

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149134.13 - phase: 0
(163 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
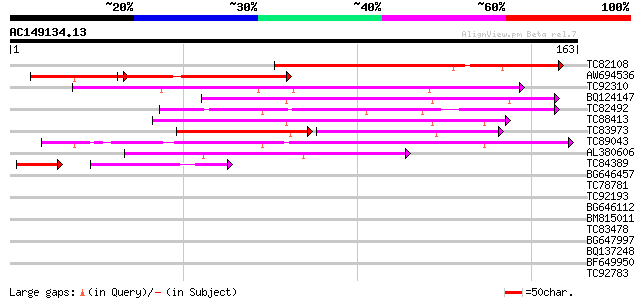
Sequences producing significant alignments: (bits) Value
TC82108 similar to GP|21537320|gb|AAM61661.1 unknown {Arabidopsi... 77 3e-15
AW694536 57 3e-11
TC92310 weakly similar to PIR|T47778|T47778 hypothetical protein... 58 2e-09
BQ124147 similar to GP|10177836|dbj emb|CAB62440.1~gene_id:MCD7.... 53 6e-08
TC82492 similar to GP|14596145|gb|AAK68800.1 Unknown protein {Ar... 49 9e-07
TC88413 weakly similar to GP|10176786|dbj|BAB09900. gene_id:MIK1... 45 1e-05
TC83973 GP|21537320|gb|AAM61661.1 unknown {Arabidopsis thaliana}... 35 2e-05
TC89043 similar to GP|10178241|dbj|BAB11673. gene_id:MDJ22.9~unk... 43 6e-05
AL380606 40 3e-04
TC84389 weakly similar to GP|10177368|dbj|BAB10659. contains sim... 32 5e-04
BG646457 similar to GP|3420058|gb|A unknown protein {Arabidopsis... 39 0.001
TC78781 weakly similar to PIR|T02291|T02291 hypothetical protein... 37 0.004
TC92193 weakly similar to PIR|A96688|A96688 hypothetical protein... 37 0.005
BG646112 weakly similar to GP|10177368|dbj contains similarity t... 33 0.066
BM815011 similar to PIR|T49178|T491 hypothetical protein T20N10.... 32 0.086
TC83478 similar to PIR|H86304|H86304 hypothetical protein AAF998... 30 0.56
BG647997 29 0.73
BQ137248 29 0.73
BF649950 similar to GP|9802755|gb|A Unknown protein {Arabidopsis... 28 1.6
TC92783 similar to PIR|T49020|T49020 hypothetical protein F3C22.... 28 2.1
>TC82108 similar to GP|21537320|gb|AAM61661.1 unknown {Arabidopsis
thaliana}, partial (7%)
Length = 817
Score = 77.0 bits (188), Expect = 3e-15
Identities = 47/91 (51%), Positives = 59/91 (64%), Gaps = 8/91 (8%)
Frame = +2
Query: 77 EVPFVNLPSLKALYLDVVTFTYYELILKLLSGCPILQYLGTNNLVVELPY-----SERPV 131
E P V LPSLK L+L+ V FTYY I KLLSGCPILQ L N+L +++P S +P
Sbjct: 2 EGPLVILPSLKILHLESVCFTYYGHIRKLLSGCPILQELEANDLTIKIPCRVFLGSSQPP 181
Query: 132 ISLSNLIRA---NICDIHIEFDWLQNVERLR 159
SLSNL+RA NIC +H D L+N + +R
Sbjct: 182 -SLSNLVRANISNICPMHNLLDRLRNAQHMR 271
>AW694536
Length = 226
Score = 57.0 bits (136), Expect(2) = 3e-11
Identities = 27/50 (54%), Positives = 39/50 (78%)
Frame = +3
Query: 32 IAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILKLKQITLNEVPFV 81
+ + GVENL+I+ H ++ TLPSF+ +SKTL+ LKLK++TLNEVP+V
Sbjct: 81 LQLHSGVENLSINLCH--LNETTLPSFILTSKTLTSLKLKRVTLNEVPYV 224
