

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149293.2 - phase: 2
(212 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
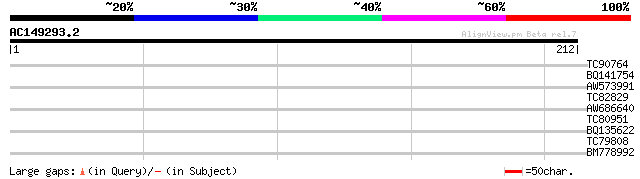
TC90764 similar to PIR|T05666|T05666 hypothetical protein F22I13... 33 0.11
BQ141754 similar to GP|1903309|emb| orf 1035 {Spodoptera littora... 29 1.2
AW573991 28 2.0
TC82829 weakly similar to GP|14334592|gb|AAK59474.1 unknown prot... 28 2.6
AW686640 28 2.6
TC80951 similar to PIR|D96519|D96519 mysoin-like protein 11013-... 28 3.4
BQ135622 27 4.4
TC79808 similar to GP|2443886|gb|AAB71479.1| Unknown protein {Ar... 27 5.8
BM778992 similar to GP|21594024|gb| unknown {Arabidopsis thalian... 26 9.8
>TC90764 similar to PIR|T05666|T05666 hypothetical protein F22I13.150 -
Arabidopsis thaliana, partial (57%)
Length = 1219
Score = 32.7 bits (73), Expect = 0.11
Identities = 36/122 (29%), Positives = 50/122 (40%), Gaps = 8/122 (6%)
Frame = +2
Query: 24 GMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAAYMMMRTLNMK- 82
G KD+ P+ L ++ +L Y G+AGAA +T+ SQ + +M+ LN +
Sbjct: 755 GFKDTKTPVICLGIGNLSAVFLFPLLMYYFKLGVAGAAISTVLSQYIGTLLMIWCLNKRA 934
Query: 83 -------GYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHTMAA 135
G F I SG +LG V TM +L A G MAA
Sbjct: 935 VLLPPKMGNLQFGGYIKSGG---FVLGRTLAVLTTM-------TLGTSMAARHGPVAMAA 1084
Query: 136 HQ 137
HQ
Sbjct: 1085HQ 1090
>BQ141754 similar to GP|1903309|emb| orf 1035 {Spodoptera littoralis
nucleopolyhedrovirus}, partial (4%)
Length = 1308
Score = 29.3 bits (64), Expect = 1.2
Identities = 16/41 (39%), Positives = 22/41 (53%)
Frame = -3
Query: 118 YSLLIYFATSMGTHTMAAHQYGVNLSRKLLNHLCLNCYMEL 158
YSLL YF S T T++ Y + L+ +C NCYM +
Sbjct: 868 YSLLYYFCRSCVTSTLSTS*YELYCFSILV--ICYNCYMSV 752
>AW573991
Length = 652
Score = 28.5 bits (62), Expect = 2.0
Identities = 12/22 (54%), Positives = 15/22 (67%)
Frame = -1
Query: 112 MSKVAFYSLLIYFATSMGTHTM 133
+ K YS+ IY TS+GTHTM
Sbjct: 601 IKKKLLYSIFIYEITSIGTHTM 536
>TC82829 weakly similar to GP|14334592|gb|AAK59474.1 unknown protein
{Arabidopsis thaliana}, partial (12%)
Length = 703
Score = 28.1 bits (61), Expect = 2.6
Identities = 16/58 (27%), Positives = 27/58 (45%)
Frame = +1
Query: 92 PSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHTMAAHQYGVNLSRKLLNH 149
PS F+ + G+ VF ++S F+S L+ F + T Q+ + L L+H
Sbjct: 169 PSSPSFLMLHGM---VFTLLISSCLFFSFLLVFLLHLFIRTRDQDQHNLLLGNHYLDH 333
>AW686640
Length = 518
Score = 28.1 bits (61), Expect = 2.6
Identities = 10/38 (26%), Positives = 23/38 (60%)
Frame = +3
Query: 97 FITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHTMA 134
F ++L P+++T+ ++FYS +I+ + H++A
Sbjct: 3 FSSLLSFILPIYITLFFSLSFYSTIIFLSYFNTFHSLA 116
>TC80951 similar to PIR|D96519|D96519 mysoin-like protein 11013-7318
[imported] - Arabidopsis thaliana, partial (14%)
Length = 1050
Score = 27.7 bits (60), Expect = 3.4
Identities = 19/60 (31%), Positives = 24/60 (39%), Gaps = 11/60 (18%)
Frame = -3
Query: 103 LAAPVFMTMMSKVAFYSLLIY-----------FATSMGTHTMAAHQYGVNLSRKLLNHLC 151
LA FM M + SLL+ FA GTHT + Q+ V + HLC
Sbjct: 433 LAVYAFMVHMDGRFWRSLLLITCLALCFQNLKFAKCFGTHTS*SLQFAVTFCQSFFQHLC 254
>BQ135622
Length = 798
Score = 27.3 bits (59), Expect = 4.4
Identities = 10/17 (58%), Positives = 15/17 (87%)
Frame = -3
Query: 141 NLSRKLLNHLCLNCYME 157
N+SR+L+N +CLN Y+E
Sbjct: 607 NISRELINIVCLNVYVE 557
>TC79808 similar to GP|2443886|gb|AAB71479.1| Unknown protein {Arabidopsis
thaliana}, partial (37%)
Length = 1001
Score = 26.9 bits (58), Expect = 5.8
Identities = 19/73 (26%), Positives = 40/73 (54%), Gaps = 2/73 (2%)
Frame = -1
Query: 61 AWATMASQVVAAYMMMRTL--NMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFY 118
A+ T V+ A M ++ +++ ++A + R+FIT L +++ + +T++S +
Sbjct: 623 AFLTAIRSVIVARSMSGSVPDSLRNFSAASTKPSVTRKFITNLFMSSFLILTLLSFFSPS 444
Query: 119 SLLIYFATSMGTH 131
SL+I F + TH
Sbjct: 443 SLVITFFDAAFTH 405
>BM778992 similar to GP|21594024|gb| unknown {Arabidopsis thaliana}, partial
(38%)
Length = 682
Score = 26.2 bits (56), Expect = 9.8
Identities = 20/93 (21%), Positives = 39/93 (41%), Gaps = 20/93 (21%)
Frame = -1
Query: 93 SGREFITILGLAAPVFMTMMSKV---------AFYSLLIYFATSMGTHTMAAHQYGVNLS 143
+G FIT +GL + ++ V + +S L+ ++ +G T+ Y +
Sbjct: 418 NGENFITKIGLVYCSLLLLVGNVILF*A*GCRSLFSFLLQYSR*VGQATI*ILYYHCLGN 239
Query: 144 RKLLNHLCL-----------NCYMELIGICQRP 165
KL + + NCY ++G+C+ P
Sbjct: 238 GKLQIVIAIVFQ*SIEGW*NNCYFNIVGVCKMP 140
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.338 0.146 0.479
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,045,114
Number of Sequences: 36976
Number of extensions: 103798
Number of successful extensions: 842
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 840
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 842
length of query: 212
length of database: 9,014,727
effective HSP length: 92
effective length of query: 120
effective length of database: 5,612,935
effective search space: 673552200
effective search space used: 673552200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC149293.2


