

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149205.5 + phase: 0 /pseudo
(738 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
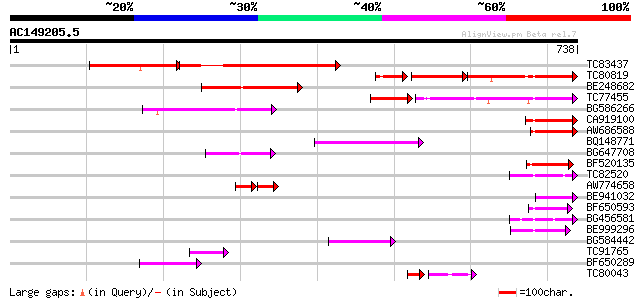
Sequences producing significant alignments: (bits) Value
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 241 1e-86
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 135 2e-62
BE248682 similar to GP|18568269|gb putative gag-pol polyprotein ... 125 8e-29
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 92 9e-28
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 110 2e-24
CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [i... 86 4e-17
AW686588 80 3e-15
BQ148771 65 9e-11
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 62 6e-10
BF520135 61 1e-09
TC82520 59 9e-09
AW774658 similar to GP|2808681|emb| Hcr9-4B {Lycopersicon hirsut... 44 1e-08
BE941032 54 3e-07
BF650593 53 4e-07
BG456581 50 4e-06
BE999296 48 2e-05
BG584442 47 3e-05
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 45 8e-05
BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgari... 44 3e-04
TC80043 similar to GP|9759561|dbj|BAB11163.1 MtN21 nodulin prote... 41 4e-04
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 241 bits (616), Expect(2) = 1e-86
Identities = 130/210 (61%), Positives = 150/210 (70%)
Frame = +2
Query: 221 MAFPLLWRKWIKECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLM 280
M+F +LWRKWIKEC+ST T S+LVNGSPT NVLM
Sbjct: 380 MSFLVLWRKWIKECVSTATTSVLVNGSPT---------------------------NVLM 478
Query: 281 KSVVENNLFKGYVVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKV 340
KS+V+ LF Y G N VVVSHLQFA+DTLLL K+WAN+RALRA L +F MSGLKV
Sbjct: 479 KSLVQTQLFTRYSFGVVNPVVVSHLQFANDTLLLETKNWANIRALRAALVIF*AMSGLKV 658
Query: 341 NFNKSLLVGVNIGESWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTR 400
NF+KS LV VNI SWL EAASVL+ KVGK+ FLYLG+ I G+ R+L+FWEP+V+ IK R
Sbjct: 659 NFHKSGLVCVNIAPSWLSEAASVLSWKVGKVPFLYLGMPIEGNSRRLSFWEPIVNRIKAR 838
Query: 401 LSGWQSCFLSFGGRLVLLKSVLTSLPVYAL 430
L+GW S FLSFGGRLVLLKSVLTSL VYAL
Sbjct: 839 LTGWNSRFLSFGGRLVLLKSVLTSLSVYAL 928
Score = 97.8 bits (242), Expect(2) = 1e-86
Identities = 64/125 (51%), Positives = 77/125 (61%), Gaps = 6/125 (4%)
Frame = +1
Query: 105 FRFSSKWKIIKRYQFYFYSSYS*SGQPSESK*FSTNFLGWESV*NFIESIG**IENDNWS 164
FR S + +II+RY +F+S +S* QPS *FS+ F G +SV*N + I * W
Sbjct: 13 FRVSPQ*EIIQRY*LHFHSPHS*G*QPSAP**FSSYFFGGKSV*NSRQIIS*SFACGYWF 192
Query: 165 GNIQV------KDMQILDGILIANEVVDEAKKTKKELLLFKVDFEKAYDSVD*GYLDAVM 218
