

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149197.6 + phase: 0 /pseudo
(174 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
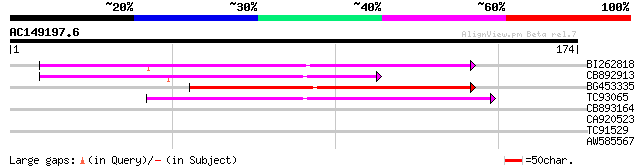
BI262818 weakly similar to GP|6642775|gb| gag-pol polyprotein {V... 79 1e-15
CB892913 similar to GP|1769897|emb lectin receptor kinase {Arabi... 69 1e-12
BG453335 weakly similar to GP|18071369|g putative gag-pol polypr... 66 8e-12
TC93065 50 5e-07
CB893164 weakly similar to GP|6016718|gb| hypothetical protein {... 26 9.3
CA920523 similar to PIR|H86313|H86 protein F2H15.10 [imported] -... 26 9.3
TC91529 similar to GP|21553452|gb|AAM62545.1 unknown {Arabidopsi... 26 9.3
AW585567 26 9.3
>BI262818 weakly similar to GP|6642775|gb| gag-pol polyprotein {Vitis
vinifera}, partial (24%)
Length = 615
Score = 78.6 bits (192), Expect = 1e-15
Identities = 44/136 (32%), Positives = 78/136 (57%), Gaps = 2/136 (1%)
Frame = +1
Query: 10 LPVFEGKNYDQWIVKMKVICRFQDVLEVVNDS--VPALASNATDMQ*TANRDAKKKDGKA 67
LPVF+G+NY W +++ D+ E V + V L+ N T Q +++ K + KA
Sbjct: 175 LPVFDGENYHIWAARIEAYLEANDLWEAVEEDYEVLPLSDNPTMAQIKNHKERKTRKSKA 354
Query: 68 MFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEE 127
+ +VS EIF +I+ +SA E W L+ Y GD +++ ++ L R++E+ M+E
Sbjct: 355 RATLFAAVSEEIFTRIMTIKSAFE-IWNFLKTEYEGDERIRGMQALNLIREFEMQKMKES 531
Query: 128 ETISQYFDKFVNLSNQ 143
ETI +Y +K ++++N+
Sbjct: 532 ETIKEYANKLISIANK 579
>CB892913 similar to GP|1769897|emb lectin receptor kinase {Arabidopsis
thaliana}, partial (35%)
Length = 837
Score = 68.6 bits (166), Expect = 1e-12
Identities = 39/107 (36%), Positives = 60/107 (55%), Gaps = 2/107 (1%)
Frame = +3
Query: 10 LPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALAS--NATDMQ*TANRDAKKKDGKA 67
+P+ +G YD +KM + DV EVV + T Q +D++K+D KA
Sbjct: 126 VPLLKGSTYDNCFIKMMALLGAHDVWEVVEKGYKESQDEDSLTKAQRDTLKDSRKRDKKA 305
Query: 68 MFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQV 114
+F ++Q++ + FEKI + SAK E WE L+ G+ K+KKVRLQ+
Sbjct: 306 LFLIYQALDEDEFEKISNATSAK-EAWEKLKTSCQGEDKVKKVRLQI 443
>BG453335 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (11%)
Length = 660
Score = 65.9 bits (159), Expect = 8e-12
Identities = 32/88 (36%), Positives = 58/88 (65%)
Frame = +1
Query: 56 ANRDAKKKDGKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVL 115
A+++ KKKD K +F + S+ F++I +KE + LEK + GD K+++++LQ L
Sbjct: 106 AHKELKKKDCKELFLIQ*SLDEGNFKRISKSTRSKEAS-NILEKYHEGDDKVEQIKLQSL 282
Query: 116 RRQYEVLTMEEEETISQYFDKFVNLSNQ 143
RR+++++ ME+E+ I+ Y K +N+ NQ
Sbjct: 283 RRKFKLMQMEDEKKIADYISKLINVVNQ 366
>TC93065
Length = 783
Score = 50.1 bits (118), Expect = 5e-07
Identities = 30/107 (28%), Positives = 57/107 (53%)
Frame = +3
Query: 43 PALASNATDMQ*TANRDAKKKDGKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYS 102
