

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149205.17 + phase: 0 /pseudo
(626 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
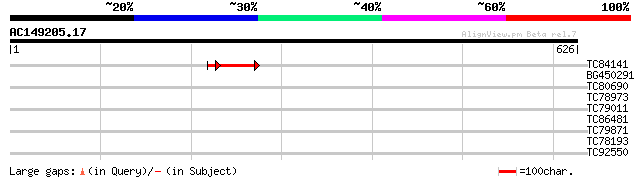
TC84141 weakly similar to PIR|H96544|H96544 hypothetical protein... 57 4e-10
BG450291 similar to GP|14335084|gb| At1g11480/T23J18_15 {Arabido... 32 0.56
TC80690 weakly similar to PIR|F84587|F84587 hypothetical protein... 31 1.2
TC78973 similar to PIR|D84745|D84745 plastid protein [imported] ... 30 2.8
TC79011 weakly similar to GP|9758298|dbj|BAB08841.1 dihydroneopt... 30 2.8
TC86481 similar to GP|9294170|dbj|BAB02072.1 emb|CAB71043.1~gene... 29 6.2
TC79871 similar to GP|19310486|gb|AAL84977.1 AT5g13970/MAC12_6 {... 29 6.2
TC78193 similar to PIR|T05166|T05166 quinone reductase homolog F... 28 8.1
TC92550 weakly similar to GP|12834399|dbj|BAB22895. coatomer pro... 28 8.1
>TC84141 weakly similar to PIR|H96544|H96544 hypothetical protein F8A12.4
[imported] - Arabidopsis thaliana, partial (3%)
Length = 720
Score = 57.4 bits (137), Expect(2) = 4e-10
Identities = 34/47 (72%), Positives = 35/47 (74%)
Frame = +2
Query: 229 KSSVSYCCSSC*RDSYGFGTCCFG*LI*GFEFV*ETNC*FEKMS**I 275
K VSYC S *R Y FGT CFG*LI*G V*ETNC*FEK+S**I
Sbjct: 50 KFFVSYCYSFG*RQYYCFGTNCFG*LI*GLNSV*ETNC*FEKIS**I 190
Score = 25.0 bits (53), Expect(2) = 4e-10
Identities = 10/14 (71%), Positives = 12/14 (85%)
Frame = +3
Query: 219 FVFPHRYNVVKSSV 232
FVF HRY +VKSS+
Sbjct: 18 FVFSHRYFIVKSSL 59
>BG450291 similar to GP|14335084|gb| At1g11480/T23J18_15 {Arabidopsis
thaliana}, partial (12%)
Length = 680
Score = 32.3 bits (72), Expect = 0.56
Identities = 26/94 (27%), Positives = 38/94 (39%)
Frame = +2
Query: 473 RSASSSKCEPSAKIPPEFPSHKLVGCTVTIGKSSDDGSNTSQGDKIVDDDAPSGSIPKLL 532
++ S P A P E P KL+ T + S +QG + V+D P ++P+
Sbjct: 125 KAVGSESQHPVASEPIERPKLKLLPRTKPLESSEPSVIEPTQGYRQVNDSVPVETVPQRH 304
Query: 533 KTMSFEKSVEDGLLAEKYVDGGAPLPAKDNTPTP 566
+F KSV G + K L K P P
Sbjct: 305 GHANFVKSVSVGTESPKDPGQRPKLNLKPLKPRP 406
>TC80690 weakly similar to PIR|F84587|F84587 hypothetical protein At2g20310
[imported] - Arabidopsis thaliana, partial (21%)
Length = 1269
Score = 31.2 bits (69), Expect = 1.2
Identities = 23/70 (32%), Positives = 33/70 (46%)
Frame = -3
Query: 469 EPRKRSASSSKCEPSAKIPPEFPSHKLVGCTVTIGKSSDDGSNTSQGDKIVDDDAPSGSI 528
+P S+SS++ P + P + T SS S TS DK+ D D PSG++
Sbjct: 592 DPSVESSSSTEVTPFSGSPSK-----------TTQSSSSTSSFTSFTDKVEDFDFPSGAL 446
Query: 529 PKLLKTMSFE 538
+KT S E
Sbjct: 445 FS*MKTTSSE 416
>TC78973 similar to PIR|D84745|D84745 plastid protein [imported] -
Arabidopsis thaliana, partial (69%)
Length = 1583
Score = 30.0 bits (66), Expect = 2.8
Identities = 11/18 (61%), Positives = 11/18 (61%)
Frame = +2
Query: 133 CVSIW*SNNHFGRCHCFG 150
CV IW HF CHCFG
Sbjct: 311 CVLIWFFFPHFNSCHCFG 364
>TC79011 weakly similar to GP|9758298|dbj|BAB08841.1 dihydroneopterin
