

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
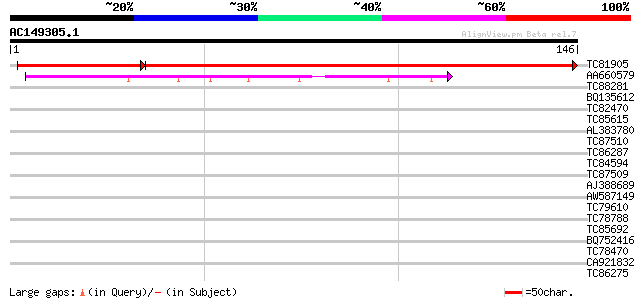
Query= AC149305.1 + phase: 0 /pseudo
(146 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC81905 weakly similar to GP|22830972|dbj|BAC15836. contains EST... 229 7e-75
AA660579 similar to PIR|T51532|T51 hypothetical protein T20K14_1... 68 1e-12
TC88281 similar to PIR|B84784|B84784 hypothetical protein At2g36... 28 1.7
BQ135612 similar to GP|15028978|emb type II keratin E2 {Oncorhyn... 28 1.7
TC82470 homologue to GP|23497460|gb|AAN37003.1 hypothetical prot... 23 2.6
TC85615 similar to GP|10178248|dbj|BAB11680. glucosyltransferase... 27 2.9
AL383780 27 2.9
TC87510 similar to GP|9758854|dbj|BAB09380.1 gene_id:K3K3.16~unk... 26 5.0
TC86287 similar to GP|23617108|dbj|BAC20790. contains ESTs AU056... 26 5.0
TC84594 similar to GP|23497092|gb|AAN36641.1 hypothetical protei... 26 5.0
TC87509 similar to GP|9758854|dbj|BAB09380.1 gene_id:K3K3.16~unk... 26 5.0
AJ388689 26 5.0
AW587149 similar to PIR|B85431|B85 trichohyalin like protein [im... 26 5.0
TC79610 similar to GP|16323294|gb|AAL15402.1 AT5g19430/F7K24_180... 26 6.5
TC78788 similar to GP|21537190|gb|AAM61531.1 putative SKP1-like ... 26 6.5
TC85692 similar to PIR|T05351|T05351 cellulose synthase (EC 2.4.... 26 6.5
BQ752416 homologue to SP|P52810|RS9_ 40S ribosomal protein S9 (S... 25 8.5
TC78470 similar to GP|21592963|gb|AAM64912.1 unknown {Arabidopsi... 25 8.5
CA921832 similar to GP|14194103|gb AT3g22390/MCB17_12 {Arabidops... 25 8.5
TC86275 similar to PIR|E96603|E96603 unknown protein F14G9.26 [i... 25 8.5
>TC81905 weakly similar to GP|22830972|dbj|BAC15836. contains ESTs
C26545(C12561) AU091755(C12561)~similar to Nipped-B gene
product, partial (8%)
Length = 1075
Score = 229 bits (585), Expect(2) = 7e-75
Identities = 111/111 (100%), Positives = 111/111 (100%)
Frame = +2
Query: 36 SEPPKPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIKR 95
SEPPKPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIKR
Sbjct: 506 SEPPKPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIKR 685
Query: 96 KRPTQTPRKVRKSGPPMVGGDNDEDDDEDWAGGSRNISFSGGRRSSLRNSR 146
KRPTQTPRKVRKSGPPMVGGDNDEDDDEDWAGGSRNISFSGGRRSSLRNSR
Sbjct: 686 KRPTQTPRKVRKSGPPMVGGDNDEDDDEDWAGGSRNISFSGGRRSSLRNSR 838
Score = 67.0 bits (162), Expect(2) = 7e-75
Identities = 33/33 (100%), Positives = 33/33 (100%)
Frame = +3
Query: 3 FCRLTVYLRLRCNFF*S*RDT*KSCIVSMMLDV 35
FCRLTVYLRLRCNFF*S*RDT*KSCIVSMMLDV
Sbjct: 393 FCRLTVYLRLRCNFF*S*RDT*KSCIVSMMLDV 491
>AA660579 similar to PIR|T51532|T51 hypothetical protein T20K14_150 -
Arabidopsis thaliana, partial (2%)
