

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
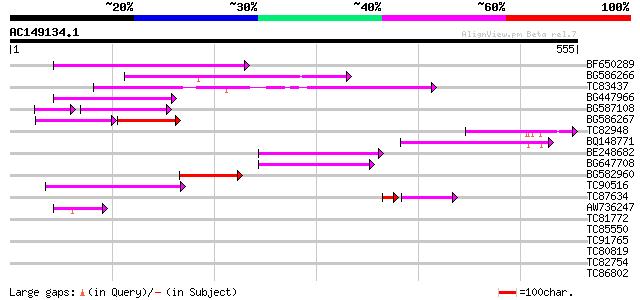
Query= AC149134.1 - phase: 0
(555 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgari... 144 7e-35
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 141 7e-34
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 112 5e-25
BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [impo... 100 1e-21
BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - ... 72 2e-16
BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At... 55 8e-16
TC82948 69 4e-12
BQ148771 66 4e-11
BE248682 similar to GP|18568269|gb putative gag-pol polyprotein ... 61 1e-09
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 58 8e-09
BG582960 54 2e-07
TC90516 48 9e-06
TC87634 40 1e-04
AW736247 weakly similar to PIR|T14619|T146 reverse transcriptase... 44 2e-04
TC81772 37 0.020
TC85550 homologue to SP|P28002|COMT_MEDSA Caffeic acid 3-O-methy... 34 0.17
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 29 0.40
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 32 0.49
TC82754 homologue to GP|18181865|emb|CAD20127. raffinose synthas... 31 1.4
TC86802 similar to GP|7939577|dbj|BAA95778.1 ATP-dependent RNA h... 30 1.9
>BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgaris}, partial
(69%)
Length = 616
Score = 144 bits (364), Expect = 7e-35
Identities = 72/191 (37%), Positives = 108/191 (55%)
Frame = +3
Query: 44 TKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFFKNGWLPNNFNANS 103
T EVKNA+F ++S APG D + FF+ WNI+ V +A+LDFFK G++P N
Sbjct: 27 TAVEVKNALFSMDSSKAPGIDGYNVHFFKCSWNIIGDSVIDAILDFFKTGFMPKIINCTY 206
Query: 104 IILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQRGFVQGRNIRD 163
+ L+PK N SV +R IA + +KII+K+L R+ +L S++S+ Q FV+GR I D
Sbjct: 207 VTLLPKEVNVTSVKNFRPIACCSVIYKIISKILTSRMQGVLNSVVSENQSAFVKGRVIFD 386
Query: 164 CIALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFNELFCNWIKTIL 223
I L+ E + K +KID+ KA+D+ W F+ ++ GF F NW+ L
Sbjct: 387 NIILSHELVKSYSRKGISPRCMVKIDLXKAYDSXEWPFIKHLMLELGFPYKFVNWVMAXL 566
Query: 224 HSSKMFISMNG 234
++ + NG
Sbjct: 567 TTASYTFNXNG 599
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 141 bits (355), Expect = 7e-34
Identities = 86/225 (38%), Positives = 125/225 (55%), Gaps = 3/225 (1%)
Frame = -3
Query: 113 ADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQRGFVQGRNIRDCIALTSEAI 172
A V +YRTIA N ++KII K+L+ R+ +L SIIS Q FV GR I D + +T + +
Sbjct: 787 AKRVSEYRTIAPCNTQYKIIAKILSKRMQPLLRSIISPSQSAFVPGRAISDNVLITHKIL 608
Query: 173 NVLDNKSFGGN--LALKIDVTKAFDTLNWDFLLLVLKTFGFNELFCNWIKTILHSSKMFI 230
+ L + +A+K D+TKA+D + W+FL VL GF+ ++ +WI + +
Sbjct: 607 HYLRQSGAKKHVSMAVKTDMTKAYDRIAWNFLREVLTRLGFHGIWISWIMECVSTVSYSF 428
Query: 231 SMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSRSISILADKGLIDLIAASRNNCLPF 290
