

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149135.13 - phase: 0
(316 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
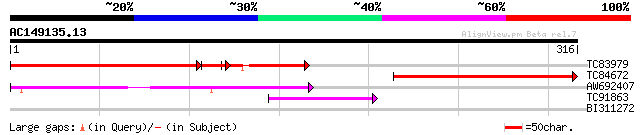
TC83979 homologue to PIR|T45963|T45963 cell division-like protei... 215 2e-75
TC84672 similar to GP|15292839|gb|AAK92790.1 putative cell divis... 216 9e-57
AW692407 similar to GP|17381234|gb AT4g25730/F14M19_10 {Arabidop... 96 2e-20
TC91863 similar to GP|1246842|emb|CAA63360.1 G2830 {Saccharomyce... 42 2e-04
BI311272 similar to GP|19550749|gb caffeic acid O-methyltransfer... 28 3.5
>TC83979 homologue to PIR|T45963|T45963 cell division-like protein -
Arabidopsis thaliana, partial (54%)
Length = 669
Score = 215 bits (547), Expect(3) = 2e-75
Identities = 107/107 (100%), Positives = 107/107 (100%)
Frame = +3
Query: 1 MGKASRDKRDIYYRKAKEEGWRARSAFKLLQIDEEFNIFEGVKRVVDLCAAPGSWSQVLS 60
MGKASRDKRDIYYRKAKEEGWRARSAFKLLQIDEEFNIFEGVKRVVDLCAAPGSWSQVLS
Sbjct: 162 MGKASRDKRDIYYRKAKEEGWRARSAFKLLQIDEEFNIFEGVKRVVDLCAAPGSWSQVLS 341
Query: 61 RKLYLPAKLAPDAKDENLPLIVAIDLQPMAPIEGVIQVQGDITNART 107
RKLYLPAKLAPDAKDENLPLIVAIDLQPMAPIEGVIQVQGDITNART
Sbjct: 342 RKLYLPAKLAPDAKDENLPLIVAIDLQPMAPIEGVIQVQGDITNART 482
Score = 69.3 bits (168), Expect(3) = 2e-75
Identities = 39/53 (73%), Positives = 41/53 (76%), Gaps = 4/53 (7%)
Frame = +2
Query: 119 KANLVVCDGA----PDVTGLHDMDEFVQSQLILAGLTIVTHVLKEGGKFIAKI 167
+ L VCDGA PD LHDMDEFVQSQLILAG+TIVTHVLKEG KFIA I
Sbjct: 518 RPTLSVCDGAS*CYPD---LHDMDEFVQSQLILAGVTIVTHVLKEGXKFIAXI 667
Score = 37.0 bits (84), Expect(3) = 2e-75
Identities = 16/16 (100%), Positives = 16/16 (100%)
Frame = +1
Query: 108 AEVVIRHFDGCKANLV 123
AEVVIRHFDGCKANLV
Sbjct: 484 AEVVIRHFDGCKANLV 531
>TC84672 similar to GP|15292839|gb|AAK92790.1 putative cell division protein
{Arabidopsis thaliana}, partial (52%)
Length = 596
Score = 216 bits (550), Expect = 9e-57
Identities = 102/102 (100%), Positives = 102/102 (100%)
Frame = +2
Query: 215 FNPKDLHRLLEKVGSPSGVDDTDCVSGWLEGPNKVYIPFLACGDLTGYDSDRSYPLPKVA 274
FNPKDLHRLLEKVGSPSGVDDTDCVSGWLEGPNKVYIPFLACGDLTGYDSDRSYPLPKVA
Sbjct: 2 FNPKDLHRLLEKVGSPSGVDDTDCVSGWLEGPNKVYIPFLACGDLTGYDSDRSYPLPKVA 181
Query: 275 GGTYQSLDPVQPPIAPPYKRALELKKASPQGFRELENLSLDS 316
GGTYQSLDPVQPPIAPPYKRALELKKASPQGFRELENLSLDS
Sbjct: 182 GGTYQSLDPVQPPIAPPYKRALELKKASPQGFRELENLSLDS 307
>AW692407 similar to GP|17381234|gb AT4g25730/F14M19_10 {Arabidopsis
thaliana}, partial (16%)
Length = 559
Score = 95.5 bits (236), Expect = 2e-20
Identities = 58/175 (33%), Positives = 97/175 (55%), Gaps = 6/175 (3%)
Frame = +1
Query: 1 MGKAS---RDKRDIYYRKAKEEGWRARSAFKLLQIDEEFNIFEGVKRVVDLCAAPGSWSQ 57
MGKA + + D YY AKE G+R+R+++KL+QI+ +F+ E + V+DLCAAPG W Q
