

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149471.21 - phase: 0
(281 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
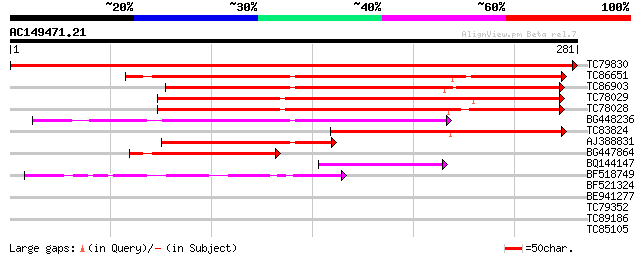
Score E
Sequences producing significant alignments: (bits) Value
TC79830 similar to PIR|G84861|G84861 hypothetical protein At2g43... 579 e-166
TC86651 similar to PIR|F84555|F84555 similar to prolyl 4-hydroxy... 217 5e-57
TC86903 similar to GP|21617881|gb|AAM66931.1 prolyl 4-hydroxylas... 191 2e-49
TC78029 similar to GP|21593296|gb|AAM65245.1 prolyl 4-hydroxylas... 189 1e-48
TC78028 similar to GP|22136524|gb|AAM91340.1 unknown protein {Ar... 181 3e-46
BG448236 similar to GP|21537370|gb putative prolyl 4-hydroxylase... 158 3e-39
TC83824 similar to GP|17381226|gb|AAL36425.1 unknown protein {Ar... 98 3e-21
AJ388831 weakly similar to GP|10177121|dbj prolyl 4-hydroxylase ... 79 1e-15
BG447864 weakly similar to GP|10177121|db prolyl 4-hydroxylase ... 67 1e-11
BQ144147 46 2e-05
BF518749 44 5e-05
BF521324 similar to GP|18086437|gb| AT3g28480/MFJ20_16 {Arabidop... 39 0.003
BE941277 29 1.8
TC79352 weakly similar to PIR|T01829|T01829 hypothetical protein... 28 3.9
TC89186 similar to PIR|G96563|G96563 probable coatomer complex s... 28 5.1
TC85105 similar to GP|20160725|dbj|BAB89667. putative transaldol... 27 6.7
>TC79830 similar to PIR|G84861|G84861 hypothetical protein At2g43080
[imported] - Arabidopsis thaliana, partial (87%)
Length = 1130
Score = 579 bits (1492), Expect = e-166
Identities = 281/281 (100%), Positives = 281/281 (100%)
Frame = +1
Query: 1 MSPPAMKIVFGLLTFVTIGMIIGALSQLAFIRRLELEEPFTTTTTTRSLLPRGYTYWNNN 60
MSPPAMKIVFGLLTFVTIGMIIGALSQLAFIRRLELEEPFTTTTTTRSLLPRGYTYWNNN
Sbjct: 94 MSPPAMKIVFGLLTFVTIGMIIGALSQLAFIRRLELEEPFTTTTTTRSLLPRGYTYWNNN 273
Query: 61 NDKEAQILRLGYVKPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKG 120
NDKEAQILRLGYVKPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKG
Sbjct: 274 NDKEAQILRLGYVKPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKG 453
Query: 121 IKSDVRTSSGMFLSHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHD 180
IKSDVRTSSGMFLSHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHD
Sbjct: 454 IKSDVRTSSGMFLSHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHD 633
Query: 181 YFSDTFNLKRGGQRIATMLMYLGDNVEGGETHFPSAGSDECSCGGKLTKGLCVKPVKGNA 240
YFSDTFNLKRGGQRIATMLMYLGDNVEGGETHFPSAGSDECSCGGKLTKGLCVKPVKGNA
Sbjct: 634 YFSDTFNLKRGGQRIATMLMYLGDNVEGGETHFPSAGSDECSCGGKLTKGLCVKPVKGNA 813
Query: 241 VLFWSMGLDGQSDPDSVHGGCPVLAGEKWSATKWMRQSVHV 281
VLFWSMGLDGQSDPDSVHGGCPVLAGEKWSATKWMRQSVHV
Sbjct: 814 VLFWSMGLDGQSDPDSVHGGCPVLAGEKWSATKWMRQSVHV 936
>TC86651 similar to PIR|F84555|F84555 similar to prolyl 4-hydroxylase alpha