+NG G +RG+RQGDPLSP LF + EVLS KG + + +R NC P
Sbjct: 427 LINGGPQGRVLPSRGLRQGDPLSPYLFILCTEVLSGLCQQALRKGTLPGVKVAR-NCPPI 251
Query: 291 -HCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCSGQIMNIRKSFI 334
H + DD M F K+ SS +L S+ +Y SG+ +N KS I
Sbjct: 250 NHLLFADDTMFFGKSNASSCAILLSIMDKYRAASGRCIN*TKSAI 116
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 112 bits (279), Expect = 5e-25
Identities = 87/347 (25%), Positives = 159/347 (45%), Gaps = 12/347 (3%)
Frame = +2
Query: 83 YEAVLDFFKNGWLPNNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAK 142
+ V +F +N L N+ I LIPK N ++ +R I+LV +KI+ K+LA+RL
Sbjct: 2 FRFVSEFHRNRKLFKGINSTFIALIPKVDNPQRLNDFRPISLVGSLYKILGKLLANRLRV 181
Query: 143 ILPSIISKEQRGFVQGRNIRDCIALTSEAINVLDNKSFGGNLALKI-DVTKAFDTLNWDF 201
++ S+IS Q FV+ R I + + L L +++ + K F L W
Sbjct: 182 VIGSVISDAQSAFVKNRQILEMVFL*QM------------RLWMRLRN*RKIFCCLRWIL 325
Query: 202 LLLVLKTFG-----------FNELFCNWIKTILHSSKMFISMNGAQHGFFNCNRGVRQGD 250
L+ + G F L+ WIK + ++ + +NG+
Sbjct: 326 KRLITLSIGLIWILF*VGMSFLVLWRKWIKECVSTATTSVLVNGS--------------- 460
Query: 251 PLSPLLFCIVEEVLSRSISILADKGLIDLIAASRNNCLPFHCFYVDDLMVFCKAKMSSLI 310
P + L+ +V+ L S G+++ + S H + +D ++ +++
Sbjct: 461 PTNVLMKSLVQTQLFTRYSF----GVVNPVVVS-------HLQFANDTLLLETKNWANIR 607
Query: 311 VLKSLFTRYADCSGQIMNIRKSFIFAGGITDTRMNNIVNILGFNVGSLPFTYLGAPIFKG 370
L++ + SG +N KS + I + ++ ++L + VG +PF YLG PI
Sbjct: 608 ALRAALVIF*AMSGLKVNFHKSGLVCVNIAPSWLSEAASVLSWKVGKVPFLYLGMPIEGN 787
Query: 371 KPKGIHFQPIADKVKAKLAKWKASLLSIAGRIQLVKSVVQSMLVHTM 417
+ ++PI +++KA+L W + LS GR+ L+KSV+ S+ V+ +
Sbjct: 788 SRRLSFWEPIVNRIKARLTGWNSRFLSFGGRLVLLKSVLTSLSVYAL 928
>BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 687
Score = 100 bits (250), Expect = 1e-21
Identities = 54/120 (45%), Positives = 70/120 (58%)
Frame = +2
Query: 44 TKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFFKNGWLPNNFNANS 103
T+EEV A+ ++ APGPD A FFQ YW+IV K+V + VL N N
Sbjct: 260 TEEEVLEAINQMHPVKAPGPDGLPALFFQKYWHIVGKEVQQMVLQVLNNSMETEELNKTF 439
Query: 104 IILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQRGFVQGRNIRD 163
I+LIPK N ++ YR I+L N KII KV+A+R+ + LP +I EQ FVQGR I D
Sbjct: 440 IVLIPKGKNPNTPKDYRPISLCNVVMKIITKVIANRVKQTLPDVIDVEQSAFVQGRLITD 619
>BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - Arabidopsis
thaliana, partial (17%)
Length = 677
Score = 72.0 bits (175), Expect(2) = 2e-16
Identities = 38/89 (42%), Positives = 51/89 (56%)
Frame = +3
Query: 70 FFQIYWNIVKKDVYEAVLDFFKNGWLPNNFNANSIILIPKTPNADSVDQYRTIALVNFKF 129
FFQ W+I+K D+ + V F +G L N +I LIPK + + R I+L N +
Sbjct: 411 FFQHSWHIIKMDLLKMVNSFLASGKLDTRLNTTNICLIPKKKRPTRMTELRPISLCNVGY 590
Query: 130 KIINKVLADRLAKILPSIISKEQRGFVQG 158
KII+KVL RL LPS+IS+ Q FV G
Sbjct: 591 KIISKVLCQRLKVCLPSLISETQSAFVHG 677
