

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149204.7 + phase: 0
(600 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
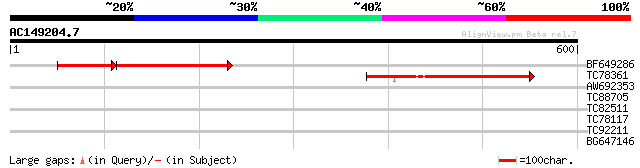
Score E
Sequences producing significant alignments: (bits) Value
BF649286 235 3e-94
TC78361 weakly similar to PIR|H86392|H86392 hypothetical protein... 128 7e-30
AW692353 35 0.11
TC88705 similar to GP|10177834|dbj|BAB11263. gene_id:MCD7.9~unkn... 35 0.11
TC82511 similar to PIR|A96606|A96606 unknown protein F13N6.17 [i... 35 0.11
TC78117 similar to GP|10177445|dbj|BAB10741. protein kinase {Ara... 31 1.5
TC92211 similar to PIR|B86182|B86182 hypothetical protein [impor... 30 2.6
BG647146 similar to GP|15146198|gb| F17F16.8/F17F16.8 {Arabidops... 28 7.7
>BF649286
Length = 562
Score = 235 bits (600), Expect(2) = 3e-94
Identities = 117/122 (95%), Positives = 117/122 (95%)
Frame = +2
Query: 114 GCRKKLCDTNRVKVVNPELNASELEKVYEYIDGRKSCGTNKMKDTNLVSNMAKKVESEGL 173
GCRKKLCDTNRVKVVNPELNASELEKVYEYIDGRKSCGTNKMKDTNLVSNMAKKVESEGL
Sbjct: 191 GCRKKLCDTNRVKVVNPELNASELEKVYEYIDGRKSCGTNKMKDTNLVSNMAKKVESEGL 370
Query: 174 VDVLMQDCKTNETSLRKLVNQNAAMEIMDDVSDVANSMPGLSDGSKSCANDDANDSGDDV 233
VDVLMQDCKTNETSL KLVNQNAAMEIMDDVSDV NSMPGLS GSKSCANDD DSGDDV
Sbjct: 371 VDVLMQDCKTNETSLXKLVNQNAAMEIMDDVSDVXNSMPGLSXGSKSCANDDPXDSGDDV 550
Query: 234 LM 235
LM
Sbjct: 551 LM 556
Score = 128 bits (322), Expect(2) = 3e-94
Identities = 63/63 (100%), Positives = 63/63 (100%)
Frame = +1
Query: 51 LVYSKYLSGLETPLKNVVDGEFPKRDLVGDRVKFGERLTELQAELVLDDYGEGDVGDEVK 110
LVYSKYLSGLETPLKNVVDGEFPKRDLVGDRVKFGERLTELQAELVLDDYGEGDVGDEVK
Sbjct: 1 LVYSKYLSGLETPLKNVVDGEFPKRDLVGDRVKFGERLTELQAELVLDDYGEGDVGDEVK 180
Query: 111 SVY 113
SVY
Sbjct: 181 SVY 189
>TC78361 weakly similar to PIR|H86392|H86392 hypothetical protein AAF98560.1
[imported] - Arabidopsis thaliana, partial (28%)
Length = 1374
Score = 128 bits (321), Expect = 7e-30
Identities = 72/189 (38%), Positives = 115/189 (60%), Gaps = 11/189 (5%)
Frame = +1
Query: 378 ILDAPPTVNIPLGPNHQAEVPEW--TGTTH--------KSDSKWLGTQIWPLQIVKSKLL 427
+L+ P + +G +HQA+VP W +G+ +++ + +GT + P+ ++ L
Sbjct: 712 LLEHSPRKPVSVGADHQADVPAWGFSGSDSNLNVRDGDEAEKRLMGTCVIPMPEMELSSL 891
Query: 428 EGEPVGKGRQDSCRCQVQGSVECVRFHIAEKSAKLKLEIGVA-FYQWNLDKAGEEVRRCW 486
+ E VGKGR D C C+ +GS+ CVR HI E+ KL G+ F + GE+V + W
Sbjct: 892 DDE-VGKGRTD-CSCEDRGSMRCVRQHIMEERKKLLKTFGIEKFIELGFAGMGEQVAQKW 1065
Query: 487 TAEEEKKFKDVVKSNPASLDRCFWDDIFTTFPKKSRESLVSYYFNVFLLQRRGYQNRHTP 546
T E+E F VV +NPASL++ FW+ + FP +S++ +VSYYFNVF+L++R QNR+
Sbjct: 1066TQEDEHLFHKVVFNNPASLNKNFWNYLSIVFPSRSKKEIVSYYFNVFMLRKRAEQNRNHL 1245
Query: 547 DNIDSDDEE 555
+ DSD++E
Sbjct: 1246LSADSDNDE 1272
>AW692353
Length = 584
Score = 34.7 bits (78), Expect = 0.11
Identities = 16/33 (48%), Positives = 21/33 (63%)
Frame = +1
Query: 13 LFMVVKDKGGYDVVCKNELWDLVGEEYGLGVKV 45
LF +VK+KGGY V + LWD V + GL + V
Sbjct: 367 LFSLVKEKGGYAAVSRKGLWDSVIVDLGLDLNV 465
>TC88705 similar to GP|10177834|dbj|BAB11263. gene_id:MCD7.9~unknown protein
{Arabidopsis thaliana}, partial (32%)
Length = 1685