Score = 26.9 bits (58), Expect(2) = 3e-11
Identities = 14/29 (48%), Positives = 18/29 (61%), Gaps = 1/29 (3%)
Frame = +2
Query: 7 TLPILSFHLQC-WNDYDCRDIYNFLYIAI 34
+LPILSF L C ++ D D N +Y AI
Sbjct: 2 SLPILSFQLNCRYSFIDKNDFNNLVYAAI 88
>TC92310 weakly similar to PIR|T47778|T47778 hypothetical protein F17J16.10
- Arabidopsis thaliana, partial (8%)
Length = 1019
Score = 57.8 bits (138), Expect = 2e-09
Identities = 44/149 (29%), Positives = 72/149 (47%), Gaps = 19/149 (12%)
Frame = +2
Query: 19 NDYDCRDIYNFLYIAIQRGVENLN----------IDFSHSLFSQMTLPSFVFSSKTLSIL 68
N + + N+LY+ +R V+ LN +D H LPS +F+ KTL +L
Sbjct: 308 NYFSILRLNNWLYLLGKRNVKYLNLHLNFLNALLVDVQHRKTLTPKLPSTIFTCKTLVVL 487
Query: 69 KL-----KQITLNEVPF-VNLPSLKALYLDVVTFTYYELILKLLSGCPILQYLGTNN--- 119
+ K + + V F PSLK L+ + + F + L LL+GCP+L+ +
Sbjct: 488 NISWFAFKGFSFSSVGFGFGFPSLKTLHFNHIYFNHLSDFLLLLAGCPVLEDFKACHVFT 667
Query: 120 LVVELPYSERPVISLSNLIRANICDIHIE 148
L E ++ ++LSNLIR +I D + +
Sbjct: 668 LEEEEEATQESTLNLSNLIRTDIIDTNFD 754
>BQ124147 similar to GP|10177836|dbj emb|CAB62440.1~gene_id:MCD7.13~similar
to unknown protein {Arabidopsis thaliana}, partial (11%)
Length = 677
Score = 52.8 bits (125), Expect = 6e-08
Identities = 39/109 (35%), Positives = 59/109 (53%), Gaps = 6/109 (5%)
Frame = +1
Query: 56 PSFVFSSKTLSILKLKQITLNEV---PFVNLPSLKALYLDVVTFTYYELILKLLSGCPIL 112
P+ +F+ +TL ILKL ++ + + LPSLK L+L + F Y + LL GCP+L
Sbjct: 259 PNTIFTRETLVILKLSELFMGSGFCNYLIRLPSLKTLHLKDIEFQQYGDLDCLLEGCPVL 438
Query: 113 QYLGTNNL-VVELPYSERPVISLSNLIRANI--CDIHIEFDWLQNVERL 158
+ L ++ V L +S +L+ L RA+I CD + L NVE L
Sbjct: 439 EDLQLYDISYVSLLFSNACRKTLTKLNRADIIQCDCGVRMKALSNVEFL 585
>TC82492 similar to GP|14596145|gb|AAK68800.1 Unknown protein {Arabidopsis
thaliana}, partial (7%)
Length = 1286
Score = 48.9 bits (115), Expect = 9e-07
Identities = 45/130 (34%), Positives = 67/130 (50%), Gaps = 15/130 (11%)
Frame = +3
Query: 44 DFSHSLFSQMTLPSFVFSSKTLSILKLK--QITLNEVPFVNLPSLKALYLDVVTFTYYEL 101
+F ++ + P +F+S+TL ILKL+ +I + + F +LPSLK L L+ F +
Sbjct: 312 EFHLGMYGHILTP-IIFTSRTLVILKLQNLKIVADNLCF-DLPSLKTLVLEREYFKNQKN 485
Query: 102 -ILKLLSGCPILQYLGT------------NNLVVELPYSERPVISLSNLIRANICDIHIE 148
+KLL+ CPIL+ L T NN V E + LS L+RA I + +
Sbjct: 486 DFVKLLNACPILEDLHTYYPRLRIMKRKDNNEVQEF-----KSLFLSKLVRAYIHSMDVP 650
Query: 149 FDWLQNVERL 158
FD + NVE L
Sbjct: 651 FDAINNVEYL 680
>TC88413 weakly similar to GP|10176786|dbj|BAB09900.
