

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149204.4 - phase: 0 /pseudo
(640 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
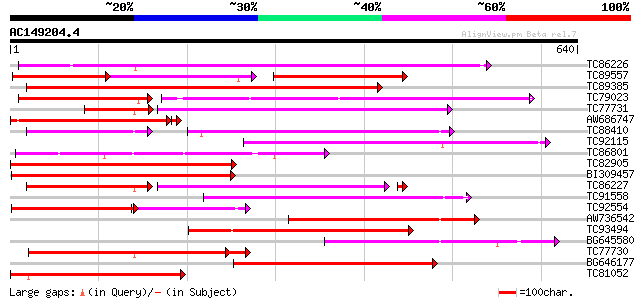
Score E
Sequences producing significant alignments: (bits) Value
TC86226 weakly similar to GP|18033111|gb|AAL56987.1 functional c... 382 e-106
TC89557 weakly similar to GP|10178211|dbj|BAB11635. TMV resistan... 127 5e-88
TC89385 similar to GP|19774173|gb|AAL99063.1 NBS-LRR-Toll resist... 320 1e-87
TC79023 weakly similar to GP|18033111|gb|AAL56987.1 functional c... 292 2e-79
TC77731 weakly similar to GP|18033111|gb|AAL56987.1 functional c... 256 1e-68
AW686747 weakly similar to PIR|A54810|A5 TMV resistance protein ... 254 3e-68
TC88410 weakly similar to GP|12056928|gb|AAG48132.1 putative res... 158 3e-64
TC92115 weakly similar to GP|18033111|gb|AAL56987.1 functional c... 233 2e-61
TC86801 similar to PIR|A54810|A54810 TMV resistance protein N - ... 226 2e-59
TC82905 weakly similar to GP|3947733|emb|CAA08797.1 NL25 {Solanu... 218 6e-57
BI309457 weakly similar to GP|15787901|gb resistance gene analog... 217 1e-56
TC86227 weakly similar to GP|18033111|gb|AAL56987.1 functional c... 209 2e-55
TC91558 weakly similar to GP|19774201|gb|AAL99077.1 NBS-LRR-Toll... 210 2e-54
TC92554 similar to GP|9758876|dbj|BAB09430.1 disease resistance ... 150 2e-54
AW736542 similar to PIR|T07656|T076 probable resistance protein ... 204 8e-53
TC93494 similar to PIR|A54810|A54810 TMV resistance protein N - ... 202 2e-52
BG645580 similar to PIR|T07656|T076 probable resistance protein ... 198 6e-51
TC77730 weakly similar to GP|7341113|gb|AAF61210.1| unknown {Gly... 177 3e-49
BG646177 weakly similar to GP|18033111|g functional candidate re... 184 1e-46
TC81052 weakly similar to GP|10178211|dbj|BAB11635. TMV resistan... 178 6e-45
>TC86226 weakly similar to GP|18033111|gb|AAL56987.1 functional candidate
resistance protein KR1 {Glycine max}, partial (24%)
Length = 1769
Score = 382 bits (981), Expect = e-106
Identities = 223/542 (41%), Positives = 325/542 (59%), Gaps = 9/542 (1%)
Frame = +3
Query: 11 SSSTTSLCTYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDD--KDLERGQVISEKLI 68
SSS + +Y VFL +G DTR GFT +L AL KGI TF DD DL+R ++ +I
Sbjct: 63 SSSFSYAFSYDVFLICKGTDTRYGFTGNLLKALIDKGIRTFHDDDDSDLQRRDKVTPIII 242
Query: 69 NAIKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCF 128
++S I I S +YASS+ CLD L I+ C VLPVF+GV+P+DVRH G +
Sbjct: 243 ---EESRILIPIFSANYASSSSCLDTLVHIIHCYKTKGCLVLPVFFGVEPTDVRHHTGRY 413
Query: 129 EEAFRKHQEKFG---QHSDRVDRWRDAFTQVASYSGW--DSKGQHEASLVENIAQHIHRK 183
+A +H+ +F ++ +R+ +W+ A + A+ + DS G +E L+ I ++I K
Sbjct: 414 GKALAEHENRFQNDTKNMERLQQWKVALSLAANLPSYHDDSHG-YEYELIGKIVKYISNK 590
Query: 184 LVPKLPSCTENLVGIVSKVEEVNKFLGMGLND-VRFIGIWGMGGIGKSTIARAVYETIRC 242
+ + VG+ S+V++V L G +D V +GI+G+GG GKST+ARA+Y +
Sbjct: 591 ISRQSLHVATYPVGLQSRVQQVKSLLDEGPDDGVHMVGIYGIGGSGKSTLARAIYNFVAD 770
Query: 243 EFELTCFLENVREISETNGLVHLQRQLLSHLSISRNDFHDLYDGKKTIQNSLCRKKVLLV 302
+FE CFLE VRE S +N L Q LLS + D+ +G I+ LCRKK+LL+
Sbjct: 771 QFEGLCFLEQVRENSASNSLKRFQEMLLSKTLQLKIKLADVSEGISIIKERLCRKKILLI 950
Query: 303 LDDVNELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFC 362
LDDV+ + QL L G DWFGPGSRVIITTRDKHLL H + KTY L +AL L
Sbjct: 951 LDDVDNMKQLNALAGGVDWFGPGSRVIITTRDKHLLACHEIEKTYAVKGLNVTEALELLR 1130
Query: 363 LKAFKGDKPQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPR 422
AFK DK Y + VV Y GLP+ +E++GS L+G+NI+ + + P+
Sbjct: 1131WMAFKNDKVPSSYEKILNRVVAYASGLPVVIEIVGSNLFGKNIEECKNTLDWYEKIPNKE 1310
Query: 423 VQDNLKISYDSLDTMEKDIFLDIACFFKGMKGDKVIDILES-CGYFPQIGIQILIERSLI 481
+Q LK+SYDSL+ E+ +FLDIAC FKG K +KV +IL + G+ +++L+E+ LI
Sbjct: 1311IQRILKVSYDSLEEEEQSVFLDIACCFKGCKWEKVKEILHAHYGHCINHHVEVLVEKCLI 1490
Query: 482 TLDSVNNKLGMHDLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSID 541
++ + +H+L++ MG+++V ESP +P +RSRLW ++DI VL +N ++ ID
Sbjct: 1491DHFEYDSHVSLHNLIENMGKELVRLESPFEPGKRSRLWFEKDIFEVLEEN----TVSKID 1658
Query: 542 MK 543
M+
Sbjct: 1659MQ 1664
>TC89557 weakly similar to GP|10178211|dbj|BAB11635. TMV resistance protein
N {Arabidopsis thaliana}, partial (8%)
