

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
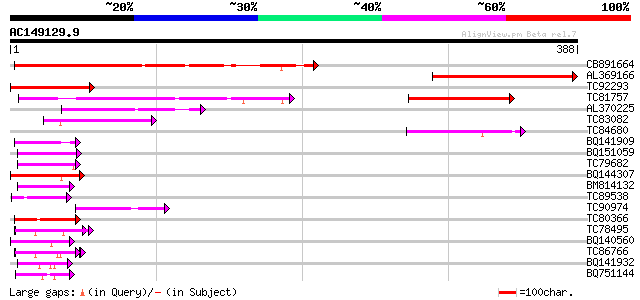
Query= AC149129.9 + phase: 0
(388 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CB891664 similar to GP|3341693|gb| unknown protein {Arabidopsis ... 227 7e-60
AL369166 similar to GP|3341693|gb|A unknown protein {Arabidopsis... 195 2e-50
TC92293 similar to SP|Q64733|FXB2_MOUSE Forkhead box protein B2 ... 120 9e-28
TC81757 similar to GP|3341693|gb|AAC27475.1| unknown protein {Ar... 114 5e-26
AL370225 homologue to GP|21536636|gb| unknown {Arabidopsis thali... 62 5e-10
TC83082 similar to GP|18149189|dbj|BAB83610. 6b-interacting prot... 52 5e-07
TC84680 similar to GP|21536636|gb|AAM60968.1 unknown {Arabidopsi... 45 4e-05
BQ141909 weakly similar to GP|14253159|emb VMP3 protein {Volvox ... 44 8e-05
BQ151059 similar to GP|7291525|gb|A CG9888-PA {Drosophila melano... 44 8e-05
TC79682 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproli... 44 1e-04
BQ144307 weakly similar to GP|6523547|emb| hydroxyproline-rich g... 44 1e-04
BM814132 weakly similar to OMNI|MT3615.1 PE_PGRS family protein ... 44 1e-04
TC89538 weakly similar to GP|21689895|gb|AAM67508.1 unknown prot... 43 2e-04
TC90974 weakly similar to GP|9294038|dbj|BAB01995.1 gene_id:MOB2... 43 2e-04
TC80366 similar to GP|13877617|gb|AAK43886.1 protein kinase-like... 43 2e-04
TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 ... 43 2e-04
BQ140560 weakly similar to GP|15128223|db hypothetical protein~s... 42 4e-04
TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E... 42 5e-04
BQ141932 weakly similar to GP|6523547|emb hydroxyproline-rich gl... 41 7e-04
BQ751144 weakly similar to PIR|PQ0479|PQ04 pistil extensin-like ... 41 7e-04
>CB891664 similar to GP|3341693|gb| unknown protein {Arabidopsis thaliana},
partial (26%)
Length = 580
Score = 227 bits (578), Expect = 7e-60
Identities = 122/210 (58%), Positives = 149/210 (70%), Gaps = 2/210 (0%)
Frame = +1
Query: 4 SSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPEETSALIDSYRDKWY 63
SSPP ++ + PP+AVPLPL+LPAP P P S+RRLPPPCW+PEET ALIDSYR+KWY
Sbjct: 34 SSPPSTTTPPPLSDPPTAVPLPLALPAPLPIPPSTRRLPPPCWSPEETLALIDSYREKWY 213
Query: 64 SLGRTNLKATHWQEVADAVAVRCPNSSPVAKTAVQCRHKMEKLRKRYRSEIQKLRSLPVP 123
SLGR NLKATHWQEVAD+V+ RCPN +P AKT VQCRHKMEKLRKRYR+EIQ+ RSL P
Sbjct: 214 SLGRGNLKATHWQEVADSVSNRCPNVTP-AKTPVQCRHKMEKLRKRYRTEIQRARSL--P 384
Query: 124 RSRSSSSWVLFKAMDSMEKGPSPPSPPHKPENPNHNANHNHIHSHNLINHRETLENDDYD 183
SR +S+WV FK MDSMEKGPSP K EN N ++ ++D+ +
Sbjct: 385 HSRFNSAWVHFKLMDSMEKGPSPV----KSEN----------------NDSDSADDDEDE 504
Query: 184 D--DDLYEELRSAAGGGSGSGNTRSLDKLY 211
D DLY ++++ G NTRS++KLY
Sbjct: 505 DHEQDLYLDIKNGHG-----SNTRSINKLY 579
>AL369166 similar to GP|3341693|gb|A unknown protein {Arabidopsis thaliana},
partial (13%)
Length = 505
Score = 195 bits (496), Expect = 2e-50
Identities = 99/99 (100%), Positives = 99/99 (100%)
Frame = +2
Query: 290 EMVNAIKVLRDGFVRMEQMKMEMAREIETMRMEMEMKRTEMILESQQRIVEAFAKAISEK 349
EMVNAIKVLRDGFVRMEQMKMEMAREIETMRMEMEMKRTEMILESQQRIVEAFAKAISEK
Sbjct: 2 EMVNAIKVLRDGFVRMEQMKMEMAREIETMRMEMEMKRTEMILESQQRIVEAFAKAISEK 181
Query: 350 NKKRKLRKGNTINNNDNNIINNNGNNNNNNNNIASPSES 388
NKKRKLRKGNTINNNDNNIINNNGNNNNNNNNIASPSES
Sbjct: 182 NKKRKLRKGNTINNNDNNIINNNGNNNNNNNNIASPSES 298
>TC92293 similar to SP|Q64733|FXB2_MOUSE Forkhead box protein B2
(Transcription factor FKH-4). [Mouse] {Mus musculus},