Length = 638
Score = 68.2 bits (165), Expect = 1e-12
Identities = 61/129 (47%), Positives = 69/129 (53%), Gaps = 19/129 (14%)
Frame = +1
Query: 5 RLTVYLRLRCNFF*S*RDT*KSCIV-----SMMLDVSEPPKPG-DVFSRQSI-PFNIGES 57
RLTVYLRLRCNFF*S*RDT*KSC SEPPKPG V SRQ+ PFNIGES
Sbjct: 229 RLTVYLRLRCNFF*S*RDT*KSCYSLDDARCQAYSPSEPPKPG*CVSSRQNYSPFNIGES 408
Query: 58 QFS--LPTSPQELIQRYQ--------EFKNALKEDTVDYSLYTANIKRK-RPTQTPRKVR 106
S S Q+LI + + KN + SL+ K K QTP K+
Sbjct: 409 SSSSFAYNSSQKLIPKISXNF*EMPLKGKNT---SGIIPSLHXQIXKXKENQHQTPPKIP 579
Query: 107 K-SGPPMVG 114
+ PPMVG
Sbjct: 580 EIXDPPMVG 606
>TC88281 similar to PIR|B84784|B84784 hypothetical protein At2g36740
[imported] - Arabidopsis thaliana, partial (59%)
Length = 1086
Score = 27.7 bits (60), Expect = 1.7
Identities = 18/63 (28%), Positives = 31/63 (48%)
Frame = +2
Query: 62 PTSPQELIQRYQEFKNALKEDTVDYSLYTANIKRKRPTQTPRKVRKSGPPMVGGDNDEDD 121
P +E Q E N K+ + Y T + K+K+ + + PP + GD+D+DD
Sbjct: 257 PEPDEEQPQNDDERMN--KKKRLVYPGKTTSAKKKKKKKILSNL--DDPPKIDGDDDQDD 424
Query: 122 DED 124
D++
Sbjct: 425 DKN 433
>BQ135612 similar to GP|15028978|emb type II keratin E2 {Oncorhynchus
mykiss}, partial (8%)
Length = 938
Score = 27.7 bits (60), Expect = 1.7
Identities = 11/24 (45%), Positives = 14/24 (57%)
Frame = +1
Query: 92 NIKRKRPTQTPRKVRKSGPPMVGG 115
N K+ + P+K RK GPP GG
Sbjct: 31 NSSTKKGGKNPKKKRKGGPPFFGG 102
>TC82470 homologue to GP|23497460|gb|AAN37003.1 hypothetical protein
{Plasmodium falciparum 3D7}, partial (8%)
Length = 1064
Score = 23.1 bits (48), Expect(2) = 2.6
Identities = 9/22 (40%), Positives = 14/22 (62%)
Frame = +2
Query: 103 RKVRKSGPPMVGGDNDEDDDED 124
+K + S P D+D+DDD+D
Sbjct: 5 KKSKVSPTPYDDDDDDDDDDDD 70
Score = 22.3 bits (46), Expect(2) = 2.6
Identities = 10/20 (50%), Positives = 14/20 (70%)
Frame = +2
Query: 116 DNDEDDDEDWAGGSRNISFS 135
D+D+DDD+D S + SFS
Sbjct: 56 DDDDDDDDDDDDMSTDKSFS 115
>TC85615 similar to GP|10178248|dbj|BAB11680. glucosyltransferase-like
protein {Arabidopsis thaliana}, partial (90%)
Length = 2067
Score = 26.9 bits (58), Expect = 2.9
Identities = 15/62 (24%), Positives = 28/62 (44%)
Frame = +2
Query: 25 KSCIVSMMLDVSEPPKPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTV 84
K C ++ D P P F R++IPF +G + +L + + ++E ++
Sbjct: 755 KHCEYVVIFDADFSPPPD--FLRRAIPFLVGNPEIALVQGRWRFVNANECLLTRMQEMSL 928
Query: 85 DY 86
DY
Sbjct: 929 DY 934
>AL383780
Length = 222
Score = 26.9 bits (58), Expect = 2.9
Identities = 16/54 (29%), Positives = 26/54 (47%)
Frame = +2
Query: 60 SLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIKRKRPTQTPRKVRKSGPPMV 113
SLP SP+ ++++ Q+ AL+ D + I+ R TP V P +V