N K DGILIANE VDEAKK KK+LLLFKVDFEKAY SVD YLD+V+
Sbjct: 193 CNFGCAVSVC*K*TNS*DGILIANEAVDEAKKLKKDLLLFKVDFEKAYHSVDWAYLDSVL 372
Query: 219 QKMAF 223
+ F
Sbjct: 373 GRYVF 387
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 135 bits (340), Expect(3) = 2e-62
Identities = 73/149 (48%), Positives = 95/149 (62%), Gaps = 6/149 (4%)
Frame = +2
Query: 596 ADMYRLGWEEGGDAWVWRRRLWAWEEECRS----LLDNVRLRLNVLDRWQWDPDTIDVYT 651
ADM RLGW G AW W RRL+ WEEEC LL+N L+ NV D+W+W D ++ Y+
Sbjct: 380 ADMGRLGWTVDGRAWEWTRRLFVWEEECVRECCILLNNFVLQDNVNDKWRWLLDPVNGYS 559
Query: 652 VRGVYQILTTS--VSNTIVKTDELVWHKQVPLN*VSIVAWRLLKDRLPTKTNLQRRDCLQ 709
V+ Y+ +T++ +S+ + D VWHK +P VS+ WRLL++RLPTK NL R L
Sbjct: 560 VKVFYRYITSTGHISDRSLVDD--VWHKHIPSK-VSLFVWRLLRNRLPTKDNLVHRGVLL 730
Query: 710 ATVNTCVSGCGIAESASHLFLHCEVFSSL 738
AT CV GC +ES +HLFLHC VF SL
Sbjct: 731 ATNAACVCGCVDSESTTHLFLHCNVFCSL 817
Score = 83.6 bits (205), Expect(3) = 2e-62
Identities = 33/72 (45%), Positives = 44/72 (60%)
Frame = +3
Query: 524 GGRSVSSWWKEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLGDVPFNRRFAR 583
GGR S WWK + K+++GVGE WF + +VGDG + FW+D W GDVP ++ R
Sbjct: 162 GGRQSSMWWKTICKVREGVGEGVGNWFEENIRMVVGDGRDAFFWYDTWAGDVPLRLKYPR 341
Query: 584 LFDLTTNKMCTV 595
L DL +K C V
Sbjct: 342 LLDLAMDKECKV 377
Score = 60.8 bits (146), Expect(3) = 2e-62
Identities = 27/41 (65%), Positives = 32/41 (77%)
Frame = +1
Query: 477 GLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEE 517
GLGV FNL+L+ KWCWRLL+D+ G W+RVL ARYGEE
Sbjct: 31 GLGVGA---FNLSLLGKWCWRLLVDKEGLWHRVLKARYGEE 144
>BE248682 similar to GP|18568269|gb putative gag-pol polyprotein {Zea mays},
partial (1%)
Length = 441
Score = 125 bits (313), Expect = 8e-29
Identities = 65/132 (49%), Positives = 91/132 (68%)
Frame = +3
Query: 250 DEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGYVVGASNSVVVSHLQFAD 309
+E S+ RGL+QGDPL+PFLF++ AEG++ LMK+ V NLF+G+ V + V SHLQ+AD
Sbjct: 6 EEISVQRGLKQGDPLAPFLFLLVAEGISGLMKNAVNRNLFQGFDVKRGGTRV-SHLQYAD 182
Query: 310 DTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLVGVNIGESWLLEAASVLNCKVG 369
DTL +G + N+ L+A+L F SGLKVNF+KS L+G+N+ ++ A LNC+
Sbjct: 183 DTLCIGMPTVDNLWTLKALLQGFEMASGLKVNFHKSSLIGINVPRDFMEAACRFLNCREE 362
Query: 370 KILFLYLGLSIG 381
I F+YLGL G
Sbjct: 363 SIPFIYLGLPGG 398
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 92.4 bits (228), Expect(2) = 9e-28
Identities = 70/224 (31%), Positives = 104/224 (46%), Gaps = 14/224 (6%)
Frame = -2