Length = 1567
Score = 127 bits (320), Expect(3) = 5e-88
Identities = 68/181 (37%), Positives = 101/181 (55%), Gaps = 11/181 (6%)
Frame = +1
Query: 109 VLPVFYGVDPSDVRHQRGCFEEAFRKHQEKFGQHSDRVDRWRDAFTQVASYSGWDSKGQH 168
V PVFY VDPS V+ Q G +E AF H E F +S +V RWR T + +GWD + +
Sbjct: 532 VFPVFYDVDPSHVKKQNGVYENAFVLHTEAFKHNSGKVARWRTTMTYLGGTAGWDVRVEP 711
Query: 169 EASLVENIAQHIHRKLVPKLPSCTENLVGIVSKVEEVNKFLGMGLND--VRFIGIWGMGG 226
E ++E I + + +KL T++L+GI +E + L + D R +GIWGM G
Sbjct: 712 EFEMIEKIVEAVIKKLGRTFSGSTDDLIGIQPHIEALENLLKLSSKDDGCRVLGIWGMDG 891
Query: 227 IGKSTIARAVYETIRCEFELTCFLENVREI---------SETNGLVHLQRQLLSHLSISR 277
IGK+T+A +Y+ I +F+ CF+ENV +I +ETN L + +R+ H+ S
Sbjct: 892 IGKTTLATVLYDKISFQFDACCFIENVSKIYEDGGACCCAETNYLSNNRRKKYRHMQCS* 1071
Query: 278 N 278
N
Sbjct: 1072N 1074
Score = 127 bits (319), Expect(3) = 5e-88
Identities = 62/152 (40%), Positives = 96/152 (62%)
Frame = +3
Query: 298 KVLLVLDDVNELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDA 357
K+L+VLD+V +L QLE L + + P S++II TRDKH+L +G + ++ ++ DA
Sbjct: 1110 KLLIVLDNVEQLEQLEKLDIEPKFLHPRSKIIIITRDKHILQAYGADEVFEAKLMNDEDA 1289
Query: 358 LVLFCLKAFKGDKPQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRS 417
L C KAFK D P G+ +L +V+ Y LPLA++VLGS+L+ RN + W S + K
Sbjct: 1290 HKLLCRKAFKSDYPSSGFAELIPKVLVYAQRLPLAVKVLGSFLFSRNANEWSSTLDKFEK 1469
Query: 418 FPHPRVQDNLKISYDSLDTMEKDIFLDIACFF 449
P ++ L++SY+ L+ EK++FL +A FF
Sbjct: 1470 NPPNKIMKALQVSYEGLEKDEKEVFLHVAXFF 1565
Score = 110 bits (274), Expect(3) = 5e-88
Identities = 55/110 (50%), Positives = 69/110 (62%)
Frame = +3
Query: 4 SSSSSCRSSSTTSLCTYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVI 63
++SSS S++ Y VF+SFRG DTR F DHL A L RKGI TFKDDK L++G+ I
Sbjct: 216 TASSSSTDSNSNQSYRYDVFISFRGSDTRNSFVDHLYAHLNRKGIFTFKDDKQLQKGEAI 395
Query: 64 SEKLINAIKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVF 113
S +L+ AI+ S +I + S DYASSTWCLDE+ I E P F
Sbjct: 396 SPQLLQAIQQSRISIVVFSKDYASSTWCLDEMAAINESRINLKTSCFPCF 545
>TC89385 similar to GP|19774173|gb|AAL99063.1 NBS-LRR-Toll resistance gene
analog protein {Medicago ruthenica}, partial (98%)
Length = 1285
Score = 320 bits (819), Expect = 1e-87
Identities = 171/405 (42%), Positives = 259/405 (63%), Gaps = 4/405 (0%)
Frame = +3
Query: 20 YHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINAIKDSMFAIT 79
Y VF++FRGEDTR FT+ L AALERKGI F+DD +L +G+ I +L+ I+ S +
Sbjct: 69 YDVFVTFRGEDTRNNFTNFLFAALERKGIYAFRDDTNLPKGESIGPELLRTIEGSQVFVA 248
Query: 80 ILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQEKF 139
+LS +YASSTWCL EL+ I EC + +VLP+FYGVDPS+V+ Q G + + F KH+++F
Sbjct: 249 VLSRNYASSTWCLQELEKICECIKGSGKYVLPIFYGVDPSEVKKQSGIYWDDFAKHEQRF 428
Query: 140 GQHSDRVDRWRDAFTQVASYSGWDSKGQHEASLVENIAQHIHRKLVPKLPSCTENLVGIV 199
Q +V RWR+A QV S +GWD + + ++ VE I Q I L K +++LVGI
Sbjct: 429 KQDPHKVSRWREALNQVGSIAGWDLRDKQQSVEVEKIVQTILNILKCKSSFVSKDLVGIN 608
Query: 200 SKVEEV-NKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETIRCEFELTCFLENVREISE 258
S+ E + ++ L ++ VR IGIWGMGG GK+T+A +Y I F+ +CF+++V +I
Sbjct: 609 SRTEALKHQLLLNSVDGVRVIGIWGMGGKGKTTLAMNLYGQICHRFDASCFIDDVSKIFR 788
Query: 259 T-NGLVHLQRQLLSH-LSISRNDFHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQLENLV 316
+G + Q+Q+L L I + + Y I++ L R+K LL+LD+V+++ QLE +
Sbjct: 789 LHDGPIDAQKQILHQTLGIEHHQICNHYSATDLIRHRLSREKTLLILDNVDQVEQLERIG 968
Query: 317 GKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDK-PQEGY 375
++W G GSR++I +RD+H+L + V YK +L ++ LF KAFK +K + Y
Sbjct: 969 VHREWLGAGSRIVIISRDEHILKEYKVDVDYKVPLLDWTESHKLFRQKAFKLEKIIMKNY 1148
Query: 376 LDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPH 420
+L+ E+++Y GLPL + VLGS+L GRN+ W SA+ +LR P+
Sbjct: 1149QNLAYEILNYANGLPLTITVLGSFLSGRNVTEWKSALARLRQRPN 1283
>TC79023 weakly similar to GP|18033111|gb|AAL56987.1 functional candidate
resistance protein KR1 {Glycine max}, partial (37%)
Length = 2072
Score = 292 bits (748), Expect = 2e-79
Identities = 181/428 (42%), Positives = 254/428 (59%), Gaps = 7/428 (1%)
Frame = +3
Query: 172 LVENIAQHIHRKL--VPKLPSCTENLVGIVSKVEEVNKFLGMGLND-VRFIGIWGMGGIG 228
+VE+I+ I R+ V K P VG+ S+V+ V L +D V +G++G GGIG
