRPLLISGGIQMIICQVIVAIILGIKF------GDNQELSKGYSLSVVV 181
Query: 435 GLAFFFANFGPN--TTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEK 492
+ F FG + + VP+E+FP RS I+ AV + I + FL K
Sbjct: 182 AICLFVLAFGWSWGPLGWTVPSEIFPLEIRSAGQSITVAVNLLFTFIIAQTFLSLLCSFK 361
Query: 493 EEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEE 534
L G + I+ +FV F ET G +EE
Sbjct: 362 ---------FGIFLFFAGWITIMTIFVV-LFLPETKGIPIEE 457
>TC77810 similar to GP|13676624|gb|AAK38197.1 phosphate transporter 2
{Lupinus albus}, partial (11%)
Length = 1035
Score = 32.0 bits (71), Expect = 0.63
Identities = 20/47 (42%), Positives = 22/47 (46%)
Frame = +3
Query: 44 QQKHNTTISRPSS*QEWDFSLTLMTFFPSHSSQRC*EEYTTPTTKKS 90
Q KHN T S+PS WD + MTF EEYTT T S
Sbjct: 372 QPKHNGTFSQPS*LLVWDSAQMHMTFSAYQMLPSYLEEYTTHTQMHS 512
>TC79655 similar to GP|10177966|dbj|BAB11349. uridine kinase-like protein
{Arabidopsis thaliana}, partial (61%)
Length = 1015
Score = 31.6 bits (70), Expect = 0.82
Identities = 20/52 (38%), Positives = 26/52 (49%)
Frame = +2
Query: 67 MTFFPSHSSQRC*EEYTTPTTKKSVKSTRSWSQPLSPLHS*EPPSVSSYSDA 118
+TF +HS QRC K V+ST W Q L P P SVS+ +D+
Sbjct: 41 LTF*LNHSDQRC--------RTKQVRSTTPWKQHLGPT---SPASVSTAADS 163
>TC77434 similar to GP|21554191|gb|AAM63270.1 unknown {Arabidopsis
thaliana}, partial (71%)
Length = 1362
Score = 30.8 bits (68), Expect = 1.4
Identities = 20/52 (38%), Positives = 24/52 (45%)
Frame = +3
Query: 436 LAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWA 487
LA F+ G P L PA FRS + G + V I GSVG LW+
Sbjct: 330 LAKFYGRAGLMNLVNAGPEHLRPAIFRSLLYEACGRI--VNPIYGSVGLLWS 479
>TC83213 weakly similar to GP|16945177|emb|CAC69071. STP5 protein
{Arabidopsis thaliana}, partial (18%)
Length = 563
Score = 30.4 bits (67), Expect = 1.8
Identities = 28/92 (30%), Positives = 43/92 (46%)
Frame = +1
Query: 442 NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPKGIG 501
++GP T+++P+E+FP + R+T I+ AV + + S FL K
Sbjct: 4 SWGP--LTWLIPSEIFPVKIRTTGQSIAVAVQFIIIFVLSQTFLTMLCHMK--------- 150
Query: 502 MKASLIILGGVCIVGMFVTYFFTKETMGRSLE 533
A + V ++ MFV FF ET G LE
Sbjct: 151 FGAFVFYAFWVIVMTMFV-IFFLPETKGIPLE 243
>BQ751129 weakly similar to GP|2258125|emb| AmMst-1 {Amanita muscaria},
partial (14%)
Length = 565
Score = 30.0 bits (66), Expect = 2.4
Identities = 14/40 (35%), Positives = 19/40 (47%)
Frame = -3
Query: 495 GYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEE 534
G +G + + GG+C TYF ET G SLE+
Sbjct: 455 GEKRGNLKSSVFFVWGGLCTCAFVYTYFLVPETKGLSLEQ 336
>BF635519 similar to GP|17064732|gb putative protein {Arabidopsis thaliana},
partial (25%)
Length = 676
Score = 29.6 bits (65), Expect = 3.1
Identities = 16/48 (33%), Positives = 26/48 (53%)
Frame = -3
Query: 436 LAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVG 483
LAF +FG + F+V + ++ HG++ G + AIIG+VG
Sbjct: 527 LAFGLKSFGHSG--FLVNFQEIAPQYSGVIHGMANTAGTLAAIIGTVG 390
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.342 0.150 0.499
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,565,648
Number of Sequences: 36976
Number of extensions: 297503
Number of successful extensions: 2739
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 2649
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2703
length of query: 548
length of database: 9,014,727
effective HSP length: 101
effective length of query: 447
effective length of database: 5,280,151
effective search space: 2360227497
effective search space used: 2360227497
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 61 (28.1 bits)
Medicago: description of AC149576.4