partial (4%)
Length = 626
Score = 120 bits (301), Expect = 9e-28
Identities = 56/58 (96%), Positives = 56/58 (96%)
Frame = +1
Query: 1 MATSSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPEETSALIDSY 58
MATSSPPPSSPTESIPSPPSAVPLPLSLPAPP PTSSRRLPPPCWTPEETSALIDSY
Sbjct: 451 MATSSPPPSSPTESIPSPPSAVPLPLSLPAPPTTPTSSRRLPPPCWTPEETSALIDSY 624
>TC81757 similar to GP|3341693|gb|AAC27475.1| unknown protein {Arabidopsis
thaliana}, partial (5%)
Length = 975
Score = 114 bits (286), Expect = 5e-26
Identities = 78/201 (38%), Positives = 105/201 (51%), Gaps = 12/201 (5%)
Frame = +3
Query: 7 PPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPEETSALIDSYRDKWYSLG 66
P S+ + + SP + LP P P P WT +ET LI +Y+DKWYSL
Sbjct: 54 PYSTNSMATDSPELTLSLPTKKPQPIP------------WTHQETINLIRAYQDKWYSLK 197
Query: 67 RTNLKATHWQEVADAVAVRCP-NSSPVAKTAVQCRHKMEKLRKRYRSEIQKLRSLPVPRS 125
R L+ + W+EVA VA RC + + +KTA+QCRHKMEKLR+R+RSE K R
Sbjct: 198 RGPLRGSQWEEVAVVVAARCGYDYNHPSKTALQCRHKMEKLRQRHRSE--KRRLTATSSV 371
Query: 126 RSSSSWVLFKAMDSMEKGPSPPSPPHKPENPNH-------NANHNHIHSHNLINHRETLE 178
SS SW + MD +E+GP P S +P + NH A HN++N ++ E
Sbjct: 372 ASSRSWQYLRLMDDLERGPLPISV--RPLSHNHPISDDSDGAAARSRSIHNILNQKQRDE 545
Query: 179 NDDYDDD----DLYEELRSAA 195
D+ +D L ELRS A
Sbjct: 546 TDEEKEDVMAKGLTAELRSFA 608
Score = 57.0 bits (136), Expect = 1e-08
Identities = 30/73 (41%), Positives = 46/73 (62%), Gaps = 1/73 (1%)
Frame = +3
Query: 274 ERERDVERERERDPIGEMVNA-IKVLRDGFVRMEQMKMEMAREIETMRMEMEMKRTEMIL 332
+++RD E + D + + + A ++ + + +E MKMEM +E E R+EME KR MIL
Sbjct: 528 QKQRDETDEEKEDVMAKGLTAELRSFAERIIGLENMKMEMMKETERFRLEMENKRIRMIL 707
Query: 333 ESQQRIVEAFAKA 345
ESQ RIV++ KA
Sbjct: 708 ESQWRIVDSIGKA 746
>AL370225 homologue to GP|21536636|gb| unknown {Arabidopsis thaliana},
partial (28%)
Length = 408
Score = 61.6 bits (148), Expect = 5e-10
Identities = 34/99 (34%), Positives = 51/99 (51%)
Frame = +3
Query: 36 TSSRRLPPPCWTPEETSALIDSYRDKWYSLGRTNLKATHWQEVADAVAVRCPNSSPVAKT 95
+ + RL W+ + L+++Y KW R LK W++VA V+ R NS+ KT
Sbjct: 141 SGNERLKRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRS-NSTKSPKT 317
Query: 96 AVQCRHKMEKLRKRYRSEIQKLRSLPVPRSRSSSSWVLF 134
QC++K+E ++KRYRSE S SSW L+
Sbjct: 318 QTQCKNKIESMKKRYRSE---------SASSDVSSWPLY 407
>TC83082 similar to GP|18149189|dbj|BAB83610. 6b-interacting protein 1
{Nicotiana tabacum}, partial (19%)
Length = 374
Score = 51.6 bits (122), Expect = 5e-07
Identities = 27/85 (31%), Positives = 46/85 (53%), Gaps = 8/85 (9%)
Frame = +2
Query: 24 LPLSLPAPPP--------PPTSSRRLPPPCWTPEETSALIDSYRDKWYSLGRTNLKATHW 75
L L+L +P P PP+ + + CW+ + +S L+D++ ++ L R NL+ W
Sbjct: 119 LTLTLISPMPDFHHDSLTPPSRTVPVREDCWSEDASSTLVDAWGRRYLELNRGNLRQKDW 298
Query: 76 QEVADAVAVRCPNSSPVAKTAVQCR 100
Q+VADAV ++ +T VQC+
Sbjct: 299 QDVADAVNALHGHTKKTHRTDVQCK 373
>TC84680 similar to GP|21536636|gb|AAM60968.1 unknown {Arabidopsis
thaliana}, partial (26%)
Length = 431
Score = 45.4 bits (106), Expect = 4e-05
Identities = 29/89 (32%), Positives = 48/89 (53%), Gaps = 7/89 (7%)
Frame = +1
Query: 272 KRERERDVERERERDPIGEMVNAIKVLRDGFVRMEQMKMEMAREIETMRME-------ME 324
+R R +R++ + +I+ + + VR EQ +ME +EIE MR+E ME
Sbjct: 19 RRMRTETNNNKRQKKETMGIAESIRWVAEVVVRSEQTRMETLKEIEKMRVEAEAKRGEME 198
Query: 325 MKRTEMILESQQRIVEAFAKAISEKNKKR 353
+KRTE+I +Q I FA ++ K+K +
Sbjct: 199 IKRTEIIANTQLEIARIFAN-VNNKDKDK 282
>BQ141909 weakly similar to GP|14253159|emb VMP3 protein {Volvox carteri f.