Sbjct: 714 IVEDISNRISREPLDVAKYP------VGLQSRVQHVKGHLDEKSDDEVHMVGLYGTGGIG 875
Query: 229 KSTIARAVYETIRCEFELTCFLENVREISETNGLVHLQRQLLSHLSISRNDFH--DLYDG 286
KST+A+A+Y I +FE+ CFLENVR S ++ L HLQ +LL L R D + G
Sbjct: 876 KSTLAKAIYNFIADQFEVLCFLENVRVNSTSDNLKHLQEKLL--LKTVRLDIKLGGVSQG 1049
Query: 287 KKTIQNSLCRKKVLLVLDDVNELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKT 346
I+ LCRKK+LL+LDDV++L+QLE L G DWFGPGSRVIITTR+KHLL HG+ T
Sbjct: 1050 IPIIKQRLCRKKILLILDDVDKLDQLEALAGGLDWFGPGSRVIITTRNKHLLKIHGIEST 1229
Query: 347 YKTGMLCKHDALVLFCLKAFKGDKPQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNID 406
+ L +AL L AFK + P + D+ + Y GLPLA+ ++GS L GR++
Sbjct: 1230 HAVEGLNATEALELLRWMAFKENVP-SSHEDILNRALTYASGLPLAIVIIGSNLVGRSVQ 1406
Query: 407 VWHSAVKKLRSFPHPRVQDNLKISYDSLDTMEKDIFLDIACFFKGMKGDKVIDILES-CG 465
S + P+ +Q LK+SYDSL+ E+ +FLDIAC FKG K +V +IL + G
Sbjct: 1407 DSMSTLDGYEEIPNKEIQRILKVSYDSLEKEEQSVFLDIACCFKGCKWPEVKEILHAHYG 1586
Query: 466 YFPQIGIQILIERSLITLDSVNNKLGMHDLLQEMGRDIVFQESPNDPCRRSRLWSQEDID 525
+ + +L E+SL+ ++ + +HDL+++MG+++V QESP++P RSRLW + DI
Sbjct: 1587 HCIVHHVAVLAEKSLMDHLKYDSYVTLHDLIEDMGKEVVRQESPDEPGERSRLWFERDIV 1766
Query: 526 RVLTKNKGTEAINSIDMKL-LQPYEAHWNTEAFSKTSQLKFLSLCEMQLPLGLSCLPSSL 584
VL KN GT I I+MK + WN AF K + LK L LPSSL
Sbjct: 1767 HVLKKNTGTRKIKMINMKFPSMESDIDWNGNAFEKMTNLKTFITENGHHSKSLEYLPSSL 1946
Query: 585 KVLHWRGC 592
+V+ +GC
Sbjct: 1947 RVM--KGC 1964
Score = 141 bits (356), Expect = 7e-34
Identities = 76/154 (49%), Positives = 97/154 (62%), Gaps = 3/154 (1%)
Frame = +1
Query: 11 SSSTTSLCTYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINA 70
++ + S TY VFLSFRG DTR GFT +L AL KGI TF DD DL+RG I+ L NA
Sbjct: 67 ATQSPSSFTYQVFLSFRGADTRHGFTGNLYKALTDKGIYTFIDDNDLQRGDEITPSLKNA 246
Query: 71 IKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEE 130
I+ S I + S +YASS++CLDEL I C VLPVF GVDP+DVRH G + E
Sbjct: 247 IEKSRIFIPVFSENYASSSFCLDELVHITHCYDTKGCLVLPVFIGVDPTDVRHHTGRYGE 426
Query: 131 AFRKHQEKFGQHSD---RVDRWRDAFTQVASYSG 161
A H++KF D R+ +W++A +Q A+ SG
Sbjct: 427 ALAVHKKKFQNDKDNTERLQQWKEALSQAANLSG 528
>TC77731 weakly similar to GP|18033111|gb|AAL56987.1 functional candidate
resistance protein KR1 {Glycine max}, partial (23%)
Length = 1651
Score = 256 bits (655), Expect = 1e-68
Identities = 141/336 (41%), Positives = 204/336 (59%), Gaps = 2/336 (0%)
Frame = +1
Query: 167 QHEASLVENIAQHIHRKLVPKLPSCTENLVGIVSKVEEVNKFLGMGLND-VRFIGIWGMG 225
++E +E I + I + + + VG+ S++E+V L MG D VR +G++G G
Sbjct: 496 RYEYKFIEKIVEDISNNINHVFLNVAKYPVGLQSRIEQVKLLLDMGSEDEVRMVGLFGTG 675
Query: 226 GIGKSTIARAVYETIRCEFELTCFLENVREISETNGLVHLQRQLLSHLSISRNDFHDLYD 285
G+GKST+A+AVY + +FE CFL NVRE S L HLQ++LLS + D+ +
Sbjct: 676 GMGKSTLAKAVYNFVADQFEGVCFLHNVRENSTHKNLKHLQKKLLSKIVKFDGKLEDVSE 855
Query: 286 GKKTIQNSLCRKKVLLVLDDVNELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHK 345
G I+ L RKK+LL+LDDV++L QLE L G DWFG GSRVIITTRDKHLL HG+
Sbjct: 856 GIPIIKERLSRKKILLILDDVDKLEQLEALAGGLDWFGHGSRVIITTRDKHLLACHGITS 1035
Query: 346 TYKTGMLCKHDALVLFCLKAFKGDKPQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNI 405
T+ L + +AL L AFK DK Y ++ VV Y GLPLA+ +G L+GR +
Sbjct: 1036THAVEELNETEALELLRRMAFKNDKVPSTYEEILNRVVTYASGLPLAIVTIGDNLFGRKV 1215
Query: 406 DVWHSAVKKLRSFPHPRVQDNLKISYDSLDTMEKDIFLDIACFFKGMKGDKVIDILES-C 464
+ W + + + P+ +Q L++SYD+L+ EK +FLDIAC FKG K KV IL +
Sbjct: 1216EDWKRILDEYENIPNKDIQRILQVSYDALEPKEKSVFLDIACCFKGCKWTKVKKILHAHY 1395
Query: 465 GYFPQIGIQILIERSLITLDSVNNKLGMHDLLQEMG 500
G+ + + +L E+SLI + ++ +HDL+++MG
Sbjct: 1396GHCIEHHVGVLAEKSLIGHWEYDTQMTLHDLIEDMG 1503
Score = 75.9 bits (185), Expect = 5e-14
Identities = 36/81 (44%), Positives = 54/81 (66%), Gaps = 3/81 (3%)
Frame = +2
Query: 85 YASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQEKF---GQ 141
YASS++CLDEL I+ C + VLPVFY V+P+ +RHQ G + E KH+E+F +
Sbjct: 2 YASSSFCLDELVHIIHCYKTKSCLVLPVFYDVEPTHIRHQSGSYGEYLTKHEERFQNNEK 181
Query: 142 HSDRVDRWRDAFTQVASYSGW 162
+ +R+ +W+ A TQ A+ SG+
Sbjct: 182 NMERLRQWKIALTQAANLSGY 244
>AW686747 weakly similar to PIR|A54810|A5 TMV resistance protein N - tobacco
(Nicotiana glutinosa), partial (10%)
Length = 644
Score = 254 bits (650), Expect(2) = 3e-68