Query: 529 SSWWKEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLGDVPFNRRFARLFDLT 588
S WWK++ +++ VG WF ++ + VGDG ++ FW D W VP F R F
Sbjct: 778 SRWWKDLMSLEE-VGR--VRWFPRELIRKVGDGRSSFFWKDAWDSSVPLRESFPRAFFPY 608
Query: 589 TNKMCTVADMYRLGWEEGGDAW--VWRR-RLWAWEE----ECRSLLDNVRLRLNVLDRWQ 641
D++ + E G W WRR L+ WE+ E L+ V LR D W
Sbjct: 607 RLLKMGCGDLWDMNAE--GVRWRLYWRRLELFEWEKERLLELLGRLEGVVLRY-WADIWV 437
Query: 642 WDPDTIDVYTVRGVYQILTT--SVSNTIVKTDELV----WHKQVPLN*VSIVAWRLLKDR 695
W PD V++V Y +L + + + +E++ W + P V +W L DR
Sbjct: 436 WKPDKEGVFSVNSCYFLLQNLRLLEDRLSYEEEVIFRELWKSKAPAK-VLAFSWTLFLDR 260
Query: 696 LPTKTNLQRRDCLQATVNTCVSGCGIA-ESASHLFLHCEVFSSL 738
+PT NL +R L+ + CG E+ HLFLHC+V S +
Sbjct: 259 IPTMVNLGKRRLLRVEDSKRCVFCGCQDETVVHLFLHCDVISKV 128
Score = 50.1 bits (118), Expect(2) = 9e-28
Identities = 26/57 (45%), Positives = 35/57 (60%), Gaps = 2/57 (3%)
Frame = -3
Query: 470 CLKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGE--EARRLEVG 524
CL + GGLGVR +R N++L++KW WRLL D+ W VL YG +R + VG
Sbjct: 969 CLPRCKGGLGVRDIRLVNVSLLAKWWWRLLQDQSSLWKEVLEDIYGPRVSSRTMMVG 799
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 110 bits (275), Expect = 2e-24
Identities = 60/177 (33%), Positives = 97/177 (53%), Gaps = 3/177 (1%)
Frame = -3
Query: 174 ILDGILIANEVVDEAKKT---KKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKW 230
I D +LI ++++ +++ K + K D KAYD + +L V+ ++ F +W W
Sbjct: 643 ISDNVLITHKILHYLRQSGAKKHVSMAVKTDMTKAYDRIAWNFLREVLTRLGFHGIWISW 464
Query: 231 IKECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFK 290
I EC+ST + S L+NG P RGLRQGDPLSP+LF++ E L+ L + +
Sbjct: 463 IMECVSTVSYSFLINGGPQGRVLPSRGLRQGDPLSPYLFILCTEVLSGLCQQALRKGTLP 284
Query: 291 GYVVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLL 347
G V A N ++HL FADDT+ G+ + ++ L +++ + SG +N KS +
Sbjct: 283 GVKV-ARNCPPINHLLFADDTMFFGKSNASSCAILLSIMDKYRAASGRCIN*TKSAI 116
>CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [imported] -
Sulfolobus solfataricus, partial (3%)
Length = 789
Score = 86.3 bits (212), Expect = 4e-17
Identities = 40/67 (59%), Positives = 48/67 (70%)
Frame = -2
Query: 672 ELVWHKQVPLN*VSIVAWRLLKDRLPTKTNLQRRDCLQATVNTCVSGCGIAESASHLFLH 731
E++WH+QVPL VS+ AWRLL+DRLPTK+NL R + CVSGCG ESA HLFL
Sbjct: 590 EMIWHRQVPLK-VSVFAWRLLRDRLPTKSNLIYRGVIPTEAGLCVSGCGALESAQHLFLS 414
Query: 732 CEVFSSL 738
C F+SL
Sbjct: 413 CSYFASL 393
>AW686588
Length = 567
Score = 80.1 bits (196), Expect = 3e-15
Identities = 38/60 (63%), Positives = 45/60 (74%)