Sbjct: 2 SLPASPESVVEKRQKLLGALQRCDEDLKVLKKIIESVR---TPEPVSSPSPAVV 154
>TC87510 similar to GP|9758854|dbj|BAB09380.1 gene_id:K3K3.16~unknown
protein {Arabidopsis thaliana}, partial (28%)
Length = 440
Score = 26.2 bits (56), Expect = 5.0
Identities = 14/39 (35%), Positives = 19/39 (47%), Gaps = 6/39 (15%)
Frame = -2
Query: 114 GGDNDEDDDEDWAGGS------RNISFSGGRRSSLRNSR 146
G +DDD+DW GG R++S S R +N R
Sbjct: 145 GSKRFDDDDDDWFGGGIG**TMRDVSSSTSERRENQN*R 29
>TC86287 similar to GP|23617108|dbj|BAC20790. contains ESTs AU056959(S21019)
AU056960(S21019)~similar to Arabidopsis thaliana
chromosome 5, partial (82%)
Length = 1302
Score = 26.2 bits (56), Expect = 5.0
Identities = 9/15 (60%), Positives = 12/15 (80%)
Frame = -3
Query: 114 GGDNDEDDDEDWAGG 128
GGD DEDDD+++ G
Sbjct: 241 GGDGDEDDDDEFEDG 197
>TC84594 similar to GP|23497092|gb|AAN36641.1 hypothetical protein
{Plasmodium falciparum 3D7}, partial (0%)
Length = 791
Score = 26.2 bits (56), Expect = 5.0
Identities = 14/32 (43%), Positives = 18/32 (55%), Gaps = 5/32 (15%)
Frame = +2
Query: 115 GDNDEDDDEDWAGGSRNIS-----FSGGRRSS 141
G++DEDDDED GG + G +RSS
Sbjct: 434 GEDDEDDDEDDDGGDTVVDRLVGVGDGSKRSS 529
>TC87509 similar to GP|9758854|dbj|BAB09380.1 gene_id:K3K3.16~unknown
protein {Arabidopsis thaliana}, partial (40%)
Length = 698
Score = 26.2 bits (56), Expect = 5.0
Identities = 14/39 (35%), Positives = 19/39 (47%), Gaps = 6/39 (15%)
Frame = -1
Query: 114 GGDNDEDDDEDWAGGS------RNISFSGGRRSSLRNSR 146
G +DDD+DW GG R++S S R +N R
Sbjct: 143 GSKRFDDDDDDWFGGGIG**TMRDVSSSTSERRENQN*R 27
>AJ388689
Length = 504
Score = 26.2 bits (56), Expect = 5.0
Identities = 12/22 (54%), Positives = 14/22 (63%)
Frame = +2
Query: 52 FNIGESQFSLPTSPQELIQRYQ 73
F+ G S FSLP P L+ RYQ
Sbjct: 275 FSHGCSVFSLPQGPYRLMMRYQ 340
>AW587149 similar to PIR|B85431|B85 trichohyalin like protein [imported] -
Arabidopsis thaliana, partial (1%)
Length = 634
Score = 26.2 bits (56), Expect = 5.0
Identities = 12/22 (54%), Positives = 16/22 (72%)
Frame = +3
Query: 115 GDNDEDDDEDWAGGSRNISFSG 136
GD+DE+DDE W+ + SFSG
Sbjct: 414 GDSDEEDDEVWSPEETD-SFSG 476
>TC79610 similar to GP|16323294|gb|AAL15402.1 AT5g19430/F7K24_180
{Arabidopsis thaliana}, partial (54%)
Length = 789
Score = 25.8 bits (55), Expect = 6.5
Identities = 13/25 (52%), Positives = 18/25 (72%), Gaps = 1/25 (4%)
Frame = +1
Query: 116 DNDED-DDEDWAGGSRNISFSGGRR 139
+ND+D DDED+ GGS ++ G RR
Sbjct: 562 ENDDDLDDEDYYGGSASVRI-GNRR 633
>TC78788 similar to GP|21537190|gb|AAM61531.1 putative SKP1-like protein
{Arabidopsis thaliana}, partial (66%)
Length = 1567
Score = 25.8 bits (55), Expect = 6.5
Identities = 16/60 (26%), Positives = 27/60 (44%), Gaps = 8/60 (13%)
