

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
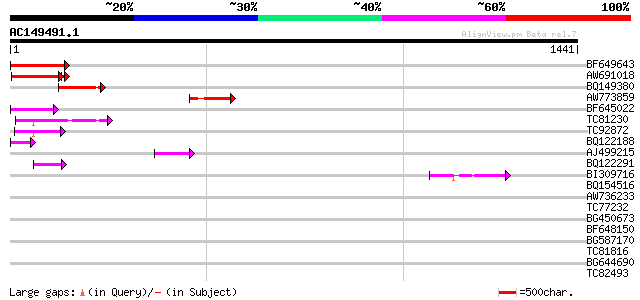
Query= AC149491.1 + phase: 0 /pseudo
(1441 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BF649643 similar to PIR|H86486|H86 protein Ty1/copia-element pol... 128 2e-29
AW691018 weakly similar to GP|2462935|em open reading frame 1 {B... 119 7e-27
BQ149380 100 4e-21
AW773859 97 6e-20
BF645022 weakly similar to GP|8777581|dbj retroelement pol polyp... 80 6e-15
TC81230 73 9e-13
TC92872 67 5e-11
BQ122188 weakly similar to GP|9758772|dbj| contains similarity t... 54 6e-07
AJ499215 weakly similar to GP|6642775|gb| gag-pol polyprotein {V... 50 6e-06
BQ122291 49 1e-05
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 44 5e-04
BQ154516 41 0.003
AW736233 GP|21105451|gb small nuclear ribonucleoprotein D1 {Dani... 40 0.007
TC77232 similar to SP|O04130|SERA_ARATH D-3-phosphoglycerate deh... 40 0.009
BG450673 homologue to GP|12055499|emb NADH dehydrogenase subunit... 37 0.042
BF648150 similar to GP|14586969|gb| pol polyprotein {Citrus x pa... 37 0.042
BG587170 similar to PIR|F86470|F8 probable retroelement polyprot... 36 0.095
TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana taba... 35 0.21
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 35 0.28
TC82493 weakly similar to GP|4056491|gb|AAC98057.1| unknown prot... 33 0.80
>BF649643 similar to PIR|H86486|H86 protein Ty1/copia-element polyprotein
[imported] - Arabidopsis thaliana, partial (2%)
Length = 608
Score = 128 bits (321), Expect = 2e-29
Identities = 60/150 (40%), Positives = 91/150 (60%), Gaps = 2/150 (1%)
Frame = +1
Query: 3 YYVHPSEGPNSVTVTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDD--LNRT 60
YY+HPS+ P + V L G NY +W+RSM AL KNK FI+GS+ P D
Sbjct: 124 YYIHPSDHPGHLLVPTKLNGTNYSSWSRSMVHALTAKNKVGFINGSIKTPSEVDQPAEYA 303
Query: 61 AWERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNL 120
W RCN++ILSW+ +SV P +A+ ++ + A VW + +++FS+ + + ++ S+ +L
Sbjct: 304 LWNRCNSMILSWLTHSVEPDLAKGVIHAKTACQVWEDFKDQFSQKNIPAIYQIQKSLASL 483
Query: 121 KQGDKSVLDYFTSIKSLWEELNSHRPMPMC 150
QG S YFT IK LW+EL ++R +P C
Sbjct: 484 SQGTVSASTYFTKIKGLWDELETYRTLPTC 573
>AW691018 weakly similar to GP|2462935|em open reading frame 1 {Brassica
oleracea}, partial (12%)
Length = 639
Score = 119 bits (298), Expect(2) = 7e-27
Identities = 55/133 (41%), Positives = 82/133 (61%)
Frame = +2
Query: 5 VHPSEGPNSVTVTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNRTAWER 64