Score = 34.7 bits (78), Expect = 0.11
Identities = 35/147 (23%), Positives = 62/147 (41%), Gaps = 6/147 (4%)
Frame = +1
Query: 183 TNETSLRKLVNQNAAMEIMDDVSDVANSMPGLSDGSKSCANDDANDSGDDVLMLDPSS-- 240
T E +K V N A++ D + + ++G S +DD + DD D +
Sbjct: 523 TGEDVGKKSVEANTALDNEDQEDILHGTNNEENEGYASETDDDTSKYDDDTGKYDDDTGK 702
Query: 241 ----VNRESFGRKRKRDDLMSEMQSWVIRTAKNPCDPVLGSMPEKSKWKSYGNQEIWKKV 296
+N E F ++ + +DL S +S V DP +KS W I++
Sbjct: 703 YDEDINDEEF-QEDEHEDLSSSYKSDVESEPDLSDDPSWLEKIQKSVWNIIQVVNIFQTP 879
Query: 297 LLFREAAFLKKDFGSDCEKLSWLAQRM 323
+ +AA ++K++ KLS + R+
Sbjct: 880 VNQSDAARIRKEYDESSAKLSKIQSRI 960
>TC82511 similar to PIR|A96606|A96606 unknown protein F13N6.17 [imported] -
Arabidopsis thaliana, partial (63%)
Length = 752
Score = 34.7 bits (78), Expect = 0.11
Identities = 20/53 (37%), Positives = 33/53 (61%)
Frame = +3
Query: 31 LWDLVGEEYGLGVKVGSSVKLVYSKYLSGLETPLKNVVDGEFPKRDLVGDRVK 83
+WDL+ + GLG ++ +V+ V+ + LSG E PL +++GE P+ D D K
Sbjct: 501 VWDLILDNNGLGKEISETVERVFCR-LSGQEPPLFPLLNGE-PQPDKEADSRK 653
>TC78117 similar to GP|10177445|dbj|BAB10741. protein kinase {Arabidopsis
thaliana}, partial (15%)
Length = 1351
Score = 30.8 bits (68), Expect = 1.5
Identities = 23/70 (32%), Positives = 30/70 (42%)
Frame = +1
Query: 216 DGSKSCANDDANDSGDDVLMLDPSSVNRESFGRKRKRDDLMSEMQSWVIRTAKNPCDPVL 275
DG ++ + GDDVL SS N D S+MQ W I +K CD L
Sbjct: 604 DGVTKLYVEEQSSEGDDVL--SGSSKNSSCV-------DEESDMQGWNITKSKWACDDDL 756
Query: 276 GSMPEKSKWK 285
M + K+K
Sbjct: 757 SPMADDGKYK 786
>TC92211 similar to PIR|B86182|B86182 hypothetical protein [imported] -
Arabidopsis thaliana, partial (31%)
Length = 798
Score = 30.0 bits (66), Expect = 2.6
Identities = 15/53 (28%), Positives = 27/53 (50%)
Frame = +3
Query: 8 LDLYKLFMVVKDKGGYDVVCKNELWDLVGEEYGLGVKVGSSVKLVYSKYLSGL 60
LDL++LF+ V +GG++ + K+ W V + ++ V KY + L
Sbjct: 252 LDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNF-PSTATNASFVLRKYYTSL 407
>BG647146 similar to GP|15146198|gb| F17F16.8/F17F16.8 {Arabidopsis
thaliana}, partial (32%)
Length = 634
Score = 28.5 bits (62), Expect = 7.7
Identities = 19/69 (27%), Positives = 34/69 (48%), Gaps = 6/69 (8%)
Frame = +1
Query: 276 GSMPEKSKWKSYGNQEIWKKVLLFREAAFLKKDFGSDCEKLSWLAQRMHP------SMYD 329
GSM E K++ GN+ + L E FL++ + ++L+ +A R H + Y
Sbjct: 253 GSMNEGDKYEGKGNRASFDDHLTPAERRFLQQTEKLELQRLAKMASRSHRDRIQQFNQYL 432
Query: 330 VNLGVNYNL 338
NL +Y++
Sbjct: 433 ANLSEHYDI 459
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.133 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,798,294
Number of Sequences: 36976
Number of extensions: 267699
Number of successful extensions: 920
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 912
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 918
length of query: 600
length of database: 9,014,727
effective HSP length: 102
effective length of query: 498
effective length of database: 5,243,175
effective search space: 2611101150
effective search space used: 2611101150
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC149204.7


