

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149205.14 - phase: 0 /pseudo
(890 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
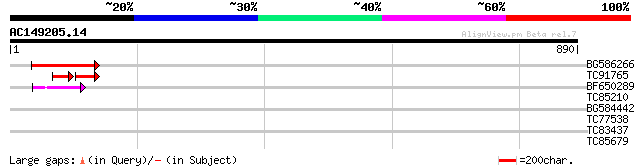
Score E
Sequences producing significant alignments: (bits) Value
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 90 3e-18
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 53 4e-12
BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgari... 51 2e-06
TC85210 similar to PIR|S56655|S56655 lipoxygenase (EC 1.13.11.12... 40 0.003
BG584442 36 0.057
TC77538 similar to SP|P54237|G6P1_CLAMI Glucose-6-phosphate isom... 30 4.1
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 28 5.9
TC85679 homologue to GP|15919089|emb|CAC86996. ATP citrate lyase... 29 9.1
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 90.1 bits (222), Expect = 3e-18
Identities = 38/106 (35%), Positives = 68/106 (63%)
Frame = -3
Query: 35 PDRSILNNAMVAIEVIHFMKTKTRGQDKYVALKLDISKAYDRMDWDYLKEIMIKMGFSQK 94
P R+I +N ++ +++H+++ + +A+K D++KAYDR+ W++L+E++ ++GF
Sbjct: 655 PGRAISDNVLITHKILHYLRQSGAKKHVSMAVKTDMTKAYDRIAWNFLREVLTRLGFHGI 476
Query: 95 WIQWMVVSTESVDYNVLVNAEHVGPVILGRGLRQGGPLSLSFYHLC 140
WI W++ +V Y+ L+N G V+ RGLRQG PLS + LC
Sbjct: 475 WISWIMECVSTVSYSFLINGGPQGRVLPSRGLRQGDPLSPYLFILC 338
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 52.8 bits (125), Expect(2) = 4e-12
Identities = 23/32 (71%), Positives = 27/32 (83%)
Frame = +1
Query: 68 LDISKAYDRMDWDYLKEIMIKMGFSQKWIQWM 99
L ISK Y+R+D DYLKEIMIKMGF+ +WI WM
Sbjct: 1 LYISKVYNRVD*DYLKEIMIKMGFNNRWIYWM 96
Score = 37.4 bits (85), Expect(2) = 4e-12
Identities = 19/37 (51%), Positives = 24/37 (64%)
Frame = +2
Query: 104 ESVDYNVLVNAEHVGPVILGRGLRQGGPLSLSFYHLC 140
ES DY VLVN + V P+I RGL+QG LS + +C
Sbjct: 110 ESNDYYVLVNNDAVDPIIPSRGLQQGDHLSPYIFIIC 220
>BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgaris}, partial
(69%)
Length = 616
Score = 51.2 bits (121), Expect = 2e-06
Identities = 22/83 (26%), Positives = 44/83 (52%)
Frame = +3
Query: 37 RSILNNAMVAIEVIHFMKTKTRGQDKYVALKLDISKAYDRMDWDYLKEIMIKMGFSQKWI 96
R I +N +++ E++ K G +K+D+ KAYD +W ++K +M+++GF K++
Sbjct: 372 RVIFDNIILSHELVKSYSRK--GISPRCMVKIDLXKAYDSXEWPFIKHLMLELGFPYKFV 545
Query: 97 QWMVVSTESVDYNVLVNAEHVGP 119
W++ + Y N + P
Sbjct: 546 NWVMAXLTTASYTFNXNGDLTXP 614
>TC85210 similar to PIR|S56655|S56655 lipoxygenase (EC 1.13.11.12) loxG -
garden pea, partial (18%)
Length = 863
Score = 40.4 bits (93), Expect = 0.003
Identities = 26/52 (50%), Positives = 34/52 (65%), Gaps = 1/52 (1%)
Frame = +2
Query: 248 HLLKHPY*L*LVE*LSWMKFNK*GYIN-EKNNKCCIGILKVTIILRKRKNCK 298
HLLKH Y *LV+*L+ +KFNK YI +K +K CI ILK + + K C+
Sbjct: 686 HLLKHLYQW*LVK*LNEVKFNKKEYIKVKKIDKYCINILK*KLF*DR*KKCQ 841
>BG584442
Length = 775
Score = 36.2 bits (82), Expect = 0.057
Identities = 31/99 (31%), Positives = 47/99 (47%)
Frame = +3
Query: 359 KN*FLEQQVSI*GGA*GHDQVGSSSYSFLCHEHFPTT*HFNYDY*TND*LVSVGSWSHLS 418
KN*FLE+Q+ + G+D V + + FLC E+F + + * + +GS S
Sbjct: 381 KN*FLEKQMFVKSYVRGND*VCTPKHIFLCDEYFSPSKFSSR*N*KDYEYFFMGSCWRKS 560
Query: 419 SWN*LAELGKAFYV*SSWWYGL*RLIGFQFSYVGQAGME 457
N L LG+ * W +G RL Q+S G++
Sbjct: 561 QRNALDVLGEIVCT*KLWRHGFYRLHDVQYSNAW*TGLK 677
>TC77538 similar to SP|P54237|G6P1_CLAMI Glucose-6-phosphate isomerase
cytosolic 1 (EC 5.3.1.9) (GPI) (Phosphoglucose isomerase)
(PGI), complete
Length = 2215
Score = 30.0 bits (66), Expect = 4.1
Identities = 17/40 (42%), Positives = 21/40 (52%), Gaps = 6/40 (15%)
Frame = +1
Query: 683 VWHSSLEC------GWFMV*CSTCCYSQ*YSYCCYFLSS* 716
VWH S C GW+ +*C CC+S + FLSS*
Sbjct: 967 VWH*S**CFCILGLGWWSI*CLQCCWSVSFIPAIRFLSS* 1086
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 28.1 bits (61), Expect(2) = 5.9
Identities = 11/26 (42%), Positives = 16/26 (61%)
Frame = +1
Query: 67 KLDISKAYDRMDWDYLKEIMIKMGFS 92
K+D KAY +DW YL ++ + FS
Sbjct: 313 KVDFEKAYHSVDWAYLDSVLGRYVFS 390
Score = 19.6 bits (39), Expect(2) = 5.9
Identities = 8/27 (29%), Positives = 13/27 (47%)
Frame = +2
Query: 87 IKMGFSQKWIQWMVVSTESVDYNVLVN 113
+ M F W +W+ + +VLVN
Sbjct: 374 VGMSFLVLWRKWIKECVSTATTSVLVN 454
>TC85679 homologue to GP|15919089|emb|CAC86996. ATP citrate lyase b-subunit
{Lupinus albus}, complete
Length = 1732
Score = 28.9 bits (63), Expect = 9.1
Identities = 14/29 (48%), Positives = 20/29 (68%)
Frame = -2
Query: 280 CCIGILKVTIILRKRKNCKETIIVRRRE* 308
C + I+K + ++KN KETI+VRR E*
Sbjct: 1710 CYMSIVK*QLF*EEKKNAKETILVRRIE* 1624
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.368 0.164 0.641
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,794,539
Number of Sequences: 36976
Number of extensions: 510398
Number of successful extensions: 6014
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1248
Number of HSP's successfully gapped in prelim test: 171
Number of HSP's that attempted gapping in prelim test: 4696
Number of HSP's gapped (non-prelim): 1578
length of query: 890
length of database: 9,014,727
effective HSP length: 105
effective length of query: 785
effective length of database: 5,132,247
effective search space: 4028813895
effective search space used: 4028813895
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC149205.14