+H S+ P V L G NY W+RSM +L KNK IDG+V P DD W+R
Sbjct: 164 LHHSDTPGISLVNQPLNGGNYGEWSRSMLLSLSAKNKLGLIDGTVKAPSADDPKLPLWKR 343
Query: 65 CNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNLKQGD 124
CN+L+L+WI++S+ P IA++++F + A VW +L +RFS+ D R+ +R I+ +QG
Sbjct: 344 CNDLVLTWILHSIEPDIARSVIFSDTAAAVWSDLHDRFSQGDESRIYQIRQEISECRQGS 523
Query: 125 KSVLDYFTSIKSL 137
+ DY+T +KSL
Sbjct: 524 LLISDYYTKLKSL 562
Score = 21.2 bits (43), Expect(2) = 7e-27
Identities = 7/20 (35%), Positives = 12/20 (60%)
Frame = +3
Query: 133 SIKSLWEELNSHRPMPMCTC 152
S + W+EL S++ C+C
Sbjct: 549 SSRVFWDELGSYQEPITCSC 608
>BQ149380
Length = 419
Score = 100 bits (249), Expect = 4e-21
Identities = 51/124 (41%), Positives = 78/124 (62%), Gaps = 5/124 (4%)
Frame = +3
Query: 125 KSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGLNDSF 184
+S L+ ++ LW+EL+ +RP P CTCP C C +MR + ED++I+FL GLND +
Sbjct: 45 ESYLNSSPDLRGLWDELDQYRPKPQCTCPIQCTCFAMRNNKALTAEDRIIKFLMGLNDDY 224
Query: 185 SVVKTQVLLIDPLPSINKVYSMVIQEESN-----IIPPTSLASNEDSSILVNASDARKPF 239
V ++VLL+ PLP IN V+SMV+Q+E + PT + ++ + LVNA D ++ F
Sbjct: 225 QGVTSKVLLMYPLPQINMVFSMVMQQERKMQ*NVVSSPTPI--DDTTCGLVNAMDGQRQF 398
Query: 240 LRGK 243
RG+
Sbjct: 399 GRGR 410
>AW773859
Length = 538
Score = 96.7 bits (239), Expect = 6e-20
Identities = 46/117 (39%), Positives = 72/117 (61%)
Frame = -3
Query: 456 LNTYVIQDVKTQKMIGLGNLCDGLYRLHPFAPASPQAHFISSAVSPTNKMSCNSVSSNNN 515
L +IQ+ ++ KMIG G L +GLY F+++ +P+ ++ + V
Sbjct: 395 LQVILIQEKRSLKMIGSGELFEGLY-------------FLTTQDTPSATIASSQVQPQPQ 255
Query: 516 VSSIPSNAIWHFRLGHLSNQRLSMMHSLYSSITIDNKAVCDICHFAKQRKLPYNLST 572
+ +P A+WHFRLGHLSN++L +HS + ITID +VCDICH+++ +KLP+ LST
Sbjct: 254 PTFLPQEALWHFRLGHLSNRKLLSLHSNFPFITIDQNSVCDICHYSRHKKLPFQLST 84
>BF645022 weakly similar to GP|8777581|dbj retroelement pol polyprotein-like
{Arabidopsis thaliana}, partial (8%)
Length = 685
Score = 80.1 bits (196), Expect = 6e-15
Identities = 31/122 (25%), Positives = 71/122 (57%)
Frame = -2
Query: 2 IYYVHPSEGPNSVTVTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNRTA 61
++++ ++ P +V L G NY W+R+++ + K K+ F++G +P P +
Sbjct: 378 VFFLGSNDNPGNVITPIQLRGLNYDEWSRAIRTSFQAKRKYGFVEGKIPKPTTPE-KLED 202
Query: 62 WERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNLK 121
W+ +++++W++N++ P + T+ +++ A +W L++RF V+ R+ L++S+ K
Sbjct: 201 WKAVQSMLIAWLLNTIEPSLRSTLSYYDDAESLWTHLKQRFCVVNGARICQLKASLGECK 22
Query: 122 QG 123
QG
Sbjct: 21 QG 16
>TC81230
Length = 958
Score = 72.8 bits (177), Expect = 9e-13