nagariensis}, partial (13%)
Length = 1223
Score = 44.3 bits (103), Expect = 8e-05
Identities = 23/45 (51%), Positives = 24/45 (53%)
Frame = -1
Query: 4 SSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
SSP PSSP S P P S P P P PPPPP PPP +P
Sbjct: 293 SSPSPSSPLPSPPPPSSLFPSPTPPPLPPPPP------PPPPASP 177
Score = 37.7 bits (86), Expect = 0.008
Identities = 21/42 (50%), Positives = 23/42 (54%), Gaps = 5/42 (11%)
Frame = -3
Query: 6 PPPSSPTESIPSP---PSAVPLPLSLP--APPPPPTSSRRLP 42
PPP P +P P P +PLPLS P APPPP S R P
Sbjct: 303 PPPFFPLPLLPPPLPPPPLLPLPLSDPPSAPPPPSPPSPRFP 178
Score = 37.7 bits (86), Expect = 0.008
Identities = 20/41 (48%), Positives = 21/41 (50%), Gaps = 9/41 (21%)
Frame = -1
Query: 6 PPPSSPTESIPSP---------PSAVPLPLSLPAPPPPPTS 37
PP SSP+ S P P PS P PL P PPPPP S
Sbjct: 302 PPLSSPSPSSPLPSPPPPSSLFPSPTPPPLPPPPPPPPPAS 180
Score = 35.8 bits (81), Expect = 0.029
Identities = 19/45 (42%), Positives = 21/45 (46%)
Frame = -2
Query: 4 SSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
SSP + P SPP P P PPPPP S L PP +P
Sbjct: 343 SSPSLTLPPSFCSSPPFLPPPPPPPSPPPPPPPPSSPLRPPLRSP 209
Score = 35.8 bits (81), Expect = 0.029
Identities = 19/41 (46%), Positives = 23/41 (55%)
Frame = -2
Query: 4 SSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
++P SSP S+ PPS P LP PPPPP+ PPP
Sbjct: 358 AAPLRSSP--SLTLPPSFCSSPPFLPPPPPPPSPPPPPPPP 242
Score = 35.4 bits (80), Expect = 0.038
Identities = 20/44 (45%), Positives = 24/44 (54%), Gaps = 1/44 (2%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPP-PCWTP 48
PPP P+ P PP + PL L +PPP P LPP P +TP
Sbjct: 286 PPPPPPSPPPPPPPPSSPLRPPLRSPPPLPP----LPPLPPFTP 167
Score = 33.1 bits (74), Expect = 0.19
Identities = 22/51 (43%), Positives = 23/51 (44%), Gaps = 2/51 (3%)
Frame = -3
Query: 4 SSPPPSS--PTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPEETS 52
S PPP S P +P PP PLPL P PPPP L P P S
Sbjct: 348 SVPPPLSRFPLPFVPPPPF-FPLPLLPPPLPPPPLLPLPLSDPPSAPPPPS 199
Score = 32.3 bits (72), Expect = 0.32
Identities = 20/45 (44%), Positives = 20/45 (44%), Gaps = 4/45 (8%)
Frame = -2
Query: 4 SSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRL----PPP 44
S PPP P S PP P PL P PP T S L PPP
Sbjct: 268 SPPPPPPPPSSPLRPPLRSPPPLPPLPPLPPFTPSSHLTTSSPPP 134
Score = 30.0 bits (66), Expect = 1.6
Identities = 22/62 (35%), Positives = 24/62 (38%), Gaps = 22/62 (35%)
Frame = -3
Query: 5 SPPPSSPT-------------------ESIPSPPSAVPLPLSLPAPPPPPTSSR---RLP 42
SPPP SP S P S P P + AP PP+SSR RL
Sbjct: 1044 SPPPLSPVLSPPHLSLTSRSSHFCYSLHSTPRHASLPPAPFPVAAPSLPPSSSRLAPRLS 865
Query: 43 PP 44
PP
Sbjct: 864 PP 859
Score = 30.0 bits (66), Expect = 1.6
Identities = 21/47 (44%), Positives = 23/47 (48%), Gaps = 4/47 (8%)
Frame = -3
Query: 6 PPPSSPTE-SIPSPPSAVPLPLSLPAPPPPPTSSRRL---PPPCWTP 48
PPP SP +P PS PLP P PPP S L PPP + P
Sbjct: 417 PPPLSPRLLPVPFLPSIFPLP---PPSVPPPLSRFPLPFVPPPPFFP 286
Score = 29.6 bits (65), Expect = 2.1
Identities = 13/30 (43%), Positives = 17/30 (56%)
Frame = -3
Query: 3 TSSPPPSSPTESIPSPPSAVPLPLSLPAPP 32
++ PPPS P+ P P + LPL LP P
Sbjct: 219 SAPPPPSPPSPRFPLSPLLLILPLPLPLLP 130
Score = 28.5 bits (62), Expect = 4.6
Identities = 18/43 (41%), Positives = 18/43 (41%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
PPPS P P VP P P P PP LPPP P
Sbjct: 357 PPPSVPPPLSRFPLPFVPPPPFFPLPLLPPP----LPPPPLLP 241
Score = 28.5 bits (62), Expect = 4.6
Identities = 22/61 (36%), Positives = 26/61 (42%), Gaps = 22/61 (36%)
Frame = -1
Query: 6 PPPSSPTESIP-----SPPSAVPL-----------PLSLPAP------PPPPTSSRRLPP 43
PP SS + S P PP +PL PLS P+P PPPP+S P
Sbjct: 404 PPASSLSHSFPLFFPCRPPPFLPLSHASPFLLFLPPLSSPSPSSPLPSPPPPSSLFPSPT 225
Query: 44 P 44
P
Sbjct: 224 P 222
>BQ151059 similar to GP|7291525|gb|A CG9888-PA {Drosophila melanogaster},
partial (26%)
Length = 308
Score = 44.3 bits (103), Expect = 8e-05
Identities = 20/44 (45%), Positives = 22/44 (49%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPE 49
PPP P P PP P P P PPPPP PPP ++PE
Sbjct: 150 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPGFSPE 19
Score = 40.4 bits (93), Expect = 0.001
Identities = 19/45 (42%), Positives = 20/45 (44%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPEE 50
PPP P P PP P P P PPPPP PPP P +
Sbjct: 152 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPRGFPRK 18
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 307 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 191
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 294 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 178