gene_id:MIK19.29~pir||T01344~similar to unknown protein
{Arabidopsis thaliana}, partial (10%)
Length = 1288
Score = 45.1 bits (105), Expect = 1e-05
Identities = 35/109 (32%), Positives = 59/109 (54%), Gaps = 6/109 (5%)
Frame = +1
Query: 42 NIDFSHSLFSQMTLPSFVFSSKTLSILKLKQITLNEV--PFVNLPSLKALYLDVVTFTYY 99
++D + S + +P V +S L +LKL +T++ V NLPSLK L+L V F
Sbjct: 556 HLDVTSSATLCLCVPYKVLTSTALVVLKLNALTIDYVHRSSTNLPSLKILHLTQVHFLKL 735
Query: 100 ELILKLLSGCPILQYLGTNNL-VVELPYSERPVISLS---NLIRANICD 144
+ ++K+LS P+L+ L +L V + ++ +L L+RA+I D
Sbjct: 736 KFLIKILSMSPLLEDLLLKDLQVTDNTLAQDDAAALKPFPKLLRADISD 882
>TC83973 GP|21537320|gb|AAM61661.1 unknown {Arabidopsis thaliana}, partial
(2%)
Length = 870
Score = 34.7 bits (78), Expect(2) = 2e-05
Identities = 24/61 (39%), Positives = 32/61 (52%), Gaps = 7/61 (11%)
Frame = +2
Query: 89 LYLDVVTFTYYELILKLLSGCPILQYLGTNNLV-------VELPYSERPVISLSNLIRAN 141
L+L V E I++LLSGCPIL+ L +L+ V L + + SL LI AN
Sbjct: 134 LHLFGVLLERAEYIIQLLSGCPILEELHAESLIVRNKEWLVSLNFVRQKFPSLPKLITAN 313
Query: 142 I 142
I
Sbjct: 314 I 316
Score = 29.3 bits (64), Expect(2) = 2e-05
Identities = 17/41 (41%), Positives = 25/41 (60%), Gaps = 2/41 (4%)
Frame = +1
Query: 49 LFSQMTLPSFVFSSKTLSILKLKQITLNEVP--FVNLPSLK 87
L +TLP VFS +TL +L L+ IT+ ++ V+ P LK
Sbjct: 10 LSGSITLPRSVFSCRTLVVLHLQWITVKDLSQLVVDFPLLK 132
>TC89043 similar to GP|10178241|dbj|BAB11673. gene_id:MDJ22.9~unknown
protein {Arabidopsis thaliana}, partial (9%)
Length = 1413
Score = 42.7 bits (99), Expect = 6e-05
Identities = 50/171 (29%), Positives = 75/171 (43%), Gaps = 18/171 (10%)
Frame = +2
Query: 10 ILSFHLQC----WNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTL 65
I + HL C W ++ D++ + A + VEN N+ + + L +F +L
Sbjct: 278 IKTLHLNCGSPNWKHFNL-DLW--IGTAKRHPVENFNLV---GTWRSIPLRPSIFRFPSL 439
Query: 66 SILKLK--QITLNEVPFVNLPSLKALYLDVVTFTYYELILKLLSGCPILQYLGTNNLVVE 123
+LKLK +I + + V+LP LK L+LD V K+L GCP+L+ L N E
Sbjct: 440 VVLKLKTLKIIVGNIT-VDLPLLKILHLDRVYLKNKTNFNKILYGCPVLEDLIANIYYKE 616
Query: 124 LPYSERPVISLS------------NLIRANICDIHIEFDWLQNVERLRATV 162
V +LS LIR I + F + NVE L T+
Sbjct: 617 PTPEPDEVFTLSKATATGEFKILPKLIRVQINADEVPFRAIHNVEFLALTM 769
>AL380606
Length = 437
Score = 40.4 bits (93), Expect = 3e-04
Identities = 29/93 (31%), Positives = 47/93 (50%), Gaps = 11/93 (11%)
Frame = +1
Query: 34 IQRGVENLNIDFSHSLFSQMT-------LPSFVFSSKTLSILKLKQITLNEVPFVNL--- 83