Identities = 64/256 (25%), Positives = 108/256 (42%), Gaps = 11/256 (4%)
Frame = +1
Query: 16 VTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPM--DDL------NRTAWERCNN 67
++ +L G NY W SM L + + ++ G P DD W+ N+
Sbjct: 226 ISIILNGSNYNHWAESMCGFLKGRRLWRYVTGDKKCPTKGKDDTADAFADKLEEWDSKNH 405
Query: 68 LILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNLK-QGDKS 126
I++W N+ P I E A +VW L++R++ D L ++NLK Q +
Sbjct: 406 QIITWFRNTSIPSIHMQFGRFENAKEVWDHLKQRYTISDLSHQYQLLKDLSNLKQQSGQP 585
Query: 127 VLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGLNDSFSV 186
V ++ ++ +W +L S P M+ R ++IQFL L D +
Sbjct: 586 VYEFLAQMEVIWNQLTSCEPSLK-------DATDMKTYETHRNRVRLIQFLMALTDEYEP 744
Query: 187 VKTQVLLIDPLPSINKVYSMVIQEES--NIIPPTSLASNEDSSILVNASDARKPFLRGKS 244
V+ L +PLP++ + EE+ ++PP + D + V ++A KP +
Sbjct: 745 VRASSLHQNPLPTLENALPCLKSEETRLQLVPPKA-----DLAFAV-TNNATKPCRHCQK 906
Query: 245 SGTSQSKNNSRYCTFC 260
SG S S + C C
Sbjct: 907 SGHSFSDCPTIECRNC 954
>TC92872
Length = 923
Score = 67.0 bits (162), Expect = 5e-11
Identities = 38/138 (27%), Positives = 69/138 (49%), Gaps = 9/138 (6%)
Frame = +1
Query: 13 SVTVTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNR---------TAWE 63
+ ++T L+ N+ AW R + L IDG+ PI +N T W
Sbjct: 88 NASITFKLSRKNFRAWKRQVTTLLAGIEVMGHIDGTTPIQTETIINNGVSAPNPDYTKWF 267
Query: 64 RCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNLKQG 123
+ LI++ +++S++ + + +E A +W+ ++ +F+ R V S+ + I +G
Sbjct: 268 TLDQLIINLLLSSMTEADSISFASYETARTLWVAIESQFNNTSRSHVMSVTNQIQRCTKG 447
Query: 124 DKSVLDYFTSIKSLWEEL 141
DKS+ DY S+KSL +EL
Sbjct: 448 DKSITDYLFSVKSLADEL 501
>BQ122188 weakly similar to GP|9758772|dbj| contains similarity to
retroelement pol polyprotein~gene_id:MNJ7.3 {Arabidopsis
thaliana}, partial (7%)
Length = 708
Score = 53.5 bits (127), Expect = 6e-07
Identities = 27/63 (42%), Positives = 36/63 (56%), Gaps = 1/63 (1%)
Frame = -3
Query: 3 YYVHPSEGPNSVTVTPLLTG-PNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNRTA 61
+++HPSE P VTP L G N+ +W RSM+ AL +KNK F+DG+ P D
Sbjct: 190 FFLHPSENPAVELVTPPLEGNKNFQSWIRSMRLALTSKNKIAFVDGTFLTPAKSDALYNQ 11
Query: 62 WER 64
W R
Sbjct: 10 WIR 2
>AJ499215 weakly similar to GP|6642775|gb| gag-pol polyprotein {Vitis
vinifera}, partial (18%)
Length = 567
Score = 50.1 bits (118), Expect = 6e-06
Identities = 33/105 (31%), Positives = 53/105 (50%), Gaps = 3/105 (2%)
Frame = +3
Query: 369 PSDHWLIDSGANEHICSSLHLFHSYYRIKPICVNLPNGSSVIVQYAGTVVFSPHF---HI 425
PS +WLIDSG H+ LF + V + NG+ + V+ GTV+ H I
