

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149305.4 - phase: 0 /partial
(542 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
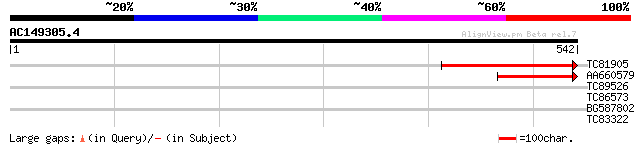
TC81905 weakly similar to GP|22830972|dbj|BAC15836. contains EST... 270 1e-72
AA660579 similar to PIR|T51532|T51 hypothetical protein T20K14_1... 160 2e-39
TC89526 similar to GP|14596139|gb|AAK68797.1 Unknown protein {Ar... 30 3.1
TC86573 similar to GP|15292907|gb|AAK92824.1 unknown protein {Ar... 28 6.9
BG587802 similar to GP|20197543|gb putative beta-1 3-glucanase {... 28 9.0
TC83322 similar to PIR|E84706|E84706 hypothetical protein At2g30... 28 9.0
>TC81905 weakly similar to GP|22830972|dbj|BAC15836. contains ESTs
C26545(C12561) AU091755(C12561)~similar to Nipped-B gene
product, partial (8%)
Length = 1075
Score = 270 bits (690), Expect = 1e-72
Identities = 130/130 (100%), Positives = 130/130 (100%)
Frame = +1
Query: 413 PDEPLYLIYTINRVVQVRAGPLEANFKAWSSSLLQSEGQGTPHGNGMYQRATDETIHSTQ 472
PDEPLYLIYTINRVVQVRAGPLEANFKAWSSSLLQSEGQGTPHGNGMYQRATDETIHSTQ
Sbjct: 1 PDEPLYLIYTINRVVQVRAGPLEANFKAWSSSLLQSEGQGTPHGNGMYQRATDETIHSTQ 180
Query: 473 GQSMDLNGPFQQNVDVQPYVDDMTSVDLNGTNHQLPDYPLSHNGRLKVKPQAAGFADSFT 532
GQSMDLNGPFQQNVDVQPYVDDMTSVDLNGTNHQLPDYPLSHNGRLKVKPQAAGFADSFT
Sbjct: 181 GQSMDLNGPFQQNVDVQPYVDDMTSVDLNGTNHQLPDYPLSHNGRLKVKPQAAGFADSFT 360
Query: 533 FSKDDLEKVQ 542
FSKDDLEKVQ
Sbjct: 361 FSKDDLEKVQ 390
>AA660579 similar to PIR|T51532|T51 hypothetical protein T20K14_150 -
Arabidopsis thaliana, partial (2%)
Length = 638
Score = 160 bits (404), Expect = 2e-39
Identities = 76/76 (100%), Positives = 76/76 (100%)
Frame = +3
Query: 467 TIHSTQGQSMDLNGPFQQNVDVQPYVDDMTSVDLNGTNHQLPDYPLSHNGRLKVKPQAAG 526
TIHSTQGQSMDLNGPFQQNVDVQPYVDDMTSVDLNGTNHQLPDYPLSHNGRLKVKPQAAG
Sbjct: 3 TIHSTQGQSMDLNGPFQQNVDVQPYVDDMTSVDLNGTNHQLPDYPLSHNGRLKVKPQAAG 182
Query: 527 FADSFTFSKDDLEKVQ 542
FADSFTFSKDDLEKVQ
Sbjct: 183 FADSFTFSKDDLEKVQ 230
>TC89526 similar to GP|14596139|gb|AAK68797.1 Unknown protein {Arabidopsis
thaliana}, partial (40%)
Length = 1105
Score = 29.6 bits (65), Expect = 3.1
Identities = 10/27 (37%), Positives = 16/27 (59%)
Frame = -3
Query: 281 TDPLESNSKMAHHLLMNMHEKYPAFFE 307
T P+ K HH N+HE++P +F+
Sbjct: 833 TFPIIPKKKQCHHFKNNLHERWPRYFK 753
>TC86573 similar to GP|15292907|gb|AAK92824.1 unknown protein {Arabidopsis
thaliana}, partial (92%)
Length = 2060
Score = 28.5 bits (62), Expect = 6.9
Identities = 29/106 (27%), Positives = 47/106 (43%), Gaps = 18/106 (16%)
Frame = +3
Query: 423 INRVVQVRAGPLEANFKA------WSSSLLQSEGQGTPHGNGMYQRATDETIHSTQGQSM 476
++ VV PLE++F+ S+SL + G + N ++R T + ST+ +
Sbjct: 1122 VSGVVTCGNVPLESSFQKVPRCVYTSTSLYEYRGWDSKAANSKHRRFT-WGLRSTERHVV 1298
Query: 477 DLN-GPFQQNVD----------VQPYVDDMTSVDLNGTNHQL-PDY 510
D FQ + V PYVDD +D+N N + PD+
Sbjct: 1299 DFYISDFQSGLRALVKTGFGARVTPYVDDSVVIDVNPENKDMSPDF 1436
>BG587802 similar to GP|20197543|gb putative beta-1 3-glucanase {Arabidopsis
thaliana}, partial (48%)
Length = 753
Score = 28.1 bits (61), Expect = 9.0
Identities = 11/36 (30%), Positives = 23/36 (63%)
Frame = -1
Query: 25 PSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSE 60
PS + + +D+ ++ +RH ++ ++ CIK +C M E
Sbjct: 651 PS*LGD*NRDISKLQIRHCKISSINNCIKNICIMQE 544
>TC83322 similar to PIR|E84706|E84706 hypothetical protein At2g30280
[imported] - Arabidopsis thaliana, partial (19%)
Length = 725
Score = 28.1 bits (61), Expect = 9.0
Identities = 11/25 (44%), Positives = 18/25 (72%)
Frame = +1
Query: 12 IIDSVLPLLRKLPPSIVEELEQDLK 36
++ S LPLLR++ P+ E+E D+K
Sbjct: 550 LLSSFLPLLREVIPNAAAEIEDDIK 624
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.137 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,560,057
Number of Sequences: 36976
Number of extensions: 210422
Number of successful extensions: 996
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 991
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 996
length of query: 542
length of database: 9,014,727
effective HSP length: 101
effective length of query: 441
effective length of database: 5,280,151
effective search space: 2328546591
effective search space used: 2328546591
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC149305.4