Score = 32.0 bits (71), Expect(2) = 2e-16
Identities = 15/40 (37%), Positives = 24/40 (59%)
Frame = +2
Query: 25 EAIPKLVDATTNRLLTMLPTKEEVKNAVFDLNSDDAPGPD 64
+ I + N+ L L T+EEV+ A+F ++ + APGPD
Sbjct: 275 DEITSSITPQMNQRLLRLATEEEVRLALFIMHPEKAPGPD 394
>BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (16%)
Length = 794
Score = 55.5 bits (132), Expect(2) = 8e-16
Identities = 27/79 (34%), Positives = 40/79 (50%)
Frame = +3
Query: 26 AIPKLVDATTNRLLTMLPTKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEA 85
+IP +V N L ++EEV+ AVFD+N PGPD FFQ +W+ + D+
Sbjct: 360 SIPAIVTEEQNAQLMAQISREEVREAVFDINPHKCPGPDGMNVFFFQQFWDTMGDDLTSM 539
Query: 86 VLDFFKNGWLPNNFNANSI 104
+F + G L N +I
Sbjct: 540 AQEFLRTGKLEEGINKTNI 596
Score = 46.2 bits (108), Expect(2) = 8e-16
Identities = 24/62 (38%), Positives = 39/62 (62%)
Frame = +1
Query: 106 LIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQRGFVQGRNIRDCI 165
L+PK A + ++R I+L N +KI++KVL+ RL +LP II++ Q F + + I D I
Sbjct: 601 LVPKKLEAKRLVEFRPISLCNVAYKIVSKVLSKRLKSVLPWIITETQAAFGRRQLISDNI 780
Query: 166 AL 167
+
Sbjct: 781 LI 786
>TC82948
Length = 705
Score = 69.3 bits (168), Expect = 4e-12
Identities = 43/131 (32%), Positives = 66/131 (49%), Gaps = 22/131 (16%)
Frame = +3
Query: 447 KMVTVAWRKICADYEEGGLGVKSLICLNEATNLKICWNLMQSDEQWANIIR-SRVIRDH- 504
K+V V+W K+C +EG LG++SL LNEA NLK+CW++M S EQW +R +R + H
Sbjct: 255 KVVKVSWEKVCRPIKEGSLGIRSLSKLNEALNLKLCWDMMISKEQWYAFMRHNRALLGHI 434
Query: 505 --RC--------MNHH------ISSSIWSG----VKAEFHVIKDNCSWIVGDGKQINFWS 544
C HH +S IW + FH+ + W + D +++ +
Sbjct: 435 DFMCSLCFKSFESTHHLFFECSVSVQIWDWFSNLINLNFHINNTDDLWRICD-REVGAAN 611
Query: 545 DSWCGQVLAQT 555
SW +L+ T
Sbjct: 612 VSWLSLLLSST 644
>BQ148771
Length = 680
Score = 65.9 bits (159), Expect = 4e-11
Identities = 40/160 (25%), Positives = 78/160 (48%), Gaps = 10/160 (6%)
Frame = -3
Query: 383 KVKAKLAKWKASLLSIAGRIQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFIWSGD 442
+V LA WKA+ LS+A R+ L KSV++++ ++ M P ++E++K + F+W
Sbjct: 588 QVHVMLANWKANHLSLARRVTLAKSVIEAVPLYPMMTTIIPKACIEEIQKLQRKFVWGDT 409
Query: 443 VTKRKMVTVAWRKICADYEEGGLGVKSLICLNEATNLKICWNLMQ-SDEQWANIIRSRVI 501
R+ V W + GLG++ L +N+A +K+ W++ S+ ++R +
Sbjct: 408 EVSRRYHAVGWETMSKPKTIYGLGLRRLDVMNKACIMKLGWSIYSGSNSLCTEVMRGKYQ 229
Query: 502 RDHRC----MNHHISSSIWSGV-----KAEFHVIKDNCSW 532
R + SS+W + + E +++ N +W
Sbjct: 228 RSESLEEIFLEKPTDSSLWKALVKLWPEIERNLVDSNGNW 109
>BE248682 similar to GP|18568269|gb putative gag-pol polyprotein {Zea mays},
partial (1%)
Length = 441
Score = 61.2 bits (147), Expect = 1e-09
Identities = 37/123 (30%), Positives = 54/123 (43%)
Frame = +3
Query: 244 RGVRQGDPLSPLLFCIVEEVLSRSISILADKGLIDLIAASRNNCLPFHCFYVDDLMVFCK 303
RG++QGDPL+P LF +V E +S + ++ L R H Y DD +
Sbjct: 24 RGLKQGDPLAPFLFLLVAEGISGLMKNAVNRNLFQGFDVKRGGTRVSHLQYADDTLCIGM 203