subunit [imported] - Arabidopsis thaliana, partial (85%)
Length = 1329
Score = 217 bits (552), Expect = 5e-57
Identities = 117/227 (51%), Positives = 150/227 (65%), Gaps = 8/227 (3%)
Frame = +2
Query: 58 NNNNDKEAQILRLGYVKPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANT 117
+ N+D+E + G EV+SW PR + HNFL+ EEC+YL +A P + STVVD+ T
Sbjct: 311 DRNDDEEGK----GEQWVEVVSWEPRAFVYHNFLTKEECEYLIDIAKPSMHKSTVVDSET 478
Query: 118 GKGIKSDVRTSSGMFLSHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRP 177
GK S VRTSSG FL+ K ++ IEK+I+ ++ IP+E+GE +QVL YE Q Y P
Sbjct: 479 GKSKDSRVRTSSGTFLARGRDK--IVRNIEKKIADFTFIPVEHGEGLQVLHYEVGQKYEP 652
Query: 178 HHDYFSDTFNLKRGGQRIATMLMYLGDNVEGGETHFPSAGS--------DECSCGGKLTK 229
H+DYF D FN K GGQRIAT+LMYL D EGGET FP+A +E S GK K
Sbjct: 653 HYDYFLDEFNTKNGGQRIATVLMYLTDVEEGGETVFPAAKGNFSNVPWYNELSDCGK--K 826
Query: 230 GLCVKPVKGNAVLFWSMGLDGQSDPDSVHGGCPVLAGEKWSATKWMR 276
GL +KP +G+A+LFWSM D D S+HGGCPV+ G KWS+TKW+R
Sbjct: 827 GLSIKPKRGDALLFWSMKPDATLDASSLHGGCPVIKGNKWSSTKWIR 967
>TC86903 similar to GP|21617881|gb|AAM66931.1 prolyl 4-hydroxylase putative
{Arabidopsis thaliana}, partial (68%)
Length = 1310
Score = 191 bits (486), Expect = 2e-49
Identities = 101/204 (49%), Positives = 136/204 (66%), Gaps = 6/204 (2%)
Frame = +3
Query: 78 LSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSHEE 137
LSWSPR L NFL+ EECD+L ++ +L+ S V D +GK I+S+VRTSSGMFL+ ++
Sbjct: 315 LSWSPRAFLYKNFLTDEECDHLIELSKDKLEKSMVADNESGKSIQSEVRTSSGMFLNKQQ 494
Query: 138 RKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRIAT 197
+ ++ IE RI+ ++ +P+ENGE MQVL Y + Y PH D+F D N + GG R+AT
Sbjct: 495 DE--IVSGIEARIAAWTFLPVENGESMQVLHYMNGEKYEPHFDFFHDKANQRLGGHRVAT 668
Query: 198 MLMYLGDNVEGGETHFP------SAGSDECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQ 251
+LMYL + +GGET FP S DE S KG VKP KG+A+LF+S+ LD
Sbjct: 669 VLMYLSNVEKGGETIFPHAEGKLSQPKDE-SWSECAHKGYAVKPRKGDALLFFSLHLDAT 845
Query: 252 SDPDSVHGGCPVLAGEKWSATKWM 275
+D S+HG CPV+ GEKWSATKW+
Sbjct: 846 TDSKSLHGSCPVIEGEKWSATKWI 917
>TC78029 similar to GP|21593296|gb|AAM65245.1 prolyl 4-hydroxylase alpha
subunit-like protein {Arabidopsis thaliana}, partial
(89%)
Length = 1126
Score = 189 bits (479), Expect = 1e-48
Identities = 101/210 (48%), Positives = 135/210 (64%), Gaps = 8/210 (3%)
Frame = +3
Query: 74 KPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFL 133
K + +SW PR + FL+ ECD+L +A LK S V D +G SDVRTSSGMF+
Sbjct: 216 KVKQISWIPRAFVYQGFLTDLECDHLISLAKSELKRSAVADNLSGDSQLSDVRTSSGMFI 395
Query: 134 SHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQ 193
S + K P++ IE RIS ++ +P ENGE +QVLRYE Q Y PH+DYF+D N+ +GG