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 301 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 185
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 290 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 174
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 283 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 167
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 258 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 142
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 279 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 163
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 276 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 160
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 273 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 157
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 293 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 177
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 268 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 152
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 266 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 150
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 210 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 94
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 215 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 99
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 257 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 141
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 252 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 136
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 249 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 133
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 247 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 131
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 244 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 128
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 240 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 124
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 239 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 123
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 236 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 120
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 231 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 115
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 272 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 156
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 226 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 110
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 224 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 108
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 220 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 104
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 216 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 100
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 263 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 147
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 285 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 169
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 207 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 91
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 204 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 88
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 203 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 87
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 198 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 82
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 196 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 80
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 194 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 78
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 189 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 73
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 186 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 70
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 183 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 67
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 181 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 65
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 178 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 62
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 176 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 60
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 171 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 55
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 169 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 53
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 166 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 50
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 163 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 47
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 159 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 43
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 158 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 42
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 154 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 38
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 298 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 182
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 303 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 187
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 228 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 112
Score = 37.7 bits (86), Expect = 0.008
Identities = 19/49 (38%), Positives = 21/49 (42%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPEETSAL 54
PPP P P PP P P P PPPPP PP + + S L
Sbjct: 148 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPGVFPGKGDSCL 2
>TC79682 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproline-rich
glycoprotein DZ-HRGP {Volvox carteri f. nagariensis},
partial (17%)
Length = 1031
Score = 43.9 bits (102), Expect = 1e-04
Identities = 22/47 (46%), Positives = 24/47 (50%), Gaps = 4/47 (8%)
Frame = +1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLP----PPCWTP 48
PPPSSP S P PP +P P PPPPP S P PP + P
Sbjct: 88 PPPSSPPPSFPFPPIVIPGLTPSPPPPPPPKSIFPAPLLPFPPLFPP 228
Score = 32.3 bits (72), Expect = 0.32
Identities = 19/47 (40%), Positives = 21/47 (44%), Gaps = 4/47 (8%)
Frame = +1
Query: 1 MATSSPPPSSPTESIPSP----PSAVPLPLSLPAPPPPPTSSRRLPP 43
+ S PPP P P+P P P PLS P PPTSS P
Sbjct: 145 LTPSPPPPPPPKSIFPAPLLPFPPLFPPPLS---PGSPPTSSSNFSP 276
Score = 29.6 bits (65), Expect = 2.1
Identities = 17/46 (36%), Positives = 20/46 (42%), Gaps = 8/46 (17%)
Frame = +1
Query: 7 PPSSPTESIP---SPPSAVPLPLSL-----PAPPPPPTSSRRLPPP 44
PP P P SPP + P P + P+PPPPP P P
Sbjct: 61 PPLIPNPFQPPPSSPPPSFPFPPIVIPGLTPSPPPPPPPKSIFPAP 198
>BQ144307 weakly similar to GP|6523547|emb| hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (24%)
Length = 1059
Score = 43.5 bits (101), Expect = 1e-04
Identities = 27/55 (49%), Positives = 34/55 (61%), Gaps = 4/55 (7%)
Frame = -1
Query: 1 MATSSPPPSSPTESIPSPPS-AVPLPLSL-PAPPPP--PTSSRRLPPPCWTPEET 51
++T PPP SP S P PP+ + P PLS PA P P PTS+ LPPP +P E+
Sbjct: 495 LSTPRPPPPSPHFSPPPPPAPSPPPPLSRHPAHPNPRSPTSTYALPPPRTSPPES 331
Score = 36.6 bits (83), Expect = 0.017
Identities = 17/41 (41%), Positives = 21/41 (50%), Gaps = 2/41 (4%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPP--PPPTSSRRLPPP 44
PPP P+ IP P+ P P+PP PPP + PPP
Sbjct: 434 PPPPLPSPDIPPTPTPAPRLQLTPSPPHAPPPLNPSTHPPP 312
Score = 31.6 bits (70), Expect = 0.54
Identities = 17/42 (40%), Positives = 24/42 (56%), Gaps = 2/42 (4%)
Frame = -3
Query: 1 MATSSPP-PSSPTESI-PSPPSAVPLPLSLPAPPPPPTSSRR 40
+ TS PP P P ++ P PP+ +P + LP PPP T+ R
Sbjct: 415 LPTSRPPQPPLPDFNLRPPPPTHLPP*IHLPTPPPTKTTKTR 290
Score = 31.2 bits (69), Expect = 0.71
Identities = 17/40 (42%), Positives = 22/40 (54%), Gaps = 3/40 (7%)
Frame = -3
Query: 8 PSSPTESIPSPPSA---VPLPLSLPAPPPPPTSSRRLPPP 44
P+S + PSPP+ +P P S P PPP P + R P P
Sbjct: 508 PNSISLHPPSPPTLSPFLPPPPSGPLPPPSPLPTSRPPQP 389
Score = 28.9 bits (63), Expect = 3.5
Identities = 20/45 (44%), Positives = 22/45 (48%), Gaps = 2/45 (4%)
Frame = -1
Query: 2 ATSSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRR--LPPP 44
A S PPP S + P+P S P S A PPP TS PPP
Sbjct: 438 APSPPPPLSRHPAHPNPRS----PTSTYALPPPRTSPPESIYPPP 316
Score = 27.7 bits (60), Expect = 7.9
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = -3
Query: 12 TESIPSPPSAVPLPLSLPAPPPPPTSSRRLPP 43
+ S+P P + + LP +P P TSS R PP
Sbjct: 754 SHSLPRPRAPLSLPSLHISPVPHTTSSNRTPP 659
>BM814132 weakly similar to OMNI|MT3615.1 PE_PGRS family protein
{Mycobacterium tuberculosis CDC1551}, partial (7%)
Length = 164
Score = 43.5 bits (101), Expect = 1e-04