Query: 304 AKMSSLIVLKSLFTRYADCSGQIMNIRKSFIFAGGITDTRMNNIVNILGFNVGSLPFTYL 363
+ +L LK+L + SG +N KS + + M L S+PF YL
Sbjct: 204 PTVDNLWTLKALLQGFEMASGLKVNFHKSSLIGINVPRDFMEAACRFLNCREESIPFIYL 383
Query: 364 GAP 366
G P
Sbjct: 384 GLP 392
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 58.2 bits (139), Expect = 8e-09
Identities = 32/115 (27%), Positives = 57/115 (48%), Gaps = 1/115 (0%)
Frame = +1
Query: 244 RGVRQGDPLSPLLFCIVEEVLSRSISILADKGLIDLIAASRNNCLPFHCFYVDDLMVFCK 303
+G+RQGDPLSP LF + VLS + +K + I +R++ H + DD ++F +
Sbjct: 10 KGLRQGDPLSPYLFILCANVLSGLLKREGNKQNLHGIQVARSDPKITHLLFADDSLLFAR 189
Query: 304 AKMSSLIVLKSLFTRYADCSGQIMNIRKSFI-FAGGITDTRMNNIVNILGFNVGS 357
A ++ + + Y SGQ++N KS + ++ + + I + GS
Sbjct: 190 ANLTEAATIMQVLHSYQSASGQLVNFEKSEVSYSQNVPNQEKEMICQQIAIKTGS 354
>BG582960
Length = 852
Score = 53.5 bits (127), Expect = 2e-07
Identities = 32/62 (51%), Positives = 38/62 (60%)
Frame = +3
Query: 167 LTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFNELFCNWIKTILHSS 226
L E IN LD K GNLALKI + KA D L +FLL VLK F F +F N K+IL+
Sbjct: 51 LILEEINFLDKKILRGNLALKITMRKALDNLV*EFLLNVLKPFVFKHIFYN*NKSILNCV 230
Query: 227 KM 228
K+
Sbjct: 231 KL 236
>TC90516
Length = 983
Score = 48.1 bits (113), Expect = 9e-06
Identities = 39/137 (28%), Positives = 64/137 (46%)
Frame = -2
Query: 36 NRLLTMLPTKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFFKNGWL 95
N LL+ +++ V L+ +++P PD F FF N++K D+ F+ L
Sbjct: 541 NVLLSAQFLVSKMELVVSSLDGNESPRPDGFNLNFFIRLRNMLKADIEIMFEQFYTPANL 362
Query: 96 PNNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQRGF 155
F+ + LI K + + ++ + K++ KVLA RLA I+ II Q F
Sbjct: 361 LKIFS*YFLTLISKVEYPILLGDFSLMSFLGSL*KLMAKVLALRLAHIMEKIIFVNQSTF 182
Query: 156 VQGRNIRDCIALTSEAI 172
V+GR D + +E I
Sbjct: 181 VRGRQHVDGVVAINEII 131
>TC87634
Length = 677
Score = 39.7 bits (91), Expect(2) = 1e-04
Identities = 22/56 (39%), Positives = 29/56 (51%), Gaps = 1/56 (1%)
Frame = +3
Query: 384 VKAKLAKWKASLLSIAGR-IQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFI 438
++AK WK SLLSI L + T + WPI +LK M+ WI+NFI
Sbjct: 396 IRAKNLAWKGSLLSILWEESNLFDQLFIVCWCTTFACILWPISLLKIMDSWIRNFI 563
Score = 23.9 bits (50), Expect(2) = 1e-04
Identities = 10/15 (66%), Positives = 11/15 (72%)
Frame = +1
Query: 366 PIFKGKPKGIHFQPI 380
PIFKG P I+ QPI
Sbjct: 343 PIFKGMP*PIYLQPI 387
>AW736247 weakly similar to PIR|T14619|T146 reverse transcriptase - beet
retrotransposon (fragment), partial (4%)
Length = 305
Score = 43.5 bits (101), Expect = 2e-04
Identities = 21/54 (38%), Positives = 32/54 (58%), Gaps = 2/54 (3%)
Frame = +1
Query: 44 TKEEVKNAVFDLNSDDA--PGPDVFGACFFQIYWNIVKKDVYEAVLDFFKNGWL 95
++EEV+ AV+D +S + PGPD F + YW ++K D V++F NG L
Sbjct: 124 SEEEVRKAVWDCDSSHS*NPGPDGVNFTFVKEYWELIKVDFLRVVMEFHTNGKL 285