Identities = 127/181 (70%), Positives = 147/181 (81%)
Frame = +3
Query: 2 EASSSSSCRSSSTTSLCTYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQ 61
EASSSSS S TS T HVFLSFRGEDTR+GFTDHL A+LER+GI TFKDD DLERG+
Sbjct: 72 EASSSSS--PSIQTSRWTNHVFLSFRGEDTRQGFTDHLFASLERRGIKTFKDDHDLERGE 245
Query: 62 VISEKLINAIKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDV 121
VIS +L AI++SMFAI ILSP+YASSTWCLDEL+ I+ECS V P+FYGVDPSDV
Sbjct: 246 VISYELNKAIEESMFAIIILSPNYASSTWCLDELKKIVECSKSFGQAVFPIFYGVDPSDV 425
Query: 122 RHQRGCFEEAFRKHQEKFGQHSDRVDRWRDAFTQVASYSGWDSKGQHEASLVENIAQHIH 181
RHQRG F+EAFRKH+EKF + +V+RWRDA +VA YSGWDSKG+HEASLVE I +HI
Sbjct: 426 RHQRGSFDEAFRKHEEKFRKDRTKVERWRDALREVAGYSGWDSKGRHEASLVETIVEHIQ 605
Query: 182 R 182
+
Sbjct: 606 K 608
Score = 22.7 bits (47), Expect(2) = 3e-68
Identities = 8/12 (66%), Positives = 10/12 (82%)
Frame = +2
Query: 183 KLVPKLPSCTEN 194
KL+PKL CT+N
Sbjct: 608 KLIPKLKVCTDN 643
>TC88410 weakly similar to GP|12056928|gb|AAG48132.1 putative resistance
protein {Glycine max}, partial (8%)
Length = 2252
Score = 158 bits (399), Expect(2) = 3e-64
Identities = 102/310 (32%), Positives = 177/310 (56%), Gaps = 8/310 (2%)
Frame = +1
Query: 201 KVEEVNKFLGMGLND----VRFIGIWGMGGIGKSTIARAVYETIR-CEFELTCFLENVRE 255
+VE+V ++L V +GI G+ GIGK+T+AR VY EF+ CF +NV E
Sbjct: 604 RVEKVMRYLNSSPRSDDDGVCLVGICGVPGIGKTTLARGVYHFGGGTEFDSCCFFDNVGE 783
Query: 256 ISETNGLVHLQRQLLSHLSISRND--FHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQLE 313
+ +GLVHLQ+ LLS + N F ++ + I++ L +KKV LVL+DV++ L+
Sbjct: 784 YVKKHGLVHLQQMLLSAIVGHNNSTMFENVDERVWKIKHMLNQKKVFLVLEDVHDSEVLK 963
Query: 314 NLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQE 373
+V +FG GS+VIIT R+K L HG+ + Y+ + K +A L LKAF
Sbjct: 964 AIVKLSTFFGSGSKVIITAREKCFLEFHGIKRIYEVERMNKTEAFQLLNLKAFDSMNISP 1143
Query: 374 GYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPRVQDNLKISYDS 433
++ + + + Y G P LE++GSYL G++++ SA+ + + + ++ L++S+D+
Sbjct: 1144CHVTILEGLETYASGHPFILEMIGSYLSGKSMEECESALHQYKEISNRDIKKILQVSFDA 1323
Query: 434 LDTMEKDIFLDIACFFKGMKGDKVIDIL-ESCGYFPQIGIQILIERSLITLDSVNNKLGM 492
L+ ++++ + IA + + + V ++L G P+ I++L+ +SLI ++ N + +
Sbjct: 1324LEKSQQNMLIHIALHLREQELEMVENLLHRKYGVCPKYDIRVLLNKSLIKINE-NGHVIV 1500
Query: 493 HDLLQEMGRD 502
H L Q+M RD
Sbjct: 1501HVLTQDMVRD 1530
Score = 106 bits (264), Expect(2) = 3e-64
Identities = 60/143 (41%), Positives = 87/143 (59%), Gaps = 1/143 (0%)
Frame = +3
Query: 20 YHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINAIKDSMFAIT 79
Y F+SFRG DTR GF+ L L +G TF DD +LERG I+ ++ AI++S I
Sbjct: 162 YSTFISFRGSDTRCGFSGFLNKYLIDRGFRTFFDDGELERGTQITVEIPKAIEESRIFIP 341
Query: 80 ILSPDYASSTWCLDELQMIMECSSK-NNLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQEK 138
+LS +YASS++CLDEL I+E K N V PVFY ++ SDV++Q G + +A H+ +
Sbjct: 342 VLSENYASSSFCLDELVKILEEFKKGNGRWVFPVFYYINISDVKNQTGSYGQALAVHKNR 521
Query: 139 FGQHSDRVDRWRDAFTQVASYSG 161
+R ++W +A VA + G
Sbjct: 522 V--MPERFEKWINALASVADFRG 584
>TC92115 weakly similar to GP|18033111|gb|AAL56987.1 functional candidate
resistance protein KR1 {Glycine max}, partial (24%)
Length = 1348
Score = 233 bits (594), Expect = 2e-61
Identities = 138/352 (39%), Positives = 204/352 (57%), Gaps = 6/352 (1%)
Frame = +2
Query: 265 LQRQLLSHLSISRNDFHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQLENLVGKQDWFGP 324
LQ +LL + + +G I+ L KK LL+LDDV+++ QL L G DWFG
Sbjct: 2 LQEELLLKALQLKIKLGGVSEGIPYIKERLHTKKTLLILDDVDDMEQLHALAGGPDWFGR 181
Query: 325 GSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQEGYLDLSKEVVD 384
GSRVIITTRDKHLL +HG+ T++ L +AL L AFK +K Y D+ V
Sbjct: 182 GSRVIITTRDKHLLRSHGIESTHEVEELYGTEALELLRWMAFKNNKVPSIYEDVLNRAVS 361
Query: 385 YCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPRVQDNLKISYDSLDTMEKDIFLD 444
Y GLPL LE++GS L+G+ I+ W + P+ ++ LK+SYD+L+ ++ +FLD
Sbjct: 362 YASGLPLVLEIVGSNLFGKTIEEWKGTLDGYEKIPNKKIHQILKVSYDALEEEQQSVFLD 541
Query: 445 IACFFKGMKGDKVIDILES-CGYFPQIGIQILIERSLITLDSVN----NKLGMHDLLQEM 499
IAC FKG ++ IL + G+ + +L E+SL+ + + +L +HDL++EM
Sbjct: 542 IACCFKGCGWEEFEYILRAHYGHRITHHLVVLAEKSLVKITHPHYGSIYELTLHDLIKEM 721
Query: 500 GRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSIDMKL-LQPYEAHWNTEAFS 558