Frame = +1
Query: 679 VPLN*VSIVAWRLLKDRLPTKTNLQRRDCLQATVNTCVSGCGIAESASHLFLHCEVFSSL 738
VPL VSI+AWRL++DRLPTK NL RR CL CV GCGIAE+A+HLFLHC F ++
Sbjct: 142 VPLK-VSILAWRLIRDRLPTKANLVRRRCLAVEAAGCVVGCGIAETANHLFLHCATFGAV 318
>BQ148771
Length = 680
Score = 65.1 bits (157), Expect = 9e-11
Identities = 39/144 (27%), Positives = 66/144 (45%), Gaps = 2/144 (1%)
Frame = -3
Query: 397 IKTRLSGWQSCFLSFGGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSE 456
+ L+ W++ LS R+ L KSV+ ++P+Y + P I I+ L KF W +E
Sbjct: 585 VHVMLANWKANHLSLARRVTLAKSVIEAVPLYPMMTTIIPKACIEEIQKLQRKFVWGDTE 406
Query: 457 DNRKISWIAWSSVCLKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARY-- 514
+R+ + W ++ K GLG+R L N A + K W + V+ +Y
Sbjct: 405 VSRRYHAVGWETMSKPKTIYGLGLRRLDVMNKACIMKLGWSIYSGSNSLCTEVMRGKYQR 226
Query: 515 GEEARRLEVGGRSVSSWWKEVAKI 538
E + + + SS WK + K+
Sbjct: 225 SESLEEIFLEKPTDSSLWKALVKL 154
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 62.4 bits (150), Expect = 6e-10
Identities = 36/90 (40%), Positives = 49/90 (54%)
Frame = +1
Query: 256 RGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGYVVGASNSVVVSHLQFADDTLLLG 315
+GLRQGDPLSP+LF++ A L+ L+K G V S+ ++HL FADD+LL
Sbjct: 10 KGLRQGDPLSPYLFILCANVLSGLLKREGNKQNLHGIQVARSDP-KITHLLFADDSLLFA 186
Query: 316 EKSWANVRALRAVLTLFAEMSGLKVNFNKS 345
+ + VL + SG VNF KS
Sbjct: 187 RANLTEAATIMQVLHSYQSASGQLVNFEKS 276
>BF520135
Length = 202
Score = 61.2 bits (147), Expect = 1e-09
Identities = 30/62 (48%), Positives = 40/62 (64%)
Frame = +3
Query: 673 LVWHKQVPLN*VSIVAWRLLKDRLPTKTNLQRRDCLQATVNTCVSGCGIAESASHLFLHC 732
L+W K+ L+ VSI AWRL RLPTK N+ +R + + CV+ C + ES HLFLHC
Sbjct: 12 LLWRKE-DLSKVSIFAWRLFHGRLPTKANVFKRGIVHHDAHMCVTRCRLIESDVHLFLHC 188
Query: 733 EV 734
+V
Sbjct: 189 DV 194
>TC82520
Length = 833
Score = 58.5 bits (140), Expect = 9e-09
Identities = 34/88 (38%), Positives = 45/88 (50%)
Frame = +3
Query: 651 TVRGVYQILTTSVSNTIVKTDELVWHKQVPLN*VSIVAWRLLKDRLPTKTNLQRRDCLQA 710
T RG Y+ LT S + VW K +P VS+ WRL +RLPTK NL +R LQ
Sbjct: 117 TRRGTYRFLTISGQPLDRNQVDDVWQKNIPSK-VSMFVWRLFHNRLPTKVNLMQRHVLQQ 293
Query: 711 TVNTCVSGCGIAESASHLFLHCEVFSSL 738
T C+SG I H+ + +F +L
Sbjct: 294 THTACISGVAI-RKRQHICFYIVIFLAL 374
Score = 38.5 bits (88), Expect = 0.009
Identities = 14/21 (66%), Positives = 19/21 (89%)
Frame = +2
Query: 718 GCGIAESASHLFLHCEVFSSL 738
GCG +E+A+HLFLHC++F SL
Sbjct: 314 GCGDSETATHLFLHCDIFGSL 376
>AW774658 similar to GP|2808681|emb| Hcr9-4B {Lycopersicon hirsutum}, partial
(4%)
Length = 665
