

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149205.3 + phase: 0 /pseudo
(523 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
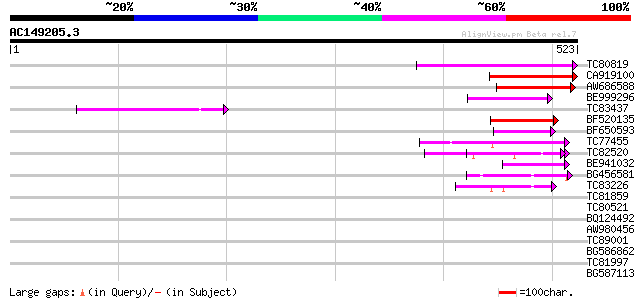
Score E
Sequences producing significant alignments: (bits) Value
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 141 7e-34
CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [i... 106 2e-23
AW686588 86 5e-17
BE999296 65 5e-11
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 64 1e-10
BF520135 62 5e-10
BF650593 57 1e-08
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 55 9e-08
TC82520 52 7e-07
BE941032 49 5e-06
BG456581 47 2e-05
TC83226 weakly similar to PIR|G86419|G86419 probable reverse tra... 44 1e-04
TC81859 similar to GP|18491135|gb|AAL69536.1 At1g04290/F19P19_27... 35 0.070
TC80521 weakly similar to GP|18377712|gb|AAL67006.1 unknown prot... 33 0.35
BQ124492 weakly similar to GP|9909168|dbj| putative transposable... 33 0.35
AW980456 32 0.45
TC89001 similar to GP|5726567|gb|AAD48471.1| glycine-rich RNA-bi... 32 0.45
BG586862 32 0.59
TC81997 31 1.0
BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At... 30 2.3
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 141 bits (355), Expect = 7e-34
Identities = 65/148 (43%), Positives = 90/148 (59%)
Frame = +2
Query: 376 AWQWRRHLWVWEEDLVEECGDLLLTVSLQELETDRWMWLSDQIRGYTVRGVYDMLTSQE* 435
AW+W R L+VWEE+ V EC LL LQ+ D+W WL D + GY+V+ Y +TS
Sbjct: 419 AWEWTRRLFVWEEECVRECCILLNNFVLQDNVNDKWRWLLDPVNGYSVKVFYRYITSTGH 598
Query: 436 PHVHQNMELIWHKQVPLKVSILALRLLRDRLPTKSNLADRGITFVEDRICVTGCGHVEDT 495
++ +WHK +P KVS+ RLLR+RLPTK NL RG+ + CV GC E T
Sbjct: 599 ISDRSLVDDVWHKHIPSKVSLFVWRLLRNRLPTKDNLVHRGVLLATNAACVCGCVDSEST 778
Query: 496 THMFLSCPIFGALLSMVQTWVGVEGVDS 523
TH+FL C +F +L S+V+ W+G+ + S
Sbjct: 779 THLFLHCNVFCSLWSLVRNWLGIPSMSS 862
>CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [imported] -
Sulfolobus solfataricus, partial (3%)
Length = 789
Score = 106 bits (264), Expect = 2e-23
Identities = 48/81 (59%), Positives = 61/81 (75%)
Frame = -2
Query: 443 ELIWHKQVPLKVSILALRLLRDRLPTKSNLADRGITFVEDRICVTGCGHVEDTTHMFLSC 502
E+IWH+QVPLKVS+ A RLLRDRLPTKSNL RG+ E +CV+GCG +E H+FLSC
Sbjct: 590 EMIWHRQVPLKVSVFAWRLLRDRLPTKSNLIYRGVIPTEAGLCVSGCGALESAQHLFLSC 411
Query: 503 PIFGALLSMVQTWVGVEGVDS 523
F +L S+V+ W+G GVD+