Sbjct: 396 S--KNKDPIVSGIEDRISAWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIVQGGH 569
Query: 194 RIATMLMYLGDNVEGGETHFPSAGSDECSCGGKLT--------KGLCVKPVKGNAVLFWS 245
R+AT+LMYL + +GGET FP A G K + KG+ VKP +G+A+LF+S
Sbjct: 570 RLATVLMYLTNVTKGGETVFPEAEEPPRRRGSKKSSDLSECAKKGIAVKPRRGDALLFFS 749
Query: 246 MGLDGQSDPDSVHGGCPVLAGEKWSATKWM 275
+ + D +S+H GCPVL GEKWSATKW+
Sbjct: 750 LDTNAIPDTNSLHAGCPVLEGEKWSATKWI 839
>TC78028 similar to GP|22136524|gb|AAM91340.1 unknown protein {Arabidopsis
thaliana}, partial (86%)
Length = 1217
Score = 181 bits (459), Expect = 3e-46
Identities = 99/215 (46%), Positives = 135/215 (62%), Gaps = 13/215 (6%)
Frame = +1
Query: 74 KPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFL 133
K + +SW PR + FL+ ECD+L +A LK S V D +G+ S+VRTSSGMF+
Sbjct: 124 KVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVRTSSGMFI 303
Query: 134 SHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQ 193
S + K ++ IE +IS ++ +P ENGE +QVLRYE Q Y PH+DYF+D N+ RGG
Sbjct: 304 S--KNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIARGGH 477
Query: 194 RIATMLMYLGDNVEGGETHFPSA-------------GSDECSCGGKLTKGLCVKPVKGNA 240
R+AT+LMYL + +GGET FP+A D CG KG+ VKP +G+A
Sbjct: 478 RVATVLMYLTNVTKGGETVFPNAELQESPRHKLSETDEDLSECG---KKGVAVKPRRGDA 648
Query: 241 VLFWSMGLDGQSDPDSVHGGCPVLAGEKWSATKWM 275
+LF+S+ + D S+H GCPV+ GEKWSATKW+
Sbjct: 649 LLFFSLHPNAIPDTLSLHAGCPVIEGEKWSATKWI 753
>BG448236 similar to GP|21537370|gb putative prolyl 4-hydroxylase alpha
subunit {Arabidopsis thaliana}, partial (52%)
Length = 682
Score = 158 bits (399), Expect = 3e-39
Identities = 92/208 (44%), Positives = 125/208 (59%)
Frame = +3
Query: 12 LLTFVTIGMIIGALSQLAFIRRLELEEPFTTTTTTRSLLPRGYTYWNNNNDKEAQILRLG 71
LLT + ++I L L + + TT SL+ + + +K+ Q
Sbjct: 57 LLTLFMLTLVIIVLLALGIL--------YLPNTTDDSLITDRRKIYESLAEKKEQWT--- 203
Query: 72 YVKPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGM 131
E+LSW PR + HNFLS EEC++L +A P L S+VVD+ TGK +S VRTSSGM
Sbjct: 204 ----EILSWEPRAFVYHNFLSKEECEHLINLAKPFLAKSSVVDSKTGKSTESRVRTSSGM 371
Query: 132 FLSHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRG 191
FL + K +I IE+RI+ ++ IP+ENGE +QVL Y + Y PH+DYF D FN K G
Sbjct: 372 FLKRGKDK--IIQNIERRIADFTFIPVENGEGLQVLHYGVGEKYEPHYDYFLDEFNTKNG 545
Query: 192 GQRIATMLMYLGDNVEGGETHFPSAGSD 219
GQR+AT+LMYL D EGGET FP+A ++
Sbjct: 546 GQRVATVLMYLSDVEEGGETVFPAAXAN 629
>TC83824 similar to GP|17381226|gb|AAL36425.1 unknown protein {Arabidopsis
thaliana}, partial (43%)
Length = 680
Score = 98.2 bits (243), Expect = 3e-21
Identities = 52/119 (43%), Positives = 73/119 (60%), Gaps = 2/119 (1%)
Frame = +3
Query: 160 NGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRIATMLMYLGDNVEGGETHFPSAG-- 217