Identities = 19/39 (48%), Positives = 19/39 (48%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP R PPP
Sbjct: 164 PPPPPPPPPPPPPPPPPPPPPPAPPPPPPPPPPRPPPPP 48
Score = 42.0 bits (97), Expect = 4e-04
Identities = 19/39 (48%), Positives = 19/39 (48%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P PAPPPPP PPP
Sbjct: 136 PPPPPPPPPPPPPPPPPPPPPPPPAPPPPPPPPPPPPPP 20
Score = 41.2 bits (95), Expect = 7e-04
Identities = 18/39 (46%), Positives = 19/39 (48%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P+PP P P P PPPPP PPP
Sbjct: 131 PPPPPPPPPPPAPPPPPPPPPPRPPPPPPPPPPPPPPPP 15
Score = 41.2 bits (95), Expect = 7e-04
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 118 PPPPPPPPPPPPPPPPPPAPPPPPPPPPPPPPPPPRPPP 2
Score = 41.2 bits (95), Expect = 7e-04
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 125 PPPPPPPPPAPPPPPPPPPPRPPPPPPPPPPPPPPPPPP 9
Score = 40.8 bits (94), Expect = 9e-04
Identities = 18/39 (46%), Positives = 19/39 (48%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P + P PPPPP PPP
Sbjct: 161 PPPPPPPPPPPPPPPPPPPPPAPPPPPPPPPPRPPPPPP 45
Score = 40.8 bits (94), Expect = 9e-04
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 158 PPPPPPPPPPPPPPPPPPPPAPPPPPPPPPPRPPPPPPP 42
Score = 40.4 bits (93), Expect = 0.001
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 153 PPPPPPPPPPPPPPPPPPRPPPPPPPPPPPPPPPPPPPP 37
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 132 PPPPPPPPPPPRPPPPPPPPPPPPPPPPPPPPPPPPPPP 16
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 141 PPPPPPPPPPPPPPRPPPPPPPPPPPPPPPPPPPPPPPP 25
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 144 PPPPPPPPPPPPPPPRPPPPPPPPPPPPPPPPPPPPPPP 28
Score = 39.7 bits (91), Expect = 0.002
Identities = 18/39 (46%), Positives = 18/39 (46%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPPP PPP
Sbjct: 147 PPPPPPPPPPPPPPPPRPPPPPPPPPPPPPPPPPPPPPP 31
Score = 38.5 bits (88), Expect = 0.004
Identities = 19/43 (44%), Positives = 19/43 (44%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
PPP P P PP P P P PPPPP PPP P
Sbjct: 120 PPPPPPPRPPPPPPPPPPPP---PPPPPPPPPPPPPPPPAPPP 1
Score = 37.4 bits (85), Expect = 0.010
Identities = 17/39 (43%), Positives = 17/39 (43%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP P P P PPPP PPP
Sbjct: 152 PPPPPPPPPPPPPPPPPPAPPPPPPPPPPRPPPPPPPPP 36
Score = 37.4 bits (85), Expect = 0.010
Identities = 17/39 (43%), Positives = 18/39 (45%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP P P PP+ P P P PPPP PPP
Sbjct: 140 PPPPPPPPPPPPPPAPPPPPPPPPPRPPPPPPPPPPPPP 24
Score = 34.7 bits (78), Expect = 0.064
Identities = 19/43 (44%), Positives = 19/43 (44%)
Frame = -2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
PPP P P PP P P P PPPPP PPP P
Sbjct: 160 PPPPPP----PPPPPPPPPP---PPPPPPPPPPPPPPPPAPPP 53
Score = 33.9 bits (76), Expect = 0.11
Identities = 16/34 (47%), Positives = 16/34 (47%)
Frame = -1
Query: 2 ATSSPPPSSPTESIPSPPSAVPLPLSLPAPPPPP 35
A PPP P P PP P P P PPPPP
Sbjct: 98 APPPPPPPPPPRPPPPPPPPPPPP---PPPPPPP 6
>TC89538 weakly similar to GP|21689895|gb|AAM67508.1 unknown protein
{Arabidopsis thaliana}, partial (37%)
Length = 990
Score = 43.1 bits (100), Expect = 2e-04
Identities = 23/41 (56%), Positives = 24/41 (58%)
Frame = +3
Query: 2 ATSSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLP 42
AT PPPS+ T S P P SA P P S PPPPP SS P
Sbjct: 213 ATPHPPPSA-TPSPPPPSSATPPPSSATPPPPPPFSSTASP 332
Score = 36.6 bits (83), Expect = 0.017
Identities = 19/45 (42%), Positives = 24/45 (53%), Gaps = 5/45 (11%)
Frame = +3
Query: 5 SPPPSSPTESIPSPPSAVPLPLSLPAPPP-----PPTSSRRLPPP 44
+P PSS S +P + P P + P+PPP PP SS PPP
Sbjct: 171 TPQPSSRFSSTAAPATPHPPPSATPSPPPPSSATPPPSSATPPPP 305
Score = 29.3 bits (64), Expect = 2.7
Identities = 18/53 (33%), Positives = 26/53 (48%), Gaps = 1/53 (1%)
Frame = +3
Query: 2 ATSSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP-CWTPEETSA 53
A+SS P ++ P P S + P PPP+++ PPP TP +SA
Sbjct: 132 ASSSSKPFRFSKVTPQPSSRFSSTAAPATPHPPPSATPSPPPPSSATPPPSSA 290
>TC90974 weakly similar to GP|9294038|dbj|BAB01995.1 gene_id:MOB24.1~unknown
protein {Arabidopsis thaliana}, partial (15%)
Length = 690
Score = 42.7 bits (99), Expect = 2e-04
Identities = 18/64 (28%), Positives = 36/64 (56%)
Frame = +1
Query: 46 WTPEETSALIDSYRDKWYSLGRTNLKATHWQEVADAVAVRCPNSSPVAKTAVQCRHKMEK 105
WT T L++ + +K+ +GR +L++ W +VA+ V+ V + QCR ++K
Sbjct: 511 WTEHATFVLLEIWGEKFLQIGRNSLRSEEWHDVAEKVS----EELKVERNVAQCRSVLDK 678
Query: 106 LRKR 109
+++R
Sbjct: 679 VKRR 690
Score = 38.5 bits (88), Expect = 0.004