aldolase-like protein {Arabidopsis thaliana}, partial
(86%)
Length = 628
Score = 30.0 bits (66), Expect = 2.8
Identities = 16/45 (35%), Positives = 27/45 (59%), Gaps = 1/45 (2%)
Frame = +2
Query: 327 LTMWGWH*IQLSMILFAVLMFNMLISVEYIIQMMKLW-YHLKKIW 370
+ M GW QL ++ +++ LI+++YI+ + KLW HLK W
Sbjct: 230 MLMLGWISNQLENLITCLIL---LITLKYIV*LRKLWKEHLKTFW 355
>TC86481 similar to GP|9294170|dbj|BAB02072.1
emb|CAB71043.1~gene_id:MJL12.8~similar to unknown
protein {Arabidopsis thaliana}, partial (86%)
Length = 2040
Score = 28.9 bits (63), Expect = 6.2
Identities = 15/38 (39%), Positives = 19/38 (49%)
Frame = +3
Query: 527 SIPKLLKTMSFEKSVEDGLLAEKYVDGGAPLPAKDNTP 564
SIPK L +S E EKY+D G P+P + P
Sbjct: 756 SIPKSLHCLSMRLMEERIAHPEKYIDEGKPIPPEVEDP 869
>TC79871 similar to GP|19310486|gb|AAL84977.1 AT5g13970/MAC12_6 {Arabidopsis
thaliana}, partial (6%)
Length = 1596
Score = 28.9 bits (63), Expect = 6.2
Identities = 27/105 (25%), Positives = 41/105 (38%)
Frame = +2
Query: 509 GSNTSQGDKIVDDDAPSGSIPKLLKTMSFEKSVEDGLLAEKYVDGGAPLPAKDNTPTPLI 568
GSN SQ D+ +DD IPK K D + E ++ G K P +
Sbjct: 998 GSNASQADEALDDLPSVTFIPK--------KKSGDVTMGENKLEVGKEGMNKKAFPVSIA 1153
Query: 569 FVDDCKHVLEDGNESEEARLSTDTICQSENQTESYSYLSELIVEE 613
DD ++ E +E + DT S+ Y ++ +EE
Sbjct: 1154 ADDDTENSDVCAMEEDEHEVIEDTKKSSQKSNRKYRKKTDDDLEE 1288
>TC78193 similar to PIR|T05166|T05166 quinone reductase homolog F18E5.200 -
Arabidopsis thaliana, partial (95%)
Length = 1379
Score = 28.5 bits (62), Expect = 8.1
Identities = 17/44 (38%), Positives = 23/44 (51%), Gaps = 1/44 (2%)
Frame = +3
Query: 153 LSLWSSCFSPNLKIRRLE-RWNRSSSALKAKR*FQKVWPLPHQC 195
LSLWS+ SP LK+R+L W S + +K W + QC
Sbjct: 891 LSLWSTNLSPFLKLRKLTGLWKAVSILERYCSWLEKPW*ILDQC 1022
>TC92550 weakly similar to GP|12834399|dbj|BAB22895. coatomer protein
complex subunit zeta 2~data source:MGD source
key:MGI:1929008 evidence:ISS~, partial (36%)
Length = 697
Score = 28.5 bits (62), Expect = 8.1
Identities = 14/40 (35%), Positives = 26/40 (65%)
Frame = -2
Query: 2 MNEQPLKDSIMEMREDFMISPDGHSKQTFRTAHFS*TYCK 41
+N +K++++ +++DF ISP G KQ+F +F YC+
Sbjct: 246 VNAITIKNNLVIIKDDFAISPSGLFKQSF-FENFLFFYCR 130
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.350 0.153 0.557
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,101,528
Number of Sequences: 36976
Number of extensions: 392444
Number of successful extensions: 3519
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 1933
Number of HSP's successfully gapped in prelim test: 192
Number of HSP's that attempted gapping in prelim test: 1544
Number of HSP's gapped (non-prelim): 2207
length of query: 626
length of database: 9,014,727
effective HSP length: 102
effective length of query: 524
effective length of database: 5,243,175
effective search space: 2747423700
effective search space used: 2747423700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC149205.17