Score = 44.3 bits (103), Expect(2) = 1e-08
Identities = 20/28 (71%), Positives = 22/28 (78%)
Frame = -1
Query: 294 VGASNSVVVSHLQFADDTLLLGEKSWAN 321
+G + V SHLQFADDTLLLG KSWAN
Sbjct: 410 IGMHSLTVFSHLQFADDTLLLGVKSWAN 327
Score = 33.5 bits (75), Expect(2) = 1e-08
Identities = 14/28 (50%), Positives = 21/28 (75%)
Frame = -3
Query: 323 RALRAVLTLFAEMSGLKVNFNKSLLVGV 350
RALR++L +F MSGLKVN + +++ V
Sbjct: 327 RALRSILVIFENMSGLKVNLREEVIIYV 244
>BE941032
Length = 435
Score = 53.5 bits (127), Expect = 3e-07
Identities = 24/54 (44%), Positives = 31/54 (56%)
Frame = +2
Query: 685 SIVAWRLLKDRLPTKTNLQRRDCLQATVNTCVSGCGIAESASHLFLHCEVFSSL 738
SI W +L +R PTK NL +R + A +CV CG A+HLFLHC F +
Sbjct: 146 SIFLWCVLLNRFPTKDNLLKRGVISAIYQSCVGECGNLYDATHLFLHCNFFRQI 307
>BF650593
Length = 486
Score = 53.1 bits (126), Expect = 4e-07
Identities = 26/57 (45%), Positives = 31/57 (53%)
Frame = +3
Query: 676 HKQVPLN*VSIVAWRLLKDRLPTKTNLQRRDCLQATVNTCVSGCGIAESASHLFLHC 732
HK VPL VS + WRL ++ L T+ NL +R L CV CG ES SH F C
Sbjct: 297 HKSVPLK-VSCLVWRLFQNXLATRDNLSKRGVLDQNSIXCVXDCGREESVSHFFFEC 464
Score = 39.3 bits (90), Expect = 0.006
Identities = 19/53 (35%), Positives = 27/53 (50%)
Frame = +1
Query: 504 GFWYRVLVARYGEEARRLEVGGRSVSSWWKEVAKIKDGVGEDDEGWFVGKVSK 556
G W+R LV +YG + + R VS WWK++ I G E + V +V K
Sbjct: 85 GLWFRALVNKYGLNRGSITIENRGVSLWWKDICYIDFGGVESLNVYSVKEVYK 243
>BG456581
Length = 683
Score = 49.7 bits (117), Expect = 4e-06
Identities = 31/88 (35%), Positives = 44/88 (49%)
Frame = +2
Query: 651 TVRGVYQILTTSVSNTIVKTDELVWHKQVPLN*VSIVAWRLLKDRLPTKTNLQRRDCLQA 710
T++ VY LT+ S +W+ +P + S + WRL DRLPT NL R C A
Sbjct: 17 TIKLVYSFLTSHTS--CAPWASTIWNSCIPPS-HSFICWRLAHDRLPTDDNLSSRGC--A 181
Query: 711 TVNTCVSGCGIAESASHLFLHCEVFSSL 738
V+ C E++ HLFL C+ +L
Sbjct: 182 LVSMCSFCLEQVETSDHLFLRCKFVVTL 265
>BE999296
Length = 384
Score = 47.8 bits (112), Expect = 2e-05
Identities = 26/79 (32%), Positives = 39/79 (48%)
Frame = -1
Query: 652 VRGVYQILTTSVSNTIVKTDELVWHKQVPLN*VSIVAWRLLKDRLPTKTNLQRRDCLQAT 711
+ G YQ L ++ + + + VWH +PL V R+L++ LPTK N RR +
Sbjct: 306 IHGAYQFLMSADAPLDREYIDSVWHNHIPLK-VCFFVLRVLRNCLPTKDNFVRRRVIHEE 130
Query: 712 VNTCVSGCGIAESASHLFL 730
C +GC E+ LFL
Sbjct: 129 HMLCPTGCSFKETTDDLFL 73
>BG584442
Length = 775
Score = 46.6 bits (109), Expect = 3e-05
Identities = 25/88 (28%), Positives = 45/88 (50%), Gaps = 1/88 (1%)
Frame = +1
Query: 416 VLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRK-ISWIAWSSVCLKKE 474
V++K L S+ Y +S F + + IE + N F W +NRK + W++ + + K