Identities = 32/110 (29%), Positives = 45/110 (40%), Gaps = 10/110 (9%)
Frame = +2
Query: 134 FKAMDSMEKGPSPPSP--PHKPENPNHNANH--NHIHSHNLINHRETLENDDYDDDDLYE 189
F +MD ME PS P + +NPN H ++ + E E+D+ ++ D Y
Sbjct: 113 FNSMDGMEDDSRYPSKGYPLRRKNPNLRQKHPIRNLQYQRYNDREEDFEDDEPEEFDDYN 292
Query: 190 ELRSAAGGG------SGSGNTRSLDKLYRNGVSGGFGGSGSGFRIRIPTG 233
G G G R K+ R+G SGGFGG +P G
Sbjct: 293 VDEDGIGNGYVGNFEQNDGFVRKKRKM-RSGPSGGFGGGSVSNYELLPWG 439
>TC80366 similar to GP|13877617|gb|AAK43886.1 protein kinase-like protein
{Arabidopsis thaliana}, partial (16%)
Length = 814
Score = 42.7 bits (99), Expect = 2e-04
Identities = 21/45 (46%), Positives = 28/45 (61%)
Frame = +3
Query: 4 SSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
SSPPP++P+ PS P A P P++ APPP ++ PPP TP
Sbjct: 3 SSPPPATPSSPPPSTP-ANPPPVTPSAPPPSTPATPSSPPPSTTP 134
Score = 40.4 bits (93), Expect = 0.001
Identities = 22/43 (51%), Positives = 26/43 (60%), Gaps = 2/43 (4%)
Frame = +3
Query: 4 SSPPPSSPTESIPSPPSAVPLPLSLPAPP--PPPTSSRRLPPP 44
SSPPPS+P P PSA P S PA P PPP+++ PPP
Sbjct: 27 SSPPPSTPANPPPVTPSAP--PPSTPATPSSPPPSTTPSAPPP 149
Score = 38.9 bits (89), Expect = 0.003
Identities = 25/66 (37%), Positives = 37/66 (55%), Gaps = 15/66 (22%)
Frame = +3
Query: 4 SSPPPSSPTESIPSPPS--AVPLPL---SLPAPP----------PPPTSSRRLPPPCWTP 48
S+PPPS+P+ S PSPP+ A+ P + P+PP P P SS+ PPP +P
Sbjct: 135 SAPPPSTPSNSPPSPPTTPAISPPSGGGTTPSPPSRTPPSSDDSPSPPSSKTPPPP--SP 308
Query: 49 EETSAL 54
+S++
Sbjct: 309 PSSSSI 326
Score = 36.6 bits (83), Expect = 0.017
Identities = 23/59 (38%), Positives = 28/59 (46%), Gaps = 12/59 (20%)
Frame = +3
Query: 2 ATSSPP------------PSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
AT S P PS+P S P+ PS+ P P + P+ PPP T S P P TP
Sbjct: 18 ATPSSPPPSTPANPPPVTPSAPPPSTPATPSSPP-PSTTPSAPPPSTPSNSPPSPPTTP 191
Score = 33.1 bits (74), Expect = 0.19
Identities = 28/95 (29%), Positives = 35/95 (36%), Gaps = 26/95 (27%)
Frame = +3
Query: 7 PPSSPTESIPSPPSAVP-LPLSLPAPPPPP--------------TSSRRLPPP------- 44
PPS S P PP+ +P P S APPPPP + LPPP
Sbjct: 537 PPSDHVVSKPPPPAPIPPRPPSHVAPPPPPPAFISSSGGSGSNYSGGELLPPPSPGIAFS 716
Query: 45 ----CWTPEETSALIDSYRDKWYSLGRTNLKATHW 75
+T EE + D + D NL + W
Sbjct: 717 SGKSTFTYEELARATDGFSD-------ANLLMSRW 800
Score = 31.6 bits (70), Expect = 0.54
Identities = 19/44 (43%), Positives = 24/44 (54%), Gaps = 4/44 (9%)
Frame = +3
Query: 4 SSPPPSSP---TESIPS-PPSAVPLPLSLPAPPPPPTSSRRLPP 43
S+PPPS+P + PS PSA P +PP PPT+ PP
Sbjct: 75 SAPPPSTPATPSSPPPSTTPSAPPPSTPSNSPPSPPTTPAISPP 206
>TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 - alfalfa,
partial (97%)
Length = 1049
Score = 42.7 bits (99), Expect = 2e-04
Identities = 23/50 (46%), Positives = 25/50 (50%), Gaps = 1/50 (2%)
Frame = +3
Query: 5 SPPPSSPTESIPSPPSAVPLPLS-LPAPPPPPTSSRRLPPPCWTPEETSA 53
+PPP T SPP A P P S PA PPP T PPP TP S+
Sbjct: 465 TPPPVQSTPPPASPPPASPPPFSPPPATPPPATPPPATPPPALTPTPLSS 614
Score = 41.2 bits (95), Expect = 7e-04
Identities = 24/59 (40%), Positives = 30/59 (50%), Gaps = 5/59 (8%)
Frame = +3
Query: 4 SSPPPSSPTESIP---SPPSAVPLPLSLPAPPPPP--TSSRRLPPPCWTPEETSALIDS 57
S+PPP+SP + P SPP A P P + P PPP T + PP TP A + S
Sbjct: 483 STPPPASPPPASPPPFSPPPATPPPATPPPATPPPALTPTPLSSPPATTPAPAPAKLKS 659
Score = 35.4 bits (80), Expect = 0.038
Identities = 19/54 (35%), Positives = 25/54 (46%), Gaps = 3/54 (5%)
Frame = +3
Query: 4 SSPPPSSPTESIP---SPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPEETSAL 54
+SPPP SP + P +PP A P P P P P ++ P P + AL
Sbjct: 513 ASPPPFSPPPATPPPATPPPATPPPALTPTPLSSPPATTPAPAPAKLKSKAPAL 674
Score = 34.7 bits (78), Expect = 0.064
Identities = 22/58 (37%), Positives = 24/58 (40%), Gaps = 13/58 (22%)
Frame = +3
Query: 4 SSPPPSS----PTESIPSPPSAVPLPLSLPAPP---------PPPTSSRRLPPPCWTP 48
SSPPP+ P +S P P P P L PP PPP S PP TP
Sbjct: 378 SSPPPAQSTPPPVQSSPPPVQQSPPPTPLTPPPVQSTPPPASPPPASPPPFSPPPATP 551
Score = 33.9 bits (76), Expect = 0.11
Identities = 20/52 (38%), Positives = 24/52 (45%), Gaps = 2/52 (3%)
Frame = +3
Query: 2 ATSSPPP--SSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPEET 51
A S+PPP SSP SPP P + + PPP + PPP P T
Sbjct: 393 AQSTPPPVQSSPPPVQQSPPPTPLTPPPVQSTPPPASPPPASPPPFSPPPAT 548
Score = 33.9 bits (76), Expect = 0.11
Identities = 20/46 (43%), Positives = 24/46 (51%), Gaps = 5/46 (10%)
Frame = +3
Query: 4 SSP-PPSSPTESIPSPPSAVPLPLSLPAPP----PPPTSSRRLPPP 44
SSP PP+ P + P+ P A P P+ PP PPP S PPP
Sbjct: 261 SSPAPPTPPANTPPTTPQASPPPVQSSPPPVQSSPPPLQSS--PPP 392
Score = 32.3 bits (72), Expect = 0.32
Identities = 20/49 (40%), Positives = 23/49 (46%), Gaps = 8/49 (16%)
Frame = +3
