

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149493.2 + phase: 0
(332 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
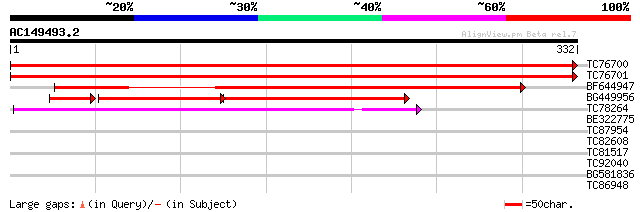
Score E
Sequences producing significant alignments: (bits) Value
TC76700 SP|O48905|MDHC_MEDSA Malate dehydrogenase cytoplasmic (... 655 0.0
TC76701 homologue to GP|10334493|emb|CAC10208. cytosolic malate ... 616 e-177
BF644947 SP|O48905|MDHC Malate dehydrogenase cytoplasmic (EC 1.... 427 e-120
BG449956 similar to SP|Q08062|MDHC Malate dehydrogenase cytopla... 191 2e-91
TC78264 homologue to SP|O48902|MDHP_MEDSA Malate dehydrogenase [... 170 6e-43
BE322775 homologue to SP|O48902|MDHP Malate dehydrogenase [NADP]... 40 0.001
TC87954 similar to GP|21928168|gb|AAM78111.1 AT5g23040/MYJ24_3 {... 29 2.9
TC82608 28 3.8
TC81517 similar to GP|16323198|gb|AAL15333.1 At1g15980/T24D18_8 ... 28 3.8
TC92040 28 4.9
BG581836 similar to GP|8778285|gb| F14D16.26 {Arabidopsis thalia... 28 4.9
TC86948 similar to GP|13194796|gb|AAK15560.1 unknown protein {Ar... 28 6.4
>TC76700 SP|O48905|MDHC_MEDSA Malate dehydrogenase cytoplasmic (EC
1.1.1.37). [Alfalfa] {Medicago sativa}, complete
Length = 1413
Score = 655 bits (1689), Expect = 0.0
Identities = 332/332 (100%), Positives = 332/332 (100%)
Frame = +2
Query: 1 MAKDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVD 60
MAKDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVD
Sbjct: 101 MAKDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVD 280
Query: 61 AAFPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKH 120
AAFPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKH
Sbjct: 281 AAFPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKH 460
Query: 121 AAANCKVLVVANPANTNALILKEFAPSIPERNISCLTRLDHNRALGQISERLNVQVSDVK 180
AAANCKVLVVANPANTNALILKEFAPSIPERNISCLTRLDHNRALGQISERLNVQVSDVK
Sbjct: 461 AAANCKVLVVANPANTNALILKEFAPSIPERNISCLTRLDHNRALGQISERLNVQVSDVK 640
Query: 181 NVIIWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAAIIKARK 240
NVIIWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAAIIKARK
Sbjct: 641 NVIIWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAAIIKARK 820
Query: 241 LSSALSAASAACDHIRDWVLGTPQGTFVSMGVYSDGSYNVPSGLIYSFPVTCANGEWKIV 300
LSSALSAASAACDHIRDWVLGTPQGTFVSMGVYSDGSYNVPSGLIYSFPVTCANGEWKIV
Sbjct: 821 LSSALSAASAACDHIRDWVLGTPQGTFVSMGVYSDGSYNVPSGLIYSFPVTCANGEWKIV 1000
Query: 301 QGLSIDEFSRKKLDLTAEELSEEKNLAYSCLS 332
QGLSIDEFSRKKLDLTAEELSEEKNLAYSCLS
Sbjct: 1001QGLSIDEFSRKKLDLTAEELSEEKNLAYSCLS 1096
>TC76701 homologue to GP|10334493|emb|CAC10208. cytosolic malate
dehydrogenase {Cicer arietinum}, complete
Length = 1440