Sbjct: 96 PSKYWLIDSGCTHHMTHDRDLFKELNKSTISKVRMLNGAHIEVEGIGTVLVKSHSGYKQI 275
Query: 426 THVLYSPSFKVYLISVSKICQSLPYHVHFLLNTYVIQDVKTQKMI 470
++VLY+P L+SV ++ + Y V F VI+D ++++
Sbjct: 276 SNVLYAPKLNQSLLSVPQLL-TKGYKVLFEHEKCVIKDQNNKEVL 407
>BQ122291
Length = 487
Score = 48.9 bits (115), Expect = 1e-05
Identities = 22/84 (26%), Positives = 45/84 (53%)
Frame = -2
Query: 60 TAWERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINN 119
T WE ++L+ +WI++ +SP + V ++ VW E+ + R LRS + +
Sbjct: 447 TEWEDKDSLLCTWILSRISPSLLSRFVLLRHSWQVWDEIHSYCFTQMKTRSRQLRSELRS 268
Query: 120 LKQGDKSVLDYFTSIKSLWEELNS 143
+ +G ++V ++ I+++ E L S
Sbjct: 267 ITKGSRTVSEFIARIRAISESLAS 196
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 43.9 bits (102), Expect = 5e-04
Identities = 66/218 (30%), Positives = 92/218 (41%), Gaps = 13/218 (5%)
Frame = +1
Query: 1067 QILIWSQTSQ*KVV*KADHSPYFQ*L*TSCI*CFFVYQINF*DFHYYFSLC**YNFSWQ- 1125
+I IW Q +* +V +F L T * F Y+I +Y S+C**+ FSW+
Sbjct: 34 KIYIWPQAGK*TMVF*VI*ISHFLWLSTILF*FFPFYKI*RFFIYYIVSIC**HCFSWK* 213
Query: 1126 ---------FSH*ISYHQECFTSGIQNQGSWHFEIFSWP*SSSFTQWDFSMPKKILFGSV 1176
FS+* NQ W IFS * F+ KKI +
Sbjct: 214 YI*NPTC*MFSY*----------SF*NQRPWFLTIFSRS*GC*KQTRHFT*SKKIYS*T- 360
Query: 1177 E*FRPSWF*AC---FYTL*SLYQTS**HFTSFP*HFSL*KTHWQIDLLEYNQT*HYFHYP 1233
FR W C Y+L* ++T F * S+*KTH Q L Y +*+
Sbjct: 361 --FRG*W*SCCKIHSYSL*YFFKTPQF*FAPLQ**NSI*KTHRQTYLFNYYSS*YILCCS 534
Query: 1234 AIESVS*QTNSNSLSCYS*SSKIPKRFTRQRNFLPTRF 1271
I+ + +T+++SLS *S I + QR L + F
Sbjct: 535 TIKPICFETSASSLSGCY*SFTISENCPCQRIILLSNF 648
>BQ154516
Length = 506
Score = 41.2 bits (95), Expect = 0.003
Identities = 39/136 (28%), Positives = 62/136 (44%), Gaps = 2/136 (1%)
Frame = +1
Query: 425 ITHVLYSPSFKVYLISVSKICQSLPYHVHFLLNTYVIQDVKTQKMIGLGNLCDGLYRLHP 484
+ H++ SPS + L+ P V +++D K + G+ DGLY+ P
Sbjct: 139 LEHLIISPSILITLLIAP------PTMVRNRC*WIMVKDSKQILLEGVVG-SDGLYQFKP 297
Query: 485 --FAPASPQAHFISSAVSPTNKMSCNSVSSNNNVSSIPSNAIWHFRLGHLSNQRLSMMHS 542
F P+ +S ++S N + +SV N++ SS S WH RLGH + + + +
Sbjct: 298 LQFLPS------MSKSLSYCNNATISSVVCNSSSSSNDSFNKWHCRLGHANPNVVKSILN 459
Query: 543 LYSSITIDNKAVCDIC 558
L I NK V D C
Sbjct: 460 L-CKIPFQNKHVLDFC 504
>AW736233 GP|21105451|gb small nuclear ribonucleoprotein D1 {Danio rerio},
partial (14%)
Length = 510
Score = 40.0 bits (92), Expect = 0.007
Identities = 21/60 (35%), Positives = 28/60 (46%)
Frame = +1
Query: 241 RGKSSGTSQSKNNSRYCTFCRRNNHTVEYCYLKHDFPNANKPTASSNAVTSEHAVDSHTS 300
RG+ G +S +N C C RNNH C ++D A+ A A SE+A S S