Sbjct: 410 SYFASLWSLVRDWIGFVGVDT 348
>AW686588
Length = 567
Score = 85.5 bits (210), Expect = 5e-17
Identities = 41/73 (56%), Positives = 48/73 (65%)
Frame = +1
Query: 450 VPLKVSILALRLLRDRLPTKSNLADRGITFVEDRICVTGCGHVEDTTHMFLSCPIFGALL 509
VPLKVSILA RL+RDRLPTK+NL R VE CV GCG E H+FL C FGA+
Sbjct: 142 VPLKVSILAWRLIRDRLPTKANLVRRRCLAVEAAGCVVGCGIAETANHLFLHCATFGAVW 321
Query: 510 SMVQTWVGVEGVD 522
++ W+GV G D
Sbjct: 322 QHIRAWIGVSGAD 360
>BE999296
Length = 384
Score = 65.5 bits (158), Expect = 5e-11
Identities = 30/78 (38%), Positives = 42/78 (53%)
Frame = -1
Query: 423 VRGVYDMLTSQE*PHVHQNMELIWHKQVPLKVSILALRLLRDRLPTKSNLADRGITFVED 482
+ G Y L S + P + ++ +WH +PLKV LR+LR+ LPTK N R + E
Sbjct: 306 IHGAYQFLMSADAPLDREYIDSVWHNHIPLKVCFFVLRVLRNCLPTKDNFVRRRVIHEEH 127
Query: 483 RICVTGCGHVEDTTHMFL 500
+C TGC E T +FL
Sbjct: 126 MLCPTGCSFKETTDDLFL 73
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 63.9 bits (154), Expect = 1e-10
Identities = 47/142 (33%), Positives = 73/142 (51%), Gaps = 1/142 (0%)
Frame = +1
Query: 62 WGR*SCSDVSFTIC**YAPDME*KLG*CTFVEGWSSFV*GYVWIESQFP*KLSRWGQY*C 121
WG *S ++ +IC + *KLG +G + + GYVW E +F *+ +Y
Sbjct: 520 WGG*SSCGITSSICKRHFVARN*KLGKHPCSKGCTCYFLGYVWFEGEFS*EWLGLCEYRS 699
Query: 122 LLVV*SGFCVGLQSG*SSVSLFGPSDWGRSTLVIILGAYC*SY*I*IVWLA*SVPFL*WS 181
LV * GFC L+SG ++ + + G+ + + LG YC S+ *+ WL *S F+ W
Sbjct: 700 FLVK*GGFCFKLESGQGTIFVLRYAYRGKFSALEFLGTYCESHKS*VDWLE*SF-FIFWR 876
Query: 182 PDST-QICLDCASCL*TFLFQS 202
P S+ ++ D CL F++
Sbjct: 877 PSSSVKVGSDLFVCLCASFFKA 942
>BF520135
Length = 202
Score = 62.0 bits (149), Expect = 5e-10
Identities = 29/63 (46%), Positives = 38/63 (60%)
Frame = +3
Query: 444 LIWHKQVPLKVSILALRLLRDRLPTKSNLADRGITFVEDRICVTGCGHVEDTTHMFLSCP 503
L+W K+ KVSI A RL RLPTK+N+ RGI + +CVT C +E H+FL C
Sbjct: 12 LLWRKEDLSKVSIFAWRLFHGRLPTKANVFKRGIVHHDAHMCVTRCRLIESDVHLFLHCD 191
Query: 504 IFG 506
+ G
Sbjct: 192 VLG 200
>BF650593
Length = 486
Score = 57.4 bits (137), Expect = 1e-08
Identities = 26/57 (45%), Positives = 32/57 (55%)
Frame = +3
Query: 447 HKQVPLKVSILALRLLRDRLPTKSNLADRGITFVEDRICVTGCGHVEDTTHMFLSCP 503
HK VPLKVS L RL ++ L T+ NL+ RG+ CV CG E +H F CP
Sbjct: 297 HKSVPLKVSCLVWRLFQNXLATRDNLSKRGVLDQNSIXCVXDCGREESVSHFFFECP 467
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 54.7 bits (130), Expect = 9e-08