Score = 616 bits (1589), Expect = e-177
Identities = 311/332 (93%), Positives = 323/332 (96%)
Frame = +3
Query: 1 MAKDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVD 60
MAKDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDI PAAESLNGVKMELVD
Sbjct: 168 MAKDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIPPAAESLNGVKMELVD 347
Query: 61 AAFPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKH 120
AAFPLLKGVVATTDVVE CTGVNIAVMVGGFPRKEGMERKDVM+KNVSIYKSQASALEKH
Sbjct: 348 AAFPLLKGVVATTDVVEGCTGVNIAVMVGGFPRKEGMERKDVMTKNVSIYKSQASALEKH 527
Query: 121 AAANCKVLVVANPANTNALILKEFAPSIPERNISCLTRLDHNRALGQISERLNVQVSDVK 180
AAANCKVLVVANPANTNALILKEFAPSIPE+NISCLTRLDHNRALGQISERLNV+VS+VK
Sbjct: 528 AAANCKVLVVANPANTNALILKEFAPSIPEKNISCLTRLDHNRALGQISERLNVEVSNVK 707
Query: 181 NVIIWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAAIIKARK 240
NVIIWGNHSS+QYPDVNHATV + EKPVRELV+DDAWLNGEFI+TVQQRGAAIIKARK
Sbjct: 708 NVIIWGNHSSSQYPDVNHATVKISSAEKPVRELVADDAWLNGEFIATVQQRGAAIIKARK 887
Query: 241 LSSALSAASAACDHIRDWVLGTPQGTFVSMGVYSDGSYNVPSGLIYSFPVTCANGEWKIV 300
LSSALSAASAACDHIRDWVLGTP+GT+VSMGVYSDGSYNVP+GLIYSFPVT NGEW+IV
Sbjct: 888 LSSALSAASAACDHIRDWVLGTPEGTWVSMGVYSDGSYNVPAGLIYSFPVTTRNGEWQIV 1067
Query: 301 QGLSIDEFSRKKLDLTAEELSEEKNLAYSCLS 332
QGLSIDEFSRKKLDLTAEELSEEK LAYSCLS
Sbjct: 1068QGLSIDEFSRKKLDLTAEELSEEKALAYSCLS 1163
>BF644947 SP|O48905|MDHC Malate dehydrogenase cytoplasmic (EC 1.1.1.37).
[Alfalfa] {Medicago sativa}, partial (68%)
Length = 679
Score = 427 bits (1098), Expect = e-120
Identities = 225/276 (81%), Positives = 225/276 (81%)
Frame = +2
Query: 27 RGVMLGPDQPVILHMLDIAPAAESLNGVKMELVDAAFPLLKGVVATTDVVEACTGVNIAV 86
RGVMLGPDQPVILHMLDIAPAAESLNGVKMELVDAAFPLLKGVV
Sbjct: 2 RGVMLGPDQPVILHMLDIAPAAESLNGVKMELVDAAFPLLKGVV---------------- 133
Query: 87 MVGGFPRKEGMERKDVMSKNVSIYKSQASALEKHAAANCKVLVVANPANTNALILKEFAP 146
AAANCKVLVVANPANTNALILKEFAP
Sbjct: 134 ----------------------------------AAANCKVLVVANPANTNALILKEFAP 211
Query: 147 SIPERNISCLTRLDHNRALGQISERLNVQVSDVKNVIIWGNHSSTQYPDVNHATVNTPAG 206
SIPERNISCLTRLDHNRALGQISERLNVQVSDVKNVIIWGNHSSTQYPDVNHATVNTPAG
Sbjct: 212 SIPERNISCLTRLDHNRALGQISERLNVQVSDVKNVIIWGNHSSTQYPDVNHATVNTPAG 391
Query: 207 EKPVRELVSDDAWLNGEFISTVQQRGAAIIKARKLSSALSAASAACDHIRDWVLGTPQGT 266
EKPVRELVSDDAWLNGEFISTVQQRGAAIIKARKLSSALSAASAACDHIRDWVLGTP GT
Sbjct: 392 EKPVRELVSDDAWLNGEFISTVQQRGAAIIKARKLSSALSAASAACDHIRDWVLGTPXGT 571
Query: 267 FVSMGVYSDGSYNVPSGLIYSFPVTCANGEWKIVQG 302
FVSMGVYSDGSYNVPSGLIYSFPVTCANGEWKIVQG
Sbjct: 572 FVSMGVYSDGSYNVPSGLIYSFPVTCANGEWKIVQG 679
>BG449956 similar to SP|Q08062|MDHC Malate dehydrogenase cytoplasmic (EC
1.1.1.37). [Maize] {Zea mays}, partial (62%)