Sbjct: 250 RGRGRGRGRSNSNRLQCQICARNNHDAARCCFRYD--QASSSQAHHRAPPSEYAASSSYS 423
>TC77232 similar to SP|O04130|SERA_ARATH D-3-phosphoglycerate dehydrogenase
chloroplast precursor (EC 1.1.1.95) (3-PGDH). [Mouse-ear
cress], partial (87%)
Length = 2474
Score = 39.7 bits (91), Expect = 0.009
Identities = 31/111 (27%), Positives = 48/111 (42%), Gaps = 6/111 (5%)
Frame = -2
Query: 256 YCTFCR-RNNHTVEY---CYLKHDFPN--ANKPTASSNAVTSEHAVDSHTSSEGTSSSSQ 309
+CT C R+N + + C+ H + N A++P +S S V SH+S + S
Sbjct: 841 FCTPCNFRSNFSKPHYC*CFPNHRYSNILASQPFSSFKGSISLRNVTSHSSKKSNPMFSS 662
Query: 310 TGLTQEQYVHLVSLLQQSSLVPSATPPNPASTNHVATSFPSSIDFTSGINT 360
+ + +H + SSL PN +STN+ SFP F T
Sbjct: 661 SNSISSRSIHNKTTKFSSSL*IHIINPNTSSTNNFQLSFPCFKHFPCNFRT 509
>BG450673 homologue to GP|12055499|emb NADH dehydrogenase subunit 2 {Sphagnum
fallax}, partial (2%)
Length = 678
Score = 37.4 bits (85), Expect = 0.042
Identities = 17/57 (29%), Positives = 26/57 (44%)
Frame = +3
Query: 39 KNKFVFIDGSVPIPPMDDLNRTAWERCNNLILSWIINSVSPQIAQTIVFHEYAIDVW 95
K+K +I+G +P PP D W N ++ WIINS+ + + VW
Sbjct: 231 KDKLGYINGDLPPPPQTDPTFRRWGTDNAIVKGWIINSMDSSLISNFIRFSTTKLVW 401
>BF648150 similar to GP|14586969|gb| pol polyprotein {Citrus x paradisi},
partial (3%)
Length = 658
Score = 37.4 bits (85), Expect = 0.042
Identities = 23/88 (26%), Positives = 39/88 (44%), Gaps = 1/88 (1%)
Frame = +2
Query: 58 NRTAWERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSI 117
+R W+ + + L I+N +S + A D+W +L+ R+ + D L S
Sbjct: 167 DRQKWDNDDYICLGHILNGMSDSLFDIYQSSPSAKDLWDKLETRYMREDATSKKFLVSHF 346
Query: 118 NNLKQGD-KSVLDYFTSIKSLWEELNSH 144
NN K D KSV++ I+ + H
Sbjct: 347 NNYKMVDNKSVMEQLYEIERILNNYKQH 430
>BG587170 similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 718
Score = 36.2 bits (82), Expect = 0.095
Identities = 21/72 (29%), Positives = 35/72 (48%)
Frame = -3
Query: 494 FISSAVSPTNKMSCNSVSSNNNVSSIPSNAIWHFRLGHLSNQRLSMMHSLYSSITIDNKA 553
++ + P + C+ SS SS+ +A+WH RLGH + L++M + +NK
Sbjct: 662 YMLEKLDPVSNYKCSFTSS----SSLNKDALWHARLGHPHGRALNLM---LPGVVFENKN 504
Query: 554 VCDICHFAKQRK 565
C+ C K K
Sbjct: 503 -CEACILGKHCK 471
>TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana tabacum},
partial (13%)
Length = 663
Score = 35.0 bits (79), Expect = 0.21
Identities = 49/170 (28%), Positives = 78/170 (45%), Gaps = 2/170 (1%)
Frame = -2
Query: 590 FLLLVFMDTSIFSPLLMILVDFYGSFC*KINQK--SQIMSKILSY*SILIIKLLPST*DL 647
FLLL+F+ +F LL++ F G F I + + +L LI+ LL T L