G+++V QESP +P RSRLW ++DI VL +N GT I I M + + +AF
Sbjct: 722 GKEVVRQESPKEPGERSRLWCEDDIVNVLKENTGTSKIEMIYMNFPSEEFVIDKKGKAFK 901
Query: 559 KTSQLKFLSLCEMQLPLGLSCLPSSLKVLHWRGCPLKTLPITTQLDELVDIT 610
K ++LK L + + GL LPSSL+VL RGC ++L I+ L + ++IT
Sbjct: 902 KMTRLKTLIIENVHFSKGLKYLPSSLRVLKLRGCLSESL-ISCSLSKAIEIT 1054
>TC86801 similar to PIR|A54810|A54810 TMV resistance protein N - tobacco
(Nicotiana glutinosa), partial (7%)
Length = 1268
Score = 226 bits (577), Expect = 2e-59
Identities = 142/366 (38%), Positives = 213/366 (57%), Gaps = 11/366 (3%)
Frame = +2
Query: 7 SSCRSSSTTSLCTYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEK 66
S SS + + Y VF+SFRGEDTR GFT HL AAL R + T+ D + +E+G + +
Sbjct: 29 SLASSSHSAAQKKYDVFISFRGEDTRAGFTSHLHAALSRTYLHTYIDYR-IEKGDEVWPE 205
Query: 67 LINAIKDSMFAITILSPDYASSTWCLDELQMIMECSSKN---NLHVLPVFYGVDPSDVRH 123
L AIK S + + S +YASSTWCL+EL +MEC +KN N+ V+PVFY VDPS VR
Sbjct: 206 LEKAIKQSTLFLVVFSENYASSTWCLNELVELMECRNKNEDDNIGVIPVFYHVDPSHVRK 385
Query: 124 QRGCFEEAFRKHQEKFGQHSDRVDRWRDAFTQVASYSGWDSKGQH-EASLVENIAQHIHR 182
Q G + A KH+++ Q + W++A Q A+ SG+ S E++++E+I + +
Sbjct: 386 QTGSYGSALAKHKQE-NQDDKMMQNWKNALFQAANLSGFHSSTYRTESNMIEDITRALLG 562
Query: 183 KLVPKL-PSCTENLVGIVSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETIR 241
KL + T NL+ + V + V+ IG+WGMGG GK+T+A A+++
Sbjct: 563 KLNHQYRDELTCNLI-LDENYWAVRSLIKFDSTTVQIIGLWGMGGTGKTTLAAAMFQRFS 739
Query: 242 CEFELTCFLENVREISETNGLVHLQRQLLSHLSISRNDFHDL-YDGKKTIQNSLCRK--- 297
++E CFLE V E+S+ +G+ + +LLS L DL D K I + R+
Sbjct: 740 FKYEGNCFLERVTEVSKKHGINYTCNKLLSKLL-----GEDLRIDTPKVIPAMIKRRLRH 904
Query: 298 -KVLLVLDDVNELNQLENLVG-KQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKH 355
K +VLDDV+ L++L+G + W GPGS VI+TTRDKH+L++ G+ + Y+ +
Sbjct: 905 MKSFIVLDDVHNSELLQDLIGVRGGWLGPGSIVIVTTRDKHVLISGGIDEIYEVKKMNSQ 1084
Query: 356 DALVLF 361
++L LF
Sbjct: 1085NSLQLF 1102
>TC82905 weakly similar to GP|3947733|emb|CAA08797.1 NL25 {Solanum
tuberosum}, partial (22%)
Length = 832
Score = 218 bits (555), Expect = 6e-57
Identities = 118/257 (45%), Positives = 163/257 (62%), Gaps = 1/257 (0%)
Frame = +1
Query: 1 MEASSSSSCRSSSTTSLCTYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERG 60
M +SS SS +++ Y VF++FRGEDTR FTD L ALE KGI F+D +L++G
Sbjct: 31 MASSSKSSSALVTSSKKNHYDVFVTFRGEDTRNNFTDFLFDALETKGIMVFRDVINLQKG 210
Query: 61 QVISEKLINAIKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSD 120
+ I +L AI+ S + I S +YASSTWCL EL+ I EC + HVLPVFY VDPS+
Sbjct: 211 ECIGPELFRAIEISQVYVAIFSKNYASSTWCLQELEKICECIKGSGKHVLPVFYDVDPSE 390
Query: 121 VRHQRGCFEEAFRKHQEKFGQHSDRVDRWRDAFTQVASYSGWDSKGQHEASLVENIAQHI 180
VR Q G + EAF KH+++F Q S +V RWR+A QV S SGWD + + A ++ I Q I
Sbjct: 391 VRKQSGIYSEAFVKHEQRFQQDSMKVSRWREALEQVGSISGWDLRDEPLAREIKEIVQKI 570
Query: 181 HRKLVPKLPSCTENLVGIVSKVEEV-NKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYET 239
L K +++LVGI S ++ + N L ++ VR IGI GMGGIGK+T+A +Y
Sbjct: 571 INILECKYSCVSKDLVGIDSPIQALQNHLLLNSVDGVRAIGICGMGGIGKTTLATTLYGQ 750
Query: 240 IRCEFELTCFLENVREI 256
I +F +CF+++V +I
Sbjct: 751 ISHQFSASCFIDDVTKI 801
>BI309457 weakly similar to GP|15787901|gb resistance gene analog NBS7
{Helianthus annuus}, partial (45%)
Length = 806
Score = 217 bits (553), Expect = 1e-56
Identities = 117/255 (45%), Positives = 165/255 (63%), Gaps = 2/255 (0%)
Frame = +3
Query: 3 ASSSSSCRSSSTTSLCTYH-VFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQ 61
AS+S+S T+S Y+ VF++FRGEDTR FTD L AL+ KGI F DD +L +G+
Sbjct: 18 ASTSNSSSVLGTSSRRNYYDVFVTFRGEDTRNNFTDFLFDALQTKGIIVFSDDTNLPKGE 197
Query: 62 VISEKLINAIKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDV 121
I +L+ AI+ S + + S +YASSTWCL EL+ I EC + HVLPVFY VDPSDV
Sbjct: 198 SIGPELLRAIEGSQVFVAVFSINYASSTWCLQELEKICECVKGSGKHVLPVFYDVDPSDV 377
Query: 122 RHQRGCFEEAFRKHQEKFGQHSDRVDRWRDAFTQVASYSGWDSKGQHEASLVENIAQHIH 181
R Q G + EAF KH+++F Q +V +WRDA QV S SGWD + + +A ++ I Q I
Sbjct: 378 RKQSGIYGEAFIKHEQRFQQEFQKVSKWRDALKQVGSISGWDLRDKPQAGEIKKIVQTIL 557
Query: 182 RKLVPKLPSCTENLVGIVSKVEEV-NKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETI 240
L K +++LVGI S+++ + N L ++ VR IGI GMGGIGK+T+A A+Y+ I