Identities = 42/146 (28%), Positives = 65/146 (43%), Gaps = 8/146 (5%)
Frame = -2
Query: 379 WRR-HLWVWEEDLVEECGDLLLTVSLQELETDRWMWLSDQIRGYTVRGVYDMLTSQE*PH 437
WRR L+ WE++ + E L V L+ D W+W D+ ++V Y +L +
Sbjct: 538 WRRLELFEWEKERLLELLGRLEGVVLRYW-ADIWVWKPDKEGVFSVNSCYFLLQNLRLLE 362
Query: 438 VHQNMEL------IWHKQVPLKVSILALRLLRDRLPTKSNLADRGITFVEDRICVTGCGH 491
+ E +W + P KV + L DR+PT NL R + VED CG
Sbjct: 361 DRLSYEEEVIFRELWKSKAPAKVLAFSWTLFLDRIPTMVNLGKRRLLRVEDSKRCVFCGC 182
Query: 492 VEDT-THMFLSCPIFGALLSMVQTWV 516
++T H+FL C + + V W+
Sbjct: 181 QDETVVHLFLHCDVISKV*REVMRWL 104
>TC82520
Length = 833
Score = 51.6 bits (122), Expect = 7e-07
Identities = 30/91 (32%), Positives = 43/91 (46%)
Frame = +3
Query: 422 TVRGVYDMLTSQE*PHVHQNMELIWHKQVPLKVSILALRLLRDRLPTKSNLADRGITFVE 481
T RG Y LT P ++ +W K +P KVS+ RL +RLPTK NL R +
Sbjct: 117 TRRGTYRFLTISGQPLDRNQVDDVWQKNIPSKVSMFVWRLFHNRLPTKVNLMQRHVLQQT 296
Query: 482 DRICVTGCGHVEDTTHMFLSCPIFGALLSMV 512
C++G + H+ IF AL ++
Sbjct: 297 HTACISGVA-IRKRQHICFYIVIFLALFGLM 386
Score = 47.8 bits (112), Expect = 1e-05
Identities = 45/141 (31%), Positives = 64/141 (44%), Gaps = 7/141 (4%)
Frame = +2
Query: 383 LWVWEEDLVEECGDLLLTVSLQELETDRWMWLSDQIRGYTVRGV---YDMLTSQE*PHVH 439
L+ WEE+ V E LL LQE D WL D I GYT + + + T+ *
Sbjct: 2 LFAWEEESVREWYALLHNTVLQENVHDVCRWLLDPI*GYTEGNISLSHYLWTTIG*ESS* 181
Query: 440 QNMELIWH-KQVPLKVSILALRLLRD---RLPTKSNLADRGITFVEDRICVTGCGHVEDT 495
+ + + K V ++V+ L+ + T S+ G+ F GCG E
Sbjct: 182 RCLAKEYSFKGVYVRVASLSQ*TSHEG*FNAATCSSTNSHGLHF--------GCGDSETA 337
Query: 496 THMFLSCPIFGALLSMVQTWV 516
TH+FL C IFG+L S V W+
Sbjct: 338 THLFLHCDIFGSLWSHVLRWL 400
>BE941032
Length = 435
Score = 48.9 bits (115), Expect = 5e-06
Identities = 24/62 (38%), Positives = 33/62 (52%)
Frame = +2
Query: 455 SILALRLLRDRLPTKSNLADRGITFVEDRICVTGCGHVEDTTHMFLSCPIFGALLSMVQT 514
SI +L +R PTK NL RG+ + CV CG++ D TH+FL C F + V
Sbjct: 146 SIFLWCVLLNRFPTKDNLLKRGVISAIYQSCVGECGNLYDATHLFLHCNFFRQIWINVSD 325
Query: 515 WV 516
W+
Sbjct: 326 WL 331
>BG456581
Length = 683
Score = 47.0 bits (110), Expect = 2e-05
Identities = 34/100 (34%), Positives = 46/100 (46%), Gaps = 2/100 (2%)
Frame = +2
Query: 422 TVRGVYDMLTSQE*PHVHQNMELIWHKQVPLKVSILALRLLRDRLPTKSNLADRGITFVE 481
T++ VY LTS IW+ +P S + RL DRLPT NL+ RG V
Sbjct: 17 TIKLVYSFLTSHT--SCAPWASTIWNSCIPPSHSFICWRLAHDRLPTDDNLSSRGCALVS 190