Sbjct: 623 FLLLLFL---VFFFLLLLFTFFRGFFYINIPYRYFTLFFLILLLLFLFLILILLFLTSIL 453
Query: 648 TMVLSF*SLNFMLLRESYTKNHVLKLPNKMVELKESINISLMLLELYFSNLNFQIFFGHM 707
T ++ F + F+LL + L + + + SL+LL L+ +
Sbjct: 452 TFIIVFILVLFLLL---------VFLRFLLFTILVFVFFSLLLLFLFL-----------L 333
Query: 708 LLFMLFS**IKCLPLFLNKNLLINFFMILYLT*IPSKFLVVYVLLLLFLL 757
LL +LF I LN L+ FF++L++ I + + +VLLLL LL
Sbjct: 332 LLILLF---ITIFFFTLNILFLLFFFLLLFVLLILTLIYISFVLLLLILL 192
Score = 29.6 bits (65), Expect = 8.9
Identities = 26/79 (32%), Positives = 40/79 (49%), Gaps = 8/79 (10%)
Frame = -2
Query: 687 SLMLLELYFSNLNFQIFFGHMLLFMLF------S**IKCLPLFLNKNLLINFFMILYLT* 740
SL+ + F L F +FF +LLF F + + LF LL+ F+IL L
Sbjct: 647 SLVFILSRFLLLLFLVFFFLLLLFTFFRGFFYINIPYRYFTLFFLILLLLFLFLILILLF 468
Query: 741 IPS--KFLVVYVLLLLFLL 757
+ S F++V++L+L LL
Sbjct: 467 LTSILTFIIVFILVLFLLL 411
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 34.7 bits (78), Expect = 0.28
Identities = 33/92 (35%), Positives = 45/92 (48%), Gaps = 1/92 (1%)
Frame = -1
Query: 993 IQSN*RVGLF*YFFTSCQNHYSQTCNCSSFN*SLVSSSIRCK*CIFAW*SSRKCLQEDST 1052
IQS R L * FFT CQN N + + C+ CI+ W S R + + ++
Sbjct: 389 IQSKRRNRL**GFFTCCQNGSY*NFNSFCCIHGVQAVPNGCEECIY*WRSQRGGVCQATS 210
Query: 1053 RIIHIQ-TKSSLQIIQILIWSQTSQ*KVV*KA 1083
I + TKS +QI IWS+ S +V*KA
Sbjct: 209 WI*RCRGTKSCVQIE*DTIWSEASSKSMV*KA 114
>TC82493 weakly similar to GP|4056491|gb|AAC98057.1| unknown protein
{Arabidopsis thaliana}, partial (35%)
Length = 1158
Score = 33.1 bits (74), Expect = 0.80
Identities = 20/62 (32%), Positives = 29/62 (46%)
Frame = +1
Query: 249 QSKNNSRYCTFCRRNNHTVEYCYLKHDFPNANKPTASSNAVTSEHAVDSHTSSEGTSSSS 308
+SKN + + FC N V+ ++ DF N K S + V H + SS TSSS
Sbjct: 655 KSKNKNSFEVFCEENKKQVKRDMVEDDFVNHRK---SFSGVVQRHCGSNKVSSLSTSSSG 825
Query: 309 QT 310
+
Sbjct: 826 SS 831
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.356 0.156 0.555
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 54,753,657
Number of Sequences: 36976
Number of extensions: 975738
Number of successful extensions: 13513
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 4426
Number of HSP's successfully gapped in prelim test: 968
Number of HSP's that attempted gapping in prelim test: 8610
Number of HSP's gapped (non-prelim): 6461
length of query: 1441
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1333
effective length of database: 5,021,319
effective search space: 6693418227
effective search space used: 6693418227
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC149491.1