Sbjct: 430 VMIKYALQSISSYVMSIFLLLNSQVDEIEKIMNTFSWVHVGENRKGMHWMS*EKLFVHKN 609
Query: 475 FGGLGVRLLREFNLALMSKWCWRLLIDR 502
+GG+G FN+ ++ K L++R
Sbjct: 610 YGGMGFTDFTTFNIPMLGKQV*SFLLNR 693
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 45.4 bits (106), Expect = 8e-05
Identities = 21/52 (40%), Positives = 30/52 (57%)
Frame = +2
Query: 234 CISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVE 285
C+ + +LVN D RGL+QGD LSP++F+I EGL+ L+ E
Sbjct: 104 CVESNDYYVLVNNDAVDPIIPSRGLQQGDHLSPYIFIICVEGLSFLIPHAKE 259
>BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgaris}, partial
(69%)
Length = 616
Score = 43.5 bits (101), Expect = 3e-04
Identities = 23/82 (28%), Positives = 44/82 (53%), Gaps = 1/82 (1%)
Frame = +3
Query: 169 VKDMQILDGILIANEVVDE-AKKTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLW 227
VK I D I++++E+V ++K + K+D KAYDS + ++ +M ++ FP +
Sbjct: 363 VKGRVIFDNIILSHELVKSYSRKGISPRCMVKIDLXKAYDSXEWPFIKHLMLELGFPYKF 542
Query: 228 RKWIKECISTTTASILVNGSPT 249
W+ ++T + + NG T
Sbjct: 543 VNWVMAXLTTASYTFNXNGDLT 608
>TC80043 similar to GP|9759561|dbj|BAB11163.1 MtN21 nodulin protein-like
{Arabidopsis thaliana}, partial (28%)
Length = 1421
Score = 41.2 bits (95), Expect(2) = 4e-04
Identities = 20/62 (32%), Positives = 34/62 (54%)
Frame = +1
Query: 546 DEGWFVGKVSKLVGDGSNTLFWFDRWLGDVPFNRRFARLFDLTTNKMCTVADMYRLGWEE 605
++ W G ++K+V +G +T FW ++W R+++RLF + +K V D L W E
Sbjct: 706 NKNWLSGSITKVVRNGRDTSFWSEKW----SRGRQYSRLFKILLDKEAKVVD---LRWRE 864
Query: 606 GG 607
G
Sbjct: 865 EG 870
Score = 20.8 bits (42), Expect(2) = 4e-04
Identities = 9/22 (40%), Positives = 14/22 (62%)
Frame = +3
Query: 519 RRLEVGGRSVSSWWKEVAKIKD 540
RR+ GGRS + WK++ + D
Sbjct: 639 RRIGKGGRSS*A*WKDLRREDD 704
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.335 0.146 0.498
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,622,742
Number of Sequences: 36976
Number of extensions: 443442
Number of successful extensions: 3686
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 2062
Number of HSP's successfully gapped in prelim test: 194
Number of HSP's that attempted gapping in prelim test: 1524
Number of HSP's gapped (non-prelim): 2390
length of query: 738
length of database: 9,014,727
effective HSP length: 103
effective length of query: 635
effective length of database: 5,206,199
effective search space: 3305936365
effective search space used: 3305936365
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC149205.5