Frame = +3
Query: 73 QEFKNALKEDTVDYSLYTAN--------IKRKRPTQTPRKVRKSGPPMVGGDNDEDDDED 124
Q+ ++LKE +V + N + RP +T PM+ D+D+DD +D
Sbjct: 939 QQKNSSLKESSVPHKKAEVNGHNNRHQSSEADRPCETSSLFHTDYDPMIEFDDDDDDIDD 1118
>TC85692 similar to PIR|T05351|T05351 cellulose synthase (EC 2.4.1.-)
catalytic chain RSW1 - Arabidopsis thaliana, partial
(18%)
Length = 854
Score = 25.8 bits (55), Expect = 6.5
Identities = 12/27 (44%), Positives = 15/27 (55%)
Frame = +3
Query: 98 PTQTPRKVRKSGPPMVGGDNDEDDDED 124
P R R G P V GD++EDD +D
Sbjct: 390 PQCKTRYKRLRGSPRVDGDDEEDDVDD 470
>BQ752416 homologue to SP|P52810|RS9_ 40S ribosomal protein S9 (S7).
{Podospora anserina}, partial (94%)
Length = 717
Score = 25.4 bits (54), Expect = 8.5
Identities = 13/39 (33%), Positives = 22/39 (56%)
Frame = +1
Query: 84 VDYSLYTANIKRKRPTQTPRKVRKSGPPMVGGDNDEDDD 122
+D++L T+ RP + RK K+ GGD +ED++
Sbjct: 472 IDFAL-TSPFGGGRPGRVRRKKAKAAEKADGGDEEEDEE 585
>TC78470 similar to GP|21592963|gb|AAM64912.1 unknown {Arabidopsis
thaliana}, partial (70%)
Length = 1111
Score = 25.4 bits (54), Expect = 8.5
Identities = 8/12 (66%), Positives = 10/12 (82%)
Frame = -1
Query: 114 GGDNDEDDDEDW 125
GGD+D+DDD W
Sbjct: 49 GGDDDDDDDGSW 14
>CA921832 similar to GP|14194103|gb AT3g22390/MCB17_12 {Arabidopsis
thaliana}, partial (3%)
Length = 802
Score = 25.4 bits (54), Expect = 8.5
Identities = 14/39 (35%), Positives = 19/39 (47%), Gaps = 4/39 (10%)
Frame = -1
Query: 94 KRKRPTQTPRKVRKSGPPMVG----GDNDEDDDEDWAGG 128
KR+R + VR+ G D ++DDDED GG
Sbjct: 394 KRRRNDRLMHGVREDGGEDTSEESINDEEDDDDEDGGGG 278
>TC86275 similar to PIR|E96603|E96603 unknown protein F14G9.26 [imported] -
Arabidopsis thaliana, partial (51%)
Length = 961
Score = 25.4 bits (54), Expect = 8.5
Identities = 15/51 (29%), Positives = 24/51 (46%), Gaps = 3/51 (5%)
Frame = +2
Query: 90 TANIKRKRPTQTPRKVRKSGPPMVGGDND---EDDDEDWAGGSRNISFSGG 137
T + R+ PT T ++ P + G + DDD+ GG ++SGG
Sbjct: 440 TQTVLREAPTITHLPGKEKSPQIDDGGSGFPPRDDDDGGGGGGGGGNWSGG 592
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.329 0.143 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,533,323
Number of Sequences: 36976
Number of extensions: 56873
Number of successful extensions: 613
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 512
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 581
length of query: 146
length of database: 9,014,727
effective HSP length: 87
effective length of query: 59
effective length of database: 5,797,815
effective search space: 342071085
effective search space used: 342071085
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC149305.1