Sbjct: 558 NILKYKSSCFSKDLVGIDSRLDGLQNHLLLDSVDSVRAIGICGMGGIGKTTLAMALYDQI 737
Query: 241 RCEFELTCFLENVRE 255
F + F+++V++
Sbjct: 738 SHRFSASWFIDDVKQ 782
>TC86227 weakly similar to GP|18033111|gb|AAL56987.1 functional candidate
resistance protein KR1 {Glycine max}, partial (22%)
Length = 1736
Score = 209 bits (532), Expect(2) = 2e-55
Identities = 112/263 (42%), Positives = 159/263 (59%), Gaps = 1/263 (0%)
Frame = +1
Query: 167 QHEASLVENIAQHIHRKLVPKLPSCTENLVGIVSKVEEVNKFLGMGLND-VRFIGIWGMG 225
++E L+ENI +HI ++ + VG+ S+V++V L ++ V +G++G
Sbjct: 679 RYEYELIENIVEHISDRINRVFLHVAKYPVGLQSRVQQVKLLLDEESDEGVNMVGLYGTR 858
Query: 226 GIGKSTIARAVYETIRCEFELTCFLENVREISETNGLVHLQRQLLSHLSISRNDFHDLYD 285
+GKST+A+A+Y I +FE CFL NVRE S L HLQ++LLS D+ +
Sbjct: 859 ALGKSTLAKAIYNFIADQFEGVCFLHNVRENSARKNLKHLQKELLSKTVQLNIKLRDVSE 1038
Query: 286 GKKTIQNSLCRKKVLLVLDDVNELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHK 345
G I+ LCRKK+LL+LDDV++L+QLE L G DWFGPGSRVIITTRDKHLL HG+ +
Sbjct: 1039GIPIIKERLCRKKILLILDDVDQLDQLEALAGGLDWFGPGSRVIITTRDKHLLTCHGIER 1218
Query: 346 TYKTGMLCKHDALVLFCLKAFKGDKPQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNI 405
TY L +AL L AFK +K Y D+ V Y G+PL LE++GS L+G+NI
Sbjct: 1219TYAVRGLYGKEALELLRWTAFKNNKVPPSYEDVLNRAVSYGSGIPLVLEIVGSNLFGKNI 1398
Query: 406 DVWHSAVKKLRSFPHPRVQDNLK 428
+VW + + P+ +Q L+
Sbjct: 1399EVWKNTLDGYDRIPNKEIQKILR 1467
Score = 140 bits (354), Expect = 1e-33
Identities = 73/145 (50%), Positives = 94/145 (64%), Gaps = 3/145 (2%)
Frame = +3
Query: 20 YHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINAIKDSMFAIT 79
YHVFLSFR DT GFT +L AL KGI TF DD DLERG + L+ AI++S I
Sbjct: 18 YHVFLSFRDIDTLYGFTGNLYKALIDKGIKTFIDDNDLERGDESTPSLVKAIEESRILIP 197
Query: 80 ILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQEKF 139
I S +YASS++CLDEL I+ C VLPVFYG DP+ VRHQ G + E KH++KF
Sbjct: 198 IFSANYASSSFCLDELVHIIHCYKTRGCSVLPVFYGADPTHVRHQTGSYGEHLTKHEDKF 377
Query: 140 ---GQHSDRVDRWRDAFTQVASYSG 161
++ +R+ +W+ A TQ A++SG
Sbjct: 378 QNNKENMERLKKWKMALTQAANFSG 452
Score = 25.8 bits (55), Expect(2) = 2e-55
Identities = 9/12 (75%), Positives = 12/12 (100%)
Frame = +2
Query: 438 EKDIFLDIACFF 449
E+++FLDIACFF
Sbjct: 1469 EQNVFLDIACFF 1504
>TC91558 weakly similar to GP|19774201|gb|AAL99077.1 NBS-LRR-Toll resistance
gene analog protein {Medicago edgeworthii}, partial
(60%)
Length = 942
Score = 210 bits (534), Expect = 2e-54
Identities = 120/306 (39%), Positives = 183/306 (59%), Gaps = 3/306 (0%)
Frame = +1
Query: 219 IGIWGMGGIGKSTIARAVYETI--RCEFELTCFLENVREISETNGLVHLQRQLLSHLSIS 276
+GIW M GIGK+T+A +Y+TI + +F+ CF+E+V +I G + +Q+++L
Sbjct: 1 VGIWRMDGIGKTTLANVLYDTISHQYQFDACCFVEDVSKIYRDGGAIAVQKRILDQTIKE 180
Query: 277 RN-DFHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQLENLVGKQDWFGPGSRVIITTRDK 335
+N + + + I N L + K+LLVLD+V++ QL+ L GSR+IITTRDK
Sbjct: 181 KNLEGYSPSEISGIISNRLYKLKLLLVLDNVDQSVQLQELHINPISLCAGSRIIITTRDK 360
Query: 336 HLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQEGYLDLSKEVVDYCGGLPLALEV 395
H+L+ + Y+ +L +DA L C KAFK D Y +L EV+ Y GLPLA+ V
Sbjct: 361 HILIEYEADIVYEAELLNDNDAHELLCRKAFKSDYSSSDYEELIPEVLKYAQGLPLAIRV 540
Query: 396 LGSYLYGRNIDVWHSAVKKLRSFPHPRVQDNLKISYDSLDTMEKDIFLDIACFFKGMKGD 455
+GS+LY R W +A++ ++ P + L+ S++ L+ EK+IFL +ACFF G + D
Sbjct: 541 MGSFLYKRKTAQWRAALEGWQNNPDSGIMKVLRSSFEGLELREKEIFLHVACFFDGERED 720
Query: 456 KVIDILESCGYFPQIGIQILIERSLITLDSVNNKLGMHDLLQEMGRDIVFQESPNDPCRR 515
V IL +CG P IGI +E+SLIT+ + N MH++LQE+G V + P++P
Sbjct: 721 YVRRILHACGLQPNIGIPTSVEKSLITIRNQENH--MHEMLQELGETNVRETHPDEP--- 885
Query: 516 SRLWSQ 521
RLWS+
Sbjct: 886 -RLWSK 900
>TC92554 similar to GP|9758876|dbj|BAB09430.1 disease resistance protein
{Arabidopsis thaliana}, partial (7%)
Length = 1043
Score = 150 bits (379), Expect(2) = 2e-54
Identities = 79/143 (55%), Positives = 100/143 (69%)
Frame = +1
Query: 3 ASSSSSCRSSSTTSLCTYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQV 62
ASSSSS +S + S +HVFL+FRGEDTR FT HL AAL K I TF DD+++ERG
Sbjct: 187 ASSSSSFGASHSNSK-QHHVFLNFRGEDTRYNFTSHLYAALCGKKIHTFMDDEEIERGDN 363
Query: 63 ISEKLINAIKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVR 122