Query: 482 DRICVTGCGHVEDTTHMFLSCPIFGALLSMV--QTWVGVE 519
+C VE + H+FL C L S + Q VG++
Sbjct: 191 --MCSFCLEQVETSDHLFLRCKFVVTLWSWLCSQLRVGLD 304
>TC83226 weakly similar to PIR|G86419|G86419 probable reverse transcriptase
100033-105622 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 885
Score = 44.3 bits (103), Expect = 1e-04
Identities = 30/102 (29%), Positives = 47/102 (45%), Gaps = 9/102 (8%)
Frame = +3
Query: 412 MWLSDQIRGYTVRGVYDMLTSQE*PHVHQNME-----LIWHKQVPLKV----SILALRLL 462
MW+ + Y+V+ Y+ L + + ++ LIW K L +L R+L
Sbjct: 3 MWMHNPTGIYSVKSGYNTLRTWQTQQINNTSTSSDETLIWKKIWSLHTIPRHKVLLWRIL 182
Query: 463 RDRLPTKSNLADRGITFVEDRICVTGCGHVEDTTHMFLSCPI 504
D LP +S+L RGI +C E TH+F+SCP+
Sbjct: 183 NDSLPVRSSLRKRGIQCYP--LCPRCHSKTETITHLFMSCPL 302
>TC81859 similar to GP|18491135|gb|AAL69536.1 At1g04290/F19P19_27
{Arabidopsis thaliana}, partial (65%)
Length = 960
Score = 35.0 bits (79), Expect = 0.070
Identities = 17/42 (40%), Positives = 23/42 (54%)
Frame = +2
Query: 464 DRLPTKSNLADRGITFVEDRICVTGCGHVEDTTHMFLSCPIF 505
DRLPTK N+ R I ++ CV G E + +F CP+F
Sbjct: 617 DRLPTKYNVLPREINNLDSSFCVGGWEVNETSQSLFF*CPLF 742
>TC80521 weakly similar to GP|18377712|gb|AAL67006.1 unknown protein
{Arabidopsis thaliana}, partial (57%)
Length = 1025
Score = 32.7 bits (73), Expect = 0.35
Identities = 10/21 (47%), Positives = 15/21 (70%)
Frame = +3
Query: 366 VLVGGGGWWGAWQWRRHLWVW 386
+L+ GG W G +WR+ +WVW
Sbjct: 915 ILIRGGWWKGLRKWRKGMWVW 977
>BQ124492 weakly similar to GP|9909168|dbj| putative transposable element
Tip100 protein {Oryza sativa (japonica cultivar-group)},
partial (8%)
Length = 694
Score = 32.7 bits (73), Expect = 0.35
Identities = 12/30 (40%), Positives = 18/30 (60%)
Frame = +2
Query: 492 VEDTTHMFLSCPIFGALLSMVQTWVGVEGV 521
++ H+FL C IF +L S V W+G+ V
Sbjct: 527 IKTARHLFLDCDIFSSLWSQVWLWLGISSV 616
>AW980456
Length = 779
Score = 32.3 bits (72), Expect = 0.45
Identities = 21/68 (30%), Positives = 33/68 (47%), Gaps = 4/68 (5%)
Frame = -2
Query: 409 DRWMWLSDQIRGYTVRGVYDMLTSQ--E*PHVHQ--NMELIWHKQVPLKVSILALRLLRD 464
DR +W ++ Y V+ Y + + ++H+ N IW +VP KV L R+ R
Sbjct: 208 DRLIWKDEKHGKYYVKSAYRFCVEELFDSSYLHRPGNWSGIWKLKVPPKVQNLVWRMCRG 29
Query: 465 RLPTKSNL 472
LPT+ L
Sbjct: 28 CLPTRIRL 5
>TC89001 similar to GP|5726567|gb|AAD48471.1| glycine-rich RNA-binding
protein {Glycine max}, partial (90%)
Length = 488