Sbjct: 55 MGKAKAKGKHRLDKYYYLAKEHGYRSRASWKLVQINSKFHFLESSRSVLDLCAAPGGWMQ 234
Query: 58 VLSRKLYLPAKLAPDAKDENLPLIVAIDLQPMAPIEGVIQVQGDITNARTAEVV--IRHF 115
V +++ + L++ +DL P+ PI G I +Q DIT V I +
Sbjct: 235 VAVQRVPVD------------HLVIGVDLTPIKPIRGAIAIQEDITRPECKSRVRKIMNE 378
Query: 116 DGCKA-NLVVCDGAPDVTGLHDMDEFVQSQLILAGLTIVTHVLKEGGKFIAKIFR 169
+G +A ++++ DG+P+V G + Q+ L++ + + T L G F + F+
Sbjct: 379 NGYRAFDVILHDGSPNVGGAWAQEATSQNSLVIDAIKLATQFLAPKGTFCYQGFQ 543
>TC91863 similar to GP|1246842|emb|CAA63360.1 G2830 {Saccharomyces
cerevisiae}, partial (13%)
Length = 1331
Score = 42.4 bits (98), Expect = 2e-04
Identities = 23/61 (37%), Positives = 32/61 (51%)
Frame = +1
Query: 145 LILAGLTIVTHVLKEGGKFIAKIFRGKDTSLLYCQLKLFFPVVTFAKPKSSRNSSIEAFA 204
L A L + L+ GG F+ K ++G + +LK F V KP+SSR+ S EAF
Sbjct: 1036 LCAAALQFASDTLRHGGHFVCKFYQGAEDRAFEKKLKTLFARVLREKPESSRSDSKEAFF 1215
Query: 205 V 205
V
Sbjct: 1216 V 1218
Score = 40.4 bits (93), Expect = 0.001
Identities = 43/169 (25%), Positives = 63/169 (36%), Gaps = 60/169 (35%)
Frame = +1
Query: 6 RDKRDIYYRKAKEEGWRARSAFKL---------------------------LQIDEEFNI 38
R +DI+ R+AK +G R+R+AFKL LQID ++ I
Sbjct: 139 RQGKDIFAREAKVQGLRSRAAFKLLEVCEKKTPSRWCSFF*LPETTRC**PLQIDAKYRI 318
Query: 39 FEGVKRVVDLCA-------------------APGSWSQVLSRKLYLPAKLAPDAKDENL- 78
F + VVDL APGSWSQV + L+ D +
Sbjct: 319 FRKGQTVVDLVILLSLPIPLVNVR*DPFQGYAPGSWSQVGGFHNVPESPLSR*ESDREV* 498
Query: 79 -------------PLIVAIDLQPMAPIEGVIQVQGDITNARTAEVVIRH 114
++ ID+ P P +GV +QG+ + +V H
Sbjct: 499 RKY*VAADRTKPSGQVLGIDIIPAQPPKGVSTIQGNFLSPDVQVLVKDH 645
>BI311272 similar to GP|19550749|gb caffeic acid O-methyltransferase II
{Nicotiana tabacum}, partial (58%)
Length = 805
Score = 28.5 bits (62), Expect = 3.5
Identities = 23/69 (33%), Positives = 32/69 (46%)
Frame = +2
Query: 34 EEFNIFEGVKRVVDLCAAPGSWSQVLSRKLYLPAKLAPDAKDENLPLIVAIDLQPMAPIE 93
E +N FEG+KR+VD+ G Q+++ K P+ N L I P P
Sbjct: 176 ESYNGFEGIKRLVDVGGGLGVNIQLVTSKY-------PNIHGINFDLPHVIQHAPSYP-- 328
Query: 94 GVIQVQGDI 102
GV V GD+
Sbjct: 329 GVEHVGGDM 355
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.139 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,588,381
Number of Sequences: 36976
Number of extensions: 139549
Number of successful extensions: 587
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 582
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 584
length of query: 316
length of database: 9,014,727
effective HSP length: 96
effective length of query: 220
effective length of database: 5,465,031
effective search space: 1202306820
effective search space used: 1202306820
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC149135.13