+QRG++ L++ L + LP +F+ KTL L L ++ F ++
Sbjct: 97 VQRGLKYLHLRLRVPLPDHFSGYPYFPKLPISIFTCKTLVSLNLSWFRVDGFSFTSVGFE 276
Query: 84 -PSLKALYLDVVTFTYYELILKLLSGCPILQYL 115
PSLK L L ++ F+ + LL+GCPIL+ L
Sbjct: 277 FPSLKTLSLRLIKFSEVRDFMLLLAGCPILEDL 375
>TC84389 weakly similar to GP|10177368|dbj|BAB10659. contains similarity to
heat shock transcription factor HSF30~gene_id:K16L22.13,
partial (8%)
Length = 692
Score = 32.3 bits (72), Expect(2) = 5e-04
Identities = 18/41 (43%), Positives = 24/41 (57%)
Frame = +1
Query: 24 RDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKT 64
R+I + QRG+ENLN++ S S+ TLP VFS T
Sbjct: 532 RNINRIGHAIAQRGIENLNLELSXSI----TLPRXVFSCXT 642
Score = 26.6 bits (57), Expect(2) = 5e-04
Identities = 11/13 (84%), Positives = 12/13 (91%)
Frame = +3
Query: 3 MRDITLPILSFHL 15
+RDITLPI SFHL
Sbjct: 462 LRDITLPIRSFHL 500
>BG646457 similar to GP|3420058|gb|A unknown protein {Arabidopsis thaliana},
partial (3%)
Length = 744
Score = 38.9 bits (89), Expect = 0.001
Identities = 32/92 (34%), Positives = 52/92 (55%), Gaps = 9/92 (9%)
Frame = +3
Query: 33 AIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILKLK---QITLNEVPFVNLPSLKAL 89
A+ V+ L++D L ++ LP + + ++L +LKL+ Q TLN V LPSLK L
Sbjct: 111 ALDLKVQELDLDLF--LHEKVLLPLQLSTCESLVVLKLRGRIQPTLNSSFHVYLPSLKIL 284
Query: 90 YL-DVVTFTYYE-----LILKLLSGCPILQYL 115
+L + VT++ ++ + LSGCP L+ L
Sbjct: 285 HLRETVTYSMFDDGMEYDLNNFLSGCPHLEEL 380
>TC78781 weakly similar to PIR|T02291|T02291 hypothetical protein T13D8.28 -
Arabidopsis thaliana, partial (12%)
Length = 1300
Score = 37.0 bits (84), Expect = 0.004
Identities = 34/99 (34%), Positives = 50/99 (50%), Gaps = 5/99 (5%)
Frame = +3
Query: 36 RGVENLNIDF-SHSLFS--QMTLPSFVFSSKTLSILKLKQITLNEVPFV-NLPSLKALYL 91
R + LN+ S S++ M L VF+S+TL LKL ++ + E V +LPSLK L L
Sbjct: 393 RNIVRLNLSLNSLSIYGILNMYLTRPVFNSRTLVELKLHRVCIAETLSVASLPSLKVLSL 572
Query: 92 DVVTFTYYELILKLLSGCP-ILQYLGTNNLVVELPYSER 129
V F + L LLS +L+ L ++ P +R
Sbjct: 573 SRVQFDTKSVFLSLLSAVSRVLEELRISSPTFSAPICDR 689
>TC92193 weakly similar to PIR|A96688|A96688 hypothetical protein T27F4.4
[imported] - Arabidopsis thaliana, partial (9%)
Length = 595
Score = 36.6 bits (83), Expect = 0.005
Identities = 30/90 (33%), Positives = 43/90 (47%), Gaps = 1/90 (1%)
Frame = +2
Query: 31 YIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILKLK-QITLNEVPFVNLPSLKAL 89