Score = 32.3 bits (72), Expect = 0.45
Identities = 8/14 (57%), Positives = 9/14 (64%)
Frame = +2
Query: 373 WWGAWQWRRHLWVW 386
WW W WRR W+W
Sbjct: 332 WWRRWLWRRRWWIW 373
>BG586862
Length = 804
Score = 32.0 bits (71), Expect = 0.59
Identities = 22/63 (34%), Positives = 30/63 (46%), Gaps = 1/63 (1%)
Frame = -1
Query: 443 ELIWH-KQVPLKVSILALRLLRDRLPTKSNLADRGITFVEDRICVTGCGHVEDTTHMFLS 501
E +W K +P S L RLL + LP K L RGI +C +E H+FL+
Sbjct: 660 EKVWGIKTIPRHKSFL-WRLLHNALPVKDELHKRGIRC--SLLCPRCESKIETVQHLFLN 490
Query: 502 CPI 504
C +
Sbjct: 489 CEV 481
>TC81997
Length = 652
Score = 31.2 bits (69), Expect = 1.0
Identities = 23/81 (28%), Positives = 37/81 (45%), Gaps = 12/81 (14%)
Frame = +2
Query: 409 DRWMWLSDQIRGYTVRGVYDMLTSQE*PHVHQNMELIWHKQVPL------------KVSI 456
D+ MW D + GYTV+ Y L HQN+ + + +V + K +
Sbjct: 164 DK*MWKYDMLCGYTVKTGYSTL--------HQNVYSLEYDEVGMVIT*VLCSAVLSKEMV 319
Query: 457 LALRLLRDRLPTKSNLADRGI 477
AL++L D+L T+ RG+
Sbjct: 320 FALKMLLDKLFTRDARG*RGL 382
>BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (10%)
Length = 767
Score = 30.0 bits (66), Expect = 2.3
Identities = 25/106 (23%), Positives = 44/106 (40%), Gaps = 11/106 (10%)
Frame = -3
Query: 409 DRWMWLSDQIRGYTVRGVYDMLTS----------QE*PHVHQNMELIWHKQVPLKVSILA 458
D + W + Y+V+ Y + T+ + P + + +W KV
Sbjct: 426 DSYSWEYSKSGHYSVKSGYYVQTNIIAAANQRGTVDQPSLDDLYQRVWKYNTSPKVRHFL 247
Query: 459 LRLLRDRLPTKSNLADRGITFVEDRICVTGCGHVEDT-THMFLSCP 503
R + + LPT +N+ R I+ +D C + CG +T H+ CP
Sbjct: 246 WRCISNSLPTAANMRSRHIS--KDGSC-SRCGMESETVNHILFQCP 118
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.364 0.164 0.688
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,852,137
Number of Sequences: 36976
Number of extensions: 342145
Number of successful extensions: 5600
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 2293
Number of HSP's successfully gapped in prelim test: 211
Number of HSP's that attempted gapping in prelim test: 2783
Number of HSP's gapped (non-prelim): 3076
length of query: 523
length of database: 9,014,727
effective HSP length: 101
effective length of query: 422
effective length of database: 5,280,151
effective search space: 2228223722
effective search space used: 2228223722
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.5 bits)
S2: 61 (28.1 bits)
Medicago: description of AC149205.3