P L +N T Q + D K+G+A+ +H ++ ++IF KI++ E AK E W L + Y
Sbjct: 3 PPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIKILNLEIAK-EAWSKLMEEYQ 179
Query: 103 GDGKLKKVRLQVLRRQYEVLTMEEEETISQYFDKFVNLSNQSQEW*K 149
G + KK+++ L + ++ + + + +F+ L ++S W K
Sbjct: 180 GSERTKKMKVLNLEESLKH*K*KKLKLLESFLTEFLKLLHKSDCWEK 320
>CB893164 weakly similar to GP|6016718|gb| hypothetical protein {Arabidopsis
thaliana}, partial (45%)
Length = 818
Score = 25.8 bits (55), Expect = 9.3
Identities = 20/88 (22%), Positives = 41/88 (45%), Gaps = 11/88 (12%)
Frame = +3
Query: 57 NRDAKKKDGKAMFFMHQSV----SNEIFEKIVHYESAKEE--TWEALEK----LYSGDGK 106
N KKDG ++ F H+++ S + + +H+ + E+ W K + G+ K
Sbjct: 372 NLPGAKKDGSSVHFFHRTMNAYSSGQRKRRKIHHRGSTEDHVRWHKTGKTKAIIEHGEHK 551
Query: 107 -LKKVRLQVLRRQYEVLTMEEEETISQY 133
KK+ + +R + E + + + + QY
Sbjct: 552 GFKKIMVLYIRSEKESKSYKSDWKMHQY 635
>CA920523 similar to PIR|H86313|H86 protein F2H15.10 [imported] - Arabidopsis
thaliana, partial (25%)
Length = 714
Score = 25.8 bits (55), Expect = 9.3
Identities = 10/20 (50%), Positives = 16/20 (80%)
Frame = +3
Query: 117 RQYEVLTMEEEETISQYFDK 136
RQY+ +TM ++ T+SQ FD+
Sbjct: 450 RQYQAITMYQKITMSQCFDQ 509
>TC91529 similar to GP|21553452|gb|AAM62545.1 unknown {Arabidopsis
thaliana}, partial (63%)
Length = 833
Score = 25.8 bits (55), Expect = 9.3
Identities = 21/72 (29%), Positives = 34/72 (47%), Gaps = 3/72 (4%)
Frame = +2
Query: 85 HYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEEETISQYFDKFVNLSNQ- 143
H E EET EA LY+G GK +K +L+ R + T+ + +K++ L +
Sbjct: 461 HQEEE*EETHEAAPCLYTG-GKRRKGKLKCKR-------LSGRRTLRR*SEKWLQLQGKE 616
Query: 144 --SQEW*KCYRL 153
++W C L
Sbjct: 617 SGQKDWLNCSAL 652
>AW585567
Length = 491
Score = 25.8 bits (55), Expect = 9.3
Identities = 22/84 (26%), Positives = 36/84 (42%)
Frame = +3
Query: 69 FFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEEE 128
FF S SN+I SA + D +L+K R ++ + T EEEE
Sbjct: 234 FFCKSSSSNQILRN----SSANSSPLATNSSTITKDPELEKKR-----KRNRI*TGEEEE 386
Query: 129 TISQYFDKFVNLSNQSQEW*KCYR 152
+ + +F + + S+ KC+R
Sbjct: 387 GTTPIYFRFQSKKSTSRTRFKCFR 458
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.332 0.140 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,622,705
Number of Sequences: 36976
Number of extensions: 46104
Number of successful extensions: 425
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 420
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 421
length of query: 174
length of database: 9,014,727
effective HSP length: 89
effective length of query: 85
effective length of database: 5,723,863
effective search space: 486528355
effective search space used: 486528355
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 55 (25.8 bits)
Medicago: description of AC149197.6