Y+ + G N N HS P+F ++L+ L+L Q TL+ + P LK L
Sbjct: 293 YLDLSLGDRNDNFVLPHSF------PAF----ESLNELRLGLQFTLHIPSGICFPILKTL 442
Query: 90 YLDVVTFTYYELILKLLSGCPILQYLGTNN 119
+ VTF +L SGCP+LQ L +N
Sbjct: 443 VVTNVTFANENSAQQLFSGCPVLQELELDN 532
>BG646112 weakly similar to GP|10177368|dbj contains similarity to heat shock
transcription factor HSF30~gene_id:K16L22.13, partial
(5%)
Length = 742
Score = 32.7 bits (73), Expect = 0.066
Identities = 34/126 (26%), Positives = 57/126 (44%), Gaps = 18/126 (14%)
Frame = +2
Query: 10 ILSFHLQCWND----YDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTL--PSFVFSSK 63
I FHL C + +Y ++ AI +E+L+++ ++ M PS + +
Sbjct: 248 IRKFHLTCCTSDVYSFPGECVYMWVRAAIGPHLEDLSLNITNCHGDDMVYLPPSLLNCTN 427
Query: 64 TLSILKLKQITLNEVPF-VNLPSLKALY---------LDVVTFTYY--ELILKLLSGCPI 111
+S+ I L P ++ PSLK L +D+ F + + IL LSGCP+
Sbjct: 428 LVSLSLFGLIHLKFQPSAIHFPSLKMLKVEFSILEHNIDIEDFGKHLTDSILVFLSGCPV 607
Query: 112 LQYLGT 117
L+ L T
Sbjct: 608 LETLDT 625
>BM815011 similar to PIR|T49178|T491 hypothetical protein T20N10.300 -
Arabidopsis thaliana, partial (4%)
Length = 628
Score = 32.3 bits (72), Expect = 0.086
Identities = 31/101 (30%), Positives = 49/101 (47%), Gaps = 3/101 (2%)
Frame = +2
Query: 18 WNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLP-SFVFSSKTLSILKL--KQIT 74
+ +Y+ I +++ AI +E L++D + + LP SF S L L+L +
Sbjct: 248 FREYEVECIEMWVHAAIGPRLEELDLDIYEA---DINLPLSFFTSCNNLVSLRLGGEYNM 418
Query: 75 LNEVPFVNLPSLKALYLDVVTFTYYELILKLLSGCPILQYL 115
+ V PSLK +YL + + I+ LSGCP LQ L
Sbjct: 419 KFKHSLVQFPSLKKVYLSTMDS---DSIVAFLSGCPKLQDL 532
>TC83478 similar to PIR|H86304|H86304 hypothetical protein AAF99838.1
[imported] - Arabidopsis thaliana, partial (3%)
Length = 906
Score = 29.6 bits (65), Expect = 0.56
Identities = 23/82 (28%), Positives = 44/82 (53%), Gaps = 1/82 (1%)
Frame = +2
Query: 35 QRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILKLKQITLNEVP-FVNLPSLKALYLDV 93
++GV+ + + + S+ +PS FS + L+ +++ + L P F SL + +
Sbjct: 5 RKGVKFIQLASNESV--PYRVPSHFFSFQKLTHVRICKFKLLVPPNFCGFKSLVHFHFER 178
Query: 94 VTFTYYELILKLLSGCPILQYL 115
+TF + L L+SGCP+L+ L
Sbjct: 179 MTFEFGALE-SLISGCPLLEEL 241
>BG647997
Length = 790
Score = 29.3 bits (64), Expect = 0.73
Identities = 37/150 (24%), Positives = 60/150 (39%), Gaps = 15/150 (10%)
Frame = +3