Length = 637
Score = 191 bits (484), Expect(3) = 2e-91
Identities = 93/109 (85%), Positives = 100/109 (91%)
Frame = +2
Query: 126 KVLVVANPANTNALILKEFAPSIPERNISCLTRLDHNRALGQISERLNVQVSDVKNVIIW 185
KVLV+ANPANTNALILKEFAPSIPE+NI+CLTRLDHNRALGQISE+LNV VSDVKNVIIW
Sbjct: 311 KVLVIANPANTNALILKEFAPSIPEKNITCLTRLDHNRALGQISEKLNVDVSDVKNVIIW 490
Query: 186 GNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAA 234
GNHSSTQYPDVNHATV T G+K VREL +DD WLN EFI+TVQQRG A
Sbjct: 491 GNHSSTQYPDVNHATVATSNGKKHVRELXADDNWLNSEFITTVQQRGGA 637
Score = 135 bits (341), Expect(3) = 2e-91
Identities = 67/74 (90%), Positives = 72/74 (96%)
Frame = +1
Query: 53 GVKMELVDAAFPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKS 112
GVKMEL+DAAFPLL+GVVATTDVVEAC VNIAVMVGGFPRKEGMERKDVMSKNVSIYK+
Sbjct: 91 GVKMELIDAAFPLLRGVVATTDVVEACKDVNIAVMVGGFPRKEGMERKDVMSKNVSIYKT 270
Query: 113 QASALEKHAAANCK 126
QASALE+HAAA+CK
Sbjct: 271 QASALEEHAAADCK 312
Score = 48.5 bits (114), Expect(3) = 2e-91
Identities = 22/27 (81%), Positives = 25/27 (92%)
Frame = +3
Query: 24 MIARGVMLGPDQPVILHMLDIAPAAES 50
MIARG+MLGP+QPVILHMLDI PA E+
Sbjct: 3 MIARGMMLGPNQPVILHMLDIEPALEA 83
>TC78264 homologue to SP|O48902|MDHP_MEDSA Malate dehydrogenase [NADP]
chloroplast precursor (EC 1.1.1.82) (NADP-MDH).
[Alfalfa], partial (91%)
Length = 1783
Score = 170 bits (431), Expect = 6e-43
Identities = 97/240 (40%), Positives = 140/240 (57%), Gaps = 1/240 (0%)
Frame = +1
Query: 3 KDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVDAA 62
K + + V+GAAG I L+ +A G + GP+QPV L +L + ++L GV MEL D+
Sbjct: 460 KKLITIAVSGAAGMISNHLLFKLASGEVFGPNQPVALKLLGSERSLQALEGVAMELEDSL 639
Query: 63 FPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKHAA 122
FPLL+ V+ + D E + A+++G PR G+ER ++ N I+ Q AL A+
Sbjct: 640 FPLLREVIISIDPYEVFQDADWALLIGAKPRGPGVERAALLDINGQIFAEQGKALNAVAS 819
Query: 123 ANCKVLVVANPANTNALILKEFAPSIPERNISCLTRLDHNRALGQISERLNVQVSDVKNV 182
N KV+VV NP NTNALI + AP+IP +N LTRLD NRA Q++ + V V N+
Sbjct: 820 RNVKVIVVGNPCNTNALICLKNAPNIPAKNFHALTRLDENRAKCQLALKAGVFYDKVSNM 999
Query: 183 IIWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFI-STVQQRGAAIIKARKL 241
IWGNHS+TQ PD +A ++ PV+E++ D WL EF + ++R A K K+
Sbjct: 1000TIWGNHSTTQVPDFLNARID----GLPVKEVIKDHKWLEEEFTEKSSKERWCAYSKMGKI 1167
Score = 34.3 bits (77), Expect = 0.068
Identities = 18/48 (37%), Positives = 27/48 (55%)
Frame = +2
Query: 225 ISTVQQRGAAIIKARKLSSALSAASAACDHIRDWVLGTPQGTFVSMGV 272
+ VQ+RG +I+ SSA S + + D IR + TP+G + S GV
Sbjct: 1115 LKKVQKRGGVLIQKWGRSSAASTSVSIVDAIRSLITPTPEGDWFSTGV 1258
>BE322775 homologue to SP|O48902|MDHP Malate dehydrogenase [NADP]
chloroplast precursor (EC 1.1.1.82) (NADP-MDH).