>TC81772
Length = 982
Score = 37.0 bits (84), Expect = 0.020
Identities = 15/38 (39%), Positives = 20/38 (52%)
Frame = -2
Query: 510 HISSSIWSGVKAEFHVIKDNCSWIVGDGKQINFWSDSW 547
H S IW V V+++ W +GDG IN W+D W
Sbjct: 711 HNPSYIWCSV*TLRMVLEEGHHWKIGDGSSINIWTDHW 598
>TC85550 homologue to SP|P28002|COMT_MEDSA Caffeic acid 3-O-methyltransferase
(EC 2.1.1.68), complete
Length = 1541
Score = 33.9 bits (76), Expect = 0.17
Identities = 11/44 (25%), Positives = 28/44 (63%)
Frame = +1
Query: 1 MWFLLIKTCFFLLTIFLQDHLLVEEAIPKLVDATTNRLLTMLPT 44
+WF L+++C+ ++ + ++H +PK++D+ ++ + ML T
Sbjct: 1039 VWFTLMQSCWLIIQVGKREHRKSLRILPKVLDSKVSKFIVMLST 1170
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 29.3 bits (64), Expect(2) = 0.40
Identities = 15/33 (45%), Positives = 21/33 (63%)
Frame = +2
Query: 243 NRGVRQGDPLSPLLFCIVEEVLSRSISILADKG 275
+RG++QGD LSP +F I E LS I ++G
Sbjct: 167 SRGLQQGDHLSPYIFIICVEGLSFLIPHAKERG 265
Score = 21.9 bits (45), Expect(2) = 0.40
Identities = 7/30 (23%), Positives = 18/30 (59%)
Frame = +1
Query: 190 VTKAFDTLNWDFLLLVLKTFGFNELFCNWI 219
++K ++ ++ D+L ++ GFN + W+
Sbjct: 7 ISKVYNRVD*DYLKEIMIKMGFNNRWIYWM 96
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 32.3 bits (72), Expect = 0.49
Identities = 11/25 (44%), Positives = 16/25 (64%)
Frame = +3
Query: 527 KDNCSWIVGDGKQINFWSDSWCGQV 551
++N +VGDG+ FW D+W G V
Sbjct: 243 EENIRMVVGDGRDAFFWYDTWAGDV 317
>TC82754 homologue to GP|18181865|emb|CAD20127. raffinose synthase {Pisum
sativum}, partial (64%)
Length = 1687
Score = 30.8 bits (68), Expect = 1.4
Identities = 16/47 (34%), Positives = 25/47 (53%)
Frame = -2
Query: 153 RGFVQGRNIRDCIALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNW 199
R F Q + R+C+ LT++ + NKSF L+I +T +NW
Sbjct: 948 RVFKQAISCRECVMLTTQDRTIRQNKSF---KQLEIVITDTIANINW 817
>TC86802 similar to GP|7939577|dbj|BAA95778.1 ATP-dependent RNA helicase
{Arabidopsis thaliana}, partial (52%)
Length = 2777
Score = 30.4 bits (67), Expect = 1.9
Identities = 8/33 (24%), Positives = 22/33 (66%)
Frame = -3
Query: 270 ILADKGLIDLIAASRNNCLPFHCFYVDDLMVFC 302
+ + ++ L+++SRN C + C++++ +V+C
Sbjct: 657 LCSPNNILPLMSSSRNTCFLYRCWFLESQLVYC 559
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.326 0.140 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,631,709
Number of Sequences: 36976
Number of extensions: 326160
Number of successful extensions: 2201
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 2176
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2199
length of query: 555
length of database: 9,014,727
effective HSP length: 101
effective length of query: 454
effective length of database: 5,280,151
effective search space: 2397188554
effective search space used: 2397188554
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC149134.1