Query: 10 ILSFHLQCWNDYDCRD-IYNFLYIAIQRGVENLNIDFS------------HSLFSQMTLP 56
I F L N C + I + QRGV++L +DFS H+LF LP
Sbjct: 285 INKFSLTVSNPQTCGETIERCVAFVTQRGVKDLTLDFSNPKWNENDLDDKHALFQ---LP 455
Query: 57 SFVFS-SKTLSILKLKQITLNEVPFVNLPSLKALYLDVVTFTYYELILKLLSGCPILQYL 115
+ V+ L LKL + + +N +LK + L + + LLS C ++ L
Sbjct: 456 THVYQLGSLLKSLKLYSCSFDMPDILNFGALKDVSLGWIEVP-MNTLKTLLSTCKTIENL 632
Query: 116 GTNNLVVELPYSERPVISLS-NLIRANICD 144
+ + + ++ L + N CD
Sbjct: 633 SLKRCWNLVDFDLKDIVQLGLKRMVLNKCD 722
>BQ137248
Length = 977
Score = 29.3 bits (64), Expect = 0.73
Identities = 13/33 (39%), Positives = 21/33 (63%)
Frame = -1
Query: 85 SLKALYLDVVTFTYYELILKLLSGCPILQYLGT 117
S ALY+ V F + ++ L+LL+GC + Y+ T
Sbjct: 917 SCAALYISVRRFDFIDVSLRLLAGCVAILYVYT 819
>BF649950 similar to GP|9802755|gb|A Unknown protein {Arabidopsis thaliana},
partial (7%)
Length = 581
Score = 28.1 bits (61), Expect = 1.6
Identities = 26/93 (27%), Positives = 46/93 (48%), Gaps = 2/93 (2%)
Frame = +2
Query: 25 DIYNFLYIAIQRGVENLNIDF-SHSLFSQMTLPSFVFSSKTLSILKLKQITLNEVP-FVN 82
D+ ++ ++ ++ L +D + L+ +P +FS ++L LKL L F
Sbjct: 221 DVDQWILYLSRKSIKELVLDVCTEELYK---IPWCLFSCQSLHHLKLHFCCLKPPTLFKG 391
Query: 83 LPSLKALYLDVVTFTYYELILKLLSGCPILQYL 115
SLK+L L+ V + L+SGCP+L+ L
Sbjct: 392 FRSLKSLDLNHVAVAQ-DAFENLISGCPLLEKL 487
>TC92783 similar to PIR|T49020|T49020 hypothetical protein F3C22.70 -
Arabidopsis thaliana (fragment), partial (7%)
Length = 679
Score = 27.7 bits (60), Expect = 2.1
Identities = 18/35 (51%), Positives = 22/35 (62%), Gaps = 2/35 (5%)
Frame = +1
Query: 83 LPSLKALYLDVVTFTYYEL--ILKLLSGCPILQYL 115
LPSLK L LD+ Y E+ + LLS CPIL+ L
Sbjct: 574 LPSLKKLQLDI---GYVEMNSMDNLLSSCPILETL 669
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.327 0.142 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,550,584
Number of Sequences: 36976
Number of extensions: 77275
Number of successful extensions: 727
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 724
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 724
length of query: 163
length of database: 9,014,727
effective HSP length: 89
effective length of query: 74
effective length of database: 5,723,863
effective search space: 423565862
effective search space used: 423565862
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 54 (25.4 bits)
Medicago: description of AC149134.13