[Alfalfa], partial (24%)
Length = 528
Score = 40.0 bits (92), Expect = 0.001
Identities = 24/66 (36%), Positives = 38/66 (57%), Gaps = 1/66 (1%)
Frame = +3
Query: 3 KDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPA-AESLNGVKMELVDA 61
K + + V+GAAG I L+ +A G + GP+QPV L +L + A+ L ++ D+
Sbjct: 330 KKLITIAVSGAAGMISNHLLFKLASGEVFGPNQPVALKLLGSERSFAKLLKVLRWSWEDS 509
Query: 62 AFPLLK 67
FPLL+
Sbjct: 510 LFPLLR 527
>TC87954 similar to GP|21928168|gb|AAM78111.1 AT5g23040/MYJ24_3 {Arabidopsis
thaliana}, partial (82%)
Length = 1200
Score = 28.9 bits (63), Expect = 2.9
Identities = 21/70 (30%), Positives = 30/70 (42%), Gaps = 2/70 (2%)
Frame = +3
Query: 237 KARKLSSALSAASAACDHIRDWVLGTPQGTFVSMGVYSDGSYNVPSGLIYSFPV--TCAN 294
K ++ S A+SA C IRDW + Q T S+ S + FP+ T
Sbjct: 75 KGKERRSFSQASSATCSSIRDWEITLVQFTLSITTTQHGFSFTFISQFPHRFPLQKTSPP 254
Query: 295 GEWKIVQGLS 304
G++K LS
Sbjct: 255 GKYKEFYNLS 284
>TC82608
Length = 798
Score = 28.5 bits (62), Expect = 3.8
Identities = 11/37 (29%), Positives = 17/37 (45%)
Frame = +2
Query: 175 QVSDVKNVIIWGNHSSTQYPDVNHATVNTPAGEKPVR 211
Q D +++WG H Q D+N + G +P R
Sbjct: 353 QPKDPNQLLVWGGHQDKQSKDLNQLFASDVHGSQPKR 463
>TC81517 similar to GP|16323198|gb|AAL15333.1 At1g15980/T24D18_8
{Arabidopsis thaliana}, partial (26%)
Length = 688
Score = 28.5 bits (62), Expect = 3.8
Identities = 15/36 (41%), Positives = 20/36 (54%)
Frame = +1
Query: 146 PSIPERNISCLTRLDHNRALGQISERLNVQVSDVKN 181
PS N+S LT LDH + RLN++V+ KN
Sbjct: 130 PSTYPTNVSLLTSLDHPSHCYNSTRRLNLRVNAKKN 237
>TC92040
Length = 1133
Score = 28.1 bits (61), Expect = 4.9
Identities = 21/106 (19%), Positives = 41/106 (37%)
Frame = +1
Query: 149 PERNISCLTRLDHNRALGQISERLNVQVSDVKNVIIWGNHSSTQYPDVNHATVNTPAGEK 208
P+R S +DH R S+ D K + +G+ + D + N +
Sbjct: 196 PKRKTSDREDVDHKRRTSHSSKAAFTPTCDAKQLAFYGDRGPKVFKDDIRSIKNKNLKDT 375
Query: 209 PVRELVSDDAWLNGEFISTVQQRGAAIIKARKLSSALSAASAACDH 254
V++ +S+D++ G +T++ + + S AL H
Sbjct: 376 VVQDHISNDSFAVG---TTIEVSNNNLTTTEESSEALYTKKTRRSH 504
>BG581836 similar to GP|8778285|gb| F14D16.26 {Arabidopsis thaliana}, partial
(18%)
Length = 749
Score = 28.1 bits (61), Expect = 4.9
Identities = 20/103 (19%), Positives = 43/103 (41%), Gaps = 8/103 (7%)
Frame = +3
Query: 236 IKARKLSSALSAASAACDHIRDWVLGTP--QGTFVSMGVYS------DGSYNVPSGLIYS 287
++ L + S D ++ TP G F+ +++ DG N+ + +S
Sbjct: 429 VEVNALRKSYSTRLVVMDECKENQSATPAQNGGFLKSNIFTLTIPQIDGGSNLSIKMRWS 608
Query: 288 FPVTCANGEWKIVQGLSIDEFSRKKLDLTAEELSEEKNLAYSC 330
V C+NGE+ + + EF ++ + +S+ + + C
Sbjct: 609 QKVVCSNGEYSLNVPFTFPEF----INPAGKRMSKREKIRLKC 725
>TC86948 similar to GP|13194796|gb|AAK15560.1 unknown protein {Arabidopsis
thaliana}, partial (78%)
Length = 802
Score = 27.7 bits (60), Expect = 6.4
Identities = 13/40 (32%), Positives = 23/40 (57%)
Frame = +2
Query: 277 SYNVPSGLIYSFPVTCANGEWKIVQGLSIDEFSRKKLDLT 316
++ PSGL SFPV+ E + G+++ E + K++ T
Sbjct: 443 TFKTPSGLFRSFPVSAFEIEEEKSSGVAVKEVAANKVNET 562
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.132 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,827,633
Number of Sequences: 36976
Number of extensions: 111482
Number of successful extensions: 504
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 499
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 503
length of query: 332
length of database: 9,014,727
effective HSP length: 97
effective length of query: 235
effective length of database: 5,428,055
effective search space: 1275592925
effective search space used: 1275592925
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC149493.2