+GE +LRYE Q Y H+D F+ + QR+A+ L+YL D EGGET FP
Sbjct: 3 HGEAFNILRYEVGQRYNSHYDAFNPDEYGPQKSQRVASFLLYLTDVEEGGETMFPFENGL 182
Query: 218 SDECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQSDPDSVHGGCPVLAGEKWSATKWMR 276
+ + + G + GL VKP +G+ +LF+S+ +G D S+HG CPV+ GEKW ATKW+R
Sbjct: 183 NMDGTYGYEDCVGLRVKPRQGDGLLFYSLLPNGTIDQTSLHGSCPVIKGEKWVATKWIR 359
>AJ388831 weakly similar to GP|10177121|dbj prolyl 4-hydroxylase alpha
subunit-like protein {Arabidopsis thaliana}, partial
(33%)
Length = 505
Score = 79.3 bits (194), Expect = 1e-15
Identities = 38/87 (43%), Positives = 56/87 (63%)
Frame = +3
Query: 76 EVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSH 135
+++SW PR L HNFL+ EEC++L +A P + S V+D TG G+ S RTSSG FL
Sbjct: 198 QIISWEPRAFLYHNFLTKEECEHLINIAKPSMHKSAVIDEETGNGVDSSERTSSGAFLKR 377
Query: 136 EERKYPMIHAIEKRISVYSQIPIENGE 162
+ ++ IE+RI+ ++ IP E+GE
Sbjct: 378 GSDR--IVKNIERRIADFTFIPXEHGE 452
>BG447864 weakly similar to GP|10177121|db prolyl 4-hydroxylase alpha
subunit-like protein {Arabidopsis thaliana}, partial
(50%)
Length = 639
Score = 66.6 bits (161), Expect = 1e-11
Identities = 35/75 (46%), Positives = 46/75 (60%)
Frame = +3
Query: 60 NNDKEAQILRLGYVKPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGK 119
N+D+E + G EV+SW P + HNFL+ EEC+YL + P + STVVD+ TGK
Sbjct: 372 NDDEEGK----GEQWVEVVSWEPXAFVYHNFLTKEECEYLIDIXKPSMHKSTVVDSETGK 539
Query: 120 GIKSDVRTSSGMFLS 134
S VRT SG FL+
Sbjct: 540 SKDSXVRTXSGTFLA 584
>BQ144147
Length = 729
Score = 45.8 bits (107), Expect = 2e-05
Identities = 19/64 (29%), Positives = 36/64 (55%)
Frame = +2
Query: 154 SQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRIATMLMYLGDNVEGGETHF 213
+ +PIENG+ + + RY+ + P+ +Y + N+ +GG R T+LM + + +G F
Sbjct: 92 TSLPIENGDNLHIWRYKHGHNHNPNDNYSTHKINIVQGGHRPPTVLMLITNETKGNRN*F 271
Query: 214 PSAG 217
+G
Sbjct: 272 SQSG 283
>BF518749
Length = 416
Score = 44.3 bits (103), Expect = 5e-05
Identities = 43/160 (26%), Positives = 71/160 (43%)
Frame = +1
Query: 8 IVFGLLTFVTIGMIIGALSQLAFIRRLELEEPFTTTTTTRSLLPRGYTYWNNNNDKEAQI 67
+ F L TF T+ ++ + S R EL + L R Y++N D +
Sbjct: 28 LTFLLFTFFTLSLLTTSFSD----SRKELRNK--NENVLKQL--RNSVYYSNRIDPSRVV 183
Query: 68 LRLGYVKPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRT 127
+SW PR+ L FLS +ECDYL IS + ++G G S
Sbjct: 184 Q---------ISWQPRVFLYKGFLSDKECDYL---------ISLAQEKSSGNGGYSKKEE 309
Query: 128 SSGMFLSHEERKYPMIHAIEKRISVYSQIPIENGELMQVL 167
+S L ++ ++ IE+R+SV++ + EN + +QV+
Sbjct: 310 TS---LDMDD---DIVKRIEERLSVWTFLSKENSKPLQVM 411
>BF521324 similar to GP|18086437|gb| AT3g28480/MFJ20_16 {Arabidopsis
thaliana}, partial (15%)
Length = 285
Score = 38.5 bits (88), Expect = 0.003
Identities = 18/37 (48%), Positives = 24/37 (64%)