IS L++AI+ S + I S DYASS+WCLDEL I+ECS K L V+PVFY +D S VR
Sbjct: 364 ISPTLLSAIESSKICVIIFSQDYASSSWCLDELVKIIECSEKKKLVVIPVFYHIDASHVR 543
Query: 123 HQRGCFEEAFRKHQEKFGQHSDR 145
HQRG + +AF KH++ F S++
Sbjct: 544 HQRGTYGDAFAKHEDPFQGQSNQ 612
Score = 81.3 bits (199), Expect(2) = 2e-54
Identities = 53/136 (38%), Positives = 73/136 (52%), Gaps = 2/136 (1%)
Frame = +2
Query: 138 KFGQHSDRVDRWRDAFTQVASYSGWDS-KGQHEASLVENIAQHIHRKLVPKLPSC-TENL 195
+F + +V WR A + A+ +GW S K + EA L++ I + I KL P + L
Sbjct: 590 RFRDNLTKVHMWRTALHKAANLAGWVSEKNRSEAVLIKEIVEVILDKLKCMSPHVQNKGL 769
Query: 196 VGIVSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETIRCEFELTCFLENVRE 255
VGI V V L G DV IGIWGMGGIGK+TIA AV+ + ++E F NVRE
Sbjct: 770 VGISVHVAHVESMLCSGSADVHIIGIWGMGGIGKTTIADAVFTKVSYQYERYYFAANVRE 949
Query: 256 ISETNGLVHLQRQLLS 271
+ LQ ++L+
Sbjct: 950 --TWGNRIKLQNEVLA 991
>AW736542 similar to PIR|T07656|T076 probable resistance protein - soybean
(fragment), partial (83%)
Length = 656
Score = 204 bits (519), Expect = 8e-53
Identities = 110/217 (50%), Positives = 143/217 (65%), Gaps = 1/217 (0%)
Frame = +2
Query: 315 LVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQEG 374
L G ++ GPGS +IITTRD LL GV Y+ L H++ LF AFK P E
Sbjct: 8 LCGNRNGIGPGSIIIITTRDARLLDILGVDFIYEAEGLNVHESRRLFNWHAFKEANPSEA 187
Query: 375 YLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPRVQDNLKISYDSL 434
+L LS +VV YCGGLPLALEVLGSYL+ R W S + KL+ P+ ++ + LKIS+D L
Sbjct: 188 FLILSGDVVSYCGGLPLALEVLGSYLFNRRKREWQSVISKLQKIPNDQIHEKLKISFDGL 367
Query: 435 -DTMEKDIFLDIACFFKGMKGDKVIDILESCGYFPQIGIQILIERSLITLDSVNNKLGMH 493
D MEK+IFLD+ CFF G V +IL CG IGI++LIERSL+ ++ NNKLGMH
Sbjct: 368 EDHMEKNIFLDVCCFFIGKDRAYVTEILNGCGLHADIGIEVLIERSLLKVEK-NNKLGMH 544
Query: 494 DLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTK 530
LL++MGR+IV + SP +P +R+RLW ED+ VL +
Sbjct: 545 ALLRDMGREIVRESSPEEPEKRTRLWCFEDVVDVLAE 655
>TC93494 similar to PIR|A54810|A54810 TMV resistance protein N - tobacco
(Nicotiana glutinosa), partial (6%)
Length = 876
Score = 202 bits (515), Expect = 2e-52
Identities = 115/254 (45%), Positives = 155/254 (60%), Gaps = 1/254 (0%)
Frame = +3
Query: 203 EEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETIRCEFELTCFLENVREISETNGL 262
+++ L +G ++VR +GIWGMGGIGK+T+ARA+Y + +FE C L NV + S GL
Sbjct: 12 DQIESKLKIGSHEVRVLGIWGMGGIGKTTLARALYAKMYSQFEGCCLL-NVMDESNKYGL 188
Query: 263 VHLQRQLLSHLSISRNDFHDL-YDGKKTIQNSLCRKKVLLVLDDVNELNQLENLVGKQDW 321
+ +LLS L N D Y + + RKKVL+VLD V L Q+E+L+ K D
Sbjct: 189 NVVPNKLLSSLLEEENIHPDASYIEAPFSERRIGRKKVLIVLDGVETLEQIEDLIPKIDG 368
Query: 322 FGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQEGYLDLSKE 381
GPGSRVIITTRDKH+ + Y+ L KHD+L L L AFK +P+ GY DLS+
Sbjct: 369 LGPGSRVIITTRDKHIFSQVSKCEIYELKELNKHDSLQLLSLTAFKEKQPKTGYEDLSES 548
Query: 382 VVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPRVQDNLKISYDSLDTMEKDI 441
V+ YC G PLAL+VLG+ L R + W + +KKL+ P+ ++ + LK SY LD +K I
Sbjct: 549 VIAYCKGNPLALKVLGANLSSRGQEAWENELKKLQKIPNQKIYNVLKFSYADLDRCQKAI 728
Query: 442 FLDIACFFKGMKGD 455
FLDIAC G D
Sbjct: 729 FLDIACLLSGQGKD 770
>BG645580 similar to PIR|T07656|T076 probable resistance protein - soybean
(fragment), partial (87%)
Length = 814
Score = 198 bits (503), Expect = 6e-51
Identities = 117/273 (42%), Positives = 158/273 (57%), Gaps = 8/273 (2%)
Frame = +1
Query: 356 DALVLFCLKAFKGDKPQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKL 415
D L LF AF+ P + + +LS+ VV YCGGLPLALEV+GSYLYGR W S + KL
Sbjct: 1 DPLELFSWHAFRQPSPIKNFSELSRTVVAYCGGLPLALEVIGSYLYGRTKQEWESVLLKL 180
Query: 416 RSFPHPRVQDNLKISYDSL-DTMEKDIFLDIACFFKGMKGDKVIDILESCGYFPQIGIQI 474
P+ +VQ+ L+ISYD L D M KDIFLDI CFF G V +IL CG + IGI +
Sbjct: 181 ERIPNDQVQEKLRISYDGLKDDMAKDIFLDICCFFIGKDRAYVTEILNGCGLYADIGITV 360
Query: 475 LIERSLITLDSVNNKLGMHDLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGT 534
L+ERSL+ ++ NNKLGMHDLL++MGR+IV Q S +P +RSRLW ED+ VLTKN GT
Sbjct: 361 LVERSLVKIEK-NNKLGMHDLLRDMGREIVRQSSAKNPGKRSRLWFHEDVHDVLTKNTGT 537
Query: 535 EAINSIDMKLLQ-PYE------AHWNTEAFSKTSQLKFLSLCEMQLPLGLSCLPSSLKVL 587
+ + + Q P E W + + + L +++ + P++
Sbjct: 538 INVEGVVF*VCQEPAEFASVLILSWK*NN*NN*NFYNLIVLTSLEI---MGVFPNN*DGS 708