Frame = +1
Query: 78 LSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVD 114
LSWSPR L +NFL+ EECD+L ++ L+ S D
Sbjct: 166 LSWSPRAFLYNNFLTDEECDHLIELSKDNLEKSMAAD 276
>BE941277
Length = 361
Score = 29.3 bits (64), Expect = 1.8
Identities = 24/74 (32%), Positives = 34/74 (45%), Gaps = 1/74 (1%)
Frame = +1
Query: 82 PRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDV-RTSSGMFLSHEERKY 140
P + LL N SY C++ RG K+ D G+GIK ++ + F S E Y
Sbjct: 70 PSVYLLPNMWSYTTCEF-RGA-----KLLGSADQGGGEGIKIELNQLKPYYFASDEGNAY 231
Query: 141 PMIHAIEKRISVYS 154
I + K I+V S
Sbjct: 232 DCIAGLTKFIAVPS 273
>TC79352 weakly similar to PIR|T01829|T01829 hypothetical protein T15F16.10 -
Arabidopsis thaliana, partial (31%)
Length = 1700
Score = 28.1 bits (61), Expect = 3.9
Identities = 18/64 (28%), Positives = 35/64 (54%), Gaps = 2/64 (3%)
Frame = +1
Query: 140 YPMIHAIEKRISVYSQIPIENGELMQVLRY--EKNQYYRPHHDYFSDTFNLKRGGQRIAT 197
+ M + +E +S+ + + +E+ LM+ L + K++Y RPHH F + GQ +
Sbjct: 1246 FSMKYVLEA*VSLKTWMNLES--LMRRLTFLLPKHRYLRPHHRLFLERMIFPLVGQLMNW 1419
Query: 198 MLMY 201
+L+Y
Sbjct: 1420 LLLY 1431
>TC89186 similar to PIR|G96563|G96563 probable coatomer complex subunit
33791-27676 [imported] - Arabidopsis thaliana, partial
(32%)
Length = 1389
Score = 27.7 bits (60), Expect = 5.1
Identities = 12/68 (17%), Positives = 36/68 (52%)
Frame = +1
Query: 132 FLSHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRG 191
+++H ++ + + + + + + P+ENG+ L E+N+ + +++++ N + G
Sbjct: 718 YINHADKSHVTLVEAFRNMQIEEEEPLENGDSNHELT-EQNEEHYTEEEHYTEEQNGEEG 894
Query: 192 GQRIATML 199
Q A ++
Sbjct: 895 SQEEAVVV 918
>TC85105 similar to GP|20160725|dbj|BAB89667. putative transaldolase {Oryza
sativa (japonica cultivar-group)}, partial (32%)
Length = 731
Score = 27.3 bits (59), Expect = 6.7
Identities = 15/55 (27%), Positives = 30/55 (54%)
Frame = +2
Query: 131 MFLSHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDT 185
+F+ +++R MI +++ + + +E+ L+ +LRY K QY HH + T
Sbjct: 221 IFMRNKDRVLTMIIYVDQFQIWFHSLLLESEVLLPILRYLKRQY---HHQMLTIT 376
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.138 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,446,535
Number of Sequences: 36976
Number of extensions: 133858
Number of successful extensions: 631
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 617
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 619
length of query: 281
length of database: 9,014,727
effective HSP length: 95
effective length of query: 186
effective length of database: 5,502,007
effective search space: 1023373302
effective search space used: 1023373302
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC149471.21