Query: 588 HWRGCPLKTLPITTQLDELVDITLSHSKIEQLW 620
+ P +T LV + L HSKI+Q+W
Sbjct: 709 VCKDLP*TVYQMTFIRKNLVALDLKHSKIKQVW 807
>TC77730 weakly similar to GP|7341113|gb|AAF61210.1| unknown {Glycine max},
partial (80%)
Length = 831
Score = 177 bits (448), Expect(2) = 3e-49
Identities = 98/233 (42%), Positives = 143/233 (61%), Gaps = 5/233 (2%)
Frame = +1
Query: 22 VFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINAIKDSMFAITIL 81
VFL+FRG DTR FT +L AL KGI TF D+ DL+RG I+ L+ AI++S I I
Sbjct: 40 VFLNFRGSDTRNNFTGNLYKALVDKGIRTFIDENDLQRGDEITPSLVKAIEESRICIPIF 219
Query: 82 SPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQEKF-- 139
S +YASS++CLDEL I+ C + VLPVFYGVDP+DVR + E KH+E F
Sbjct: 220 SANYASSSFCLDELVHIIHCYKTKSCLVLPVFYGVDPTDVRRHTCSYGEYLTKHEEGFQN 399
Query: 140 -GQHSDRVDRWRDAFTQVASYSGWD-SKGQHEASLVENIAQHIHRKLVPKLPSCTENLVG 197
++ +R+ +W+ A TQ A+ SG+ S ++E +E I Q+I + + + VG
Sbjct: 400 NEKNMERLRQWKMALTQAANLSGYHYSPHEYEHKFIEKIVQYISNNINHDFLNVAKYPVG 579
Query: 198 IVSKVEEVNKFLGMGLND-VRFIGIWGMGGIGKSTIARAVYETIRCEFELTCF 249
+ S++E+V L MG D VR +G++G GG+GKST+A+AV+ + ++ F
Sbjct: 580 LQSRIEQVKLLLDMGSEDEVRMVGLFGTGGMGKSTLAKAVFNFLADHLKVYVF 738
Score = 37.0 bits (84), Expect(2) = 3e-49
Identities = 18/28 (64%), Positives = 19/28 (67%)
Frame = +3
Query: 244 FELTCFLENVREISETNGLVHLQRQLLS 271
FE CFL NVRE S N L HLQ+ LLS
Sbjct: 720 FEGVCFLHNVRENSSHNNLKHLQKMLLS 803
>BG646177 weakly similar to GP|18033111|g functional candidate resistance
protein KR1 {Glycine max}, partial (17%)
Length = 708
Score = 184 bits (466), Expect = 1e-46
Identities = 106/232 (45%), Positives = 142/232 (60%), Gaps = 1/232 (0%)
Frame = +3
Query: 253 VREISETNGLVHLQRQLLSHLSISRNDFHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQL 312
VREIS +GL LQ +LLS F + +G I+ L KKVLL+LDDV+EL QL
Sbjct: 3 VREISAKHGLEDLQEKLLSKTVGLSVKFGHVSEGIPIIKERLRLKKVLLILDDVDELKQL 182
Query: 313 ENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQ 372
+ L G +W G GSRV++TTRDKHLL HG+ +TY+ L K +AL L KAFK +K
Sbjct: 183 KVLAGDPNWLGHGSRVVVTTRDKHLLACHGIERTYELDGLNKEEALELLKWKAFKNNKVD 362
Query: 373 EGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPRVQDNLKISYD 432
Y + V Y GLPLALEV+GS L+G++ D W S + + PH V LK+S+D
Sbjct: 363 SSYEHILNRAVTYASGLPLALEVVGSSLFGKHKDEWKSTLDRYERIPHKEVLKILKVSFD 542
Query: 433 SLDTMEKDIFLDIACFFKGMKGDKVIDILES-CGYFPQIGIQILIERSLITL 483
SL+ E+ +FLDIAC F+G +V DIL + G + I++LIE+ LI +
Sbjct: 543 SLEKDEQSVFLDIACCFRGYILAEVEDILYAHYGECMKYHIRVLIEKCLIKI 698
>TC81052 weakly similar to GP|10178211|dbj|BAB11635. TMV resistance protein
N {Arabidopsis thaliana}, partial (8%)
Length = 656
Score = 178 bits (451), Expect = 6e-45
Identities = 93/202 (46%), Positives = 131/202 (64%), Gaps = 4/202 (1%)
Frame = +2
Query: 1 MEASSSSSCRSSSTTSLCT----YHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKD 56
++ +SSS+ SS+ +L Y VF++FRGEDTR F DHL AAL+RKGI F+DD +
Sbjct: 41 IQMASSSNSSSSALMALPRRKNYYDVFVTFRGEDTRFNFIDHLFAALQRKGIFAFRDDAN 220
Query: 57 LERGQVISEKLINAIKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGV 116
L++G+ I +LI AI+ S I +LS +Y+SSTWCL EL I++CS + VLPVFY V
Sbjct: 221 LQKGESIPPELIRAIEGSQVFIAVLSKNYSSSTWCLRELVHILDCSQVSGRRVLPVFYDV 400
Query: 117 DPSDVRHQRGCFEEAFRKHQEKFGQHSDRVDRWRDAFTQVASYSGWDSKGQHEASLVENI 176
DPS+VRHQ+G + EAF KH++ F S V WR+A TQV + SGWD + + + + ++ I
Sbjct: 401 DPSEVRHQKGIYGEAFSKHEQTFQHDSHVVQSWREALTQVGNISGWDLRDKPQYAEIKKI 580
Query: 177 AQHIHRKLVPKLPSCTENLVGI 198
+ I L S + LVG+
Sbjct: 581 VEEILNILGHNFSSLPKELVGM 646
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.137 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,972,966
Number of Sequences: 36976
Number of extensions: 327109
Number of successful extensions: 2085
Number of sequences better than 10.0: 235
Number of HSP's better than 10.0 without gapping: 1910
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1939
length of query: 640
length of database: 9,014,727
effective HSP length: 102
effective length of query: 538
effective length of database: 5,243,175
effective search space: 2820828150
effective search space used: 2820828150
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC149204.4