Query: 4 SSPPPSS----PTESIPSPPSAVPLPLSLPAPP----PPPTSSRRLPPP 44
SSPPP P +S P P + P P+ PP PPPT PPP
Sbjct: 336 SSPPPVQSSPPPLQSSPPPAQSTPPPVQSSPPPVQQSPPPTP--LTPPP 476
Score = 29.6 bits (65), Expect = 2.1
Identities = 19/43 (44%), Positives = 21/43 (48%), Gaps = 2/43 (4%)
Frame = +3
Query: 4 SSPPP--SSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
+SPPP SSP SPP P + PPP SS PPP
Sbjct: 315 ASPPPVQSSPPPVQSSPPPLQSSPPPAQSTPPPVQSS---PPP 434
Score = 29.3 bits (64), Expect = 2.7
Identities = 17/45 (37%), Positives = 23/45 (50%), Gaps = 4/45 (8%)
Frame = +3
Query: 4 SSPPPSSPTESIP----SPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
++ PP++P S P SPP P L + PPP S+ PPP
Sbjct: 288 ANTPPTTPQASPPPVQSSPPPVQSSPPPLQSSPPPAQST---PPP 413
>BQ140560 weakly similar to GP|15128223|db hypothetical protein~similar to
Oryza sativa chromosome 1 P0489A01.2, partial (5%)
Length = 215
Score = 42.0 bits (97), Expect = 4e-04
Identities = 20/46 (43%), Positives = 23/46 (49%), Gaps = 2/46 (4%)
Frame = +1
Query: 1 MATSSPPPSSPTESIPSPPSAVPLPLS--LPAPPPPPTSSRRLPPP 44
M+ + PPP P P PP LP S P PPPPP +PPP
Sbjct: 4 MSKAPPPPPPPFYGAPPPPPPSYLPASSGAPLPPPPPPPPPSMPPP 141
Score = 30.8 bits (68), Expect = 0.93
Identities = 15/30 (50%), Positives = 15/30 (50%)
Frame = +1
Query: 4 SSPPPSSPTESIPSPPSAVPLPLSLPAPPP 33
S PPP P P PP LP S APPP
Sbjct: 127 SMPPPPPPFHGAPPPPXLY-LPASTGAPPP 213
>TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E8.80 -
Arabidopsis thaliana, partial (18%)
Length = 1260
Score = 41.6 bits (96), Expect = 5e-04
Identities = 22/51 (43%), Positives = 23/51 (44%), Gaps = 4/51 (7%)
Frame = +2
Query: 5 SPPPSSPTESIP----SPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPEET 51
SPPP P S P SPP P P P PPPPP PP P +T
Sbjct: 215 SPPPPPPVHSPPPPVYSPPPPSPPPCVEPPPPPPPPCVEPPPPSSPAPHQT 367
Score = 41.2 bits (95), Expect = 7e-04
Identities = 23/56 (41%), Positives = 26/56 (46%), Gaps = 7/56 (12%)
Frame = +2
Query: 4 SSPPPSSPTESIPSPPSAVPLPLSLPAP-------PPPPTSSRRLPPPCWTPEETS 52
SSPPP P S P PP + P P P P PPPP PPP ++P S
Sbjct: 116 SSPPPP-PAHSPPPPPXSPPPPPHSPTPPVYPYLSPPPPPPVHSPPPPVYSPPPPS 280
Score = 40.8 bits (94), Expect = 9e-04
Identities = 21/48 (43%), Positives = 24/48 (49%), Gaps = 3/48 (6%)
Frame = +2
Query: 4 SSPPPSSPTESIPSPPSAVPLPLSLPAPPP---PPTSSRRLPPPCWTP 48
+SPPP P + SPP VP S P PPP PP PPP +P
Sbjct: 47 NSPPPPPPPAPVFSPPPPVPYYYSSPPPPPAHSPPPPPXSPPPPPHSP 190
Score = 37.7 bits (86), Expect = 0.008
Identities = 20/48 (41%), Positives = 23/48 (47%), Gaps = 5/48 (10%)
Frame = +2
Query: 4 SSPPPSSPTESIPSPPSAV-----PLPLSLPAPPPPPTSSRRLPPPCW 46
S PP SP + PSPP V P P+ PPPP S PPP +
Sbjct: 386 SPSPPPSPVYAYPSPPPPVYTSPPPSPVYAYPSPPPPVYSSPPPPPVY 529
Score = 36.2 bits (82), Expect = 0.022
Identities = 20/47 (42%), Positives = 22/47 (46%), Gaps = 8/47 (17%)
Frame = +2
Query: 6 PPPSSPTESI-----PSPPSAV---PLPLSLPAPPPPPTSSRRLPPP 44
PPP SPT + P PP V P P+ P PP PP PPP
Sbjct: 173 PPPHSPTPPVYPYLSPPPPPPVHSPPPPVYSPPPPSPPPCVEPPPPP 313
Score = 34.7 bits (78), Expect = 0.064
Identities = 22/50 (44%), Positives = 24/50 (48%), Gaps = 10/50 (20%)
Frame = +2
Query: 5 SPPPSSPTESI-----PSPPSAVPLPLSLPAP-----PPPPTSSRRLPPP 44
SPPP SP + P PP P P S PAP PPP+ S PPP
Sbjct: 263 SPPPPSPPPCVEPPPPPPPPCVEPPPPSSPAPHQTPYHPPPSPS---PPP 403
Score = 34.3 bits (77), Expect = 0.084
Identities = 19/44 (43%), Positives = 21/44 (47%), Gaps = 2/44 (4%)
Frame = +2
Query: 2 ATSSPPPSSPTESIPSPPSAVPLPLS--LPAPPPPPTSSRRLPP 43
A SPPP T PSP A P P +PPPPP +PP
Sbjct: 416 AYPSPPPPVYTSPPPSPVYAYPSPPPPVYSSPPPPPVYEGPIPP 547
Score = 33.9 bits (76), Expect = 0.11
Identities = 16/46 (34%), Positives = 20/46 (42%), Gaps = 7/46 (15%)
Frame = +2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLP-------APPPPPTSSRRLPPP 44
PPP P + PP + P P P +PPP P + PPP
Sbjct: 299 PPPPPPPPCVEPPPPSSPAPHQTPYHPPPSPSPPPSPVYAYPSPPP 436
Score = 33.9 bits (76), Expect = 0.11
Identities = 21/47 (44%), Positives = 23/47 (48%), Gaps = 3/47 (6%)
Frame = +2
Query: 5 SPPPSSPTESIPSPPSAVPLPLSLPAP---PPPPTSSRRLPPPCWTP 48
SPPP + PSPP V P P P PPPP+S P P TP
Sbjct: 242 SPPPPVYSPPPPSPPPCVEPPPPPPPPCVEPPPPSS----PAPHQTP 370
Score = 33.1 bits (74), Expect = 0.19
Identities = 22/45 (48%), Positives = 26/45 (56%), Gaps = 6/45 (13%)
Frame = +2
Query: 6 PPPSSP----TESIPSP-PSAVPLPL-SLPAPPPPPTSSRRLPPP 44
PPPSSP T P P PS P P+ + P+PPPP +S PPP
Sbjct: 335 PPPSSPAPHQTPYHPPPSPSPPPSPVYAYPSPPPPVYTS---PPP 460
Score = 31.6 bits (70), Expect = 0.54
Identities = 16/40 (40%), Positives = 18/40 (45%)
Frame = +2
Query: 5 SPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
SPPP P P PP P + +PPPP PPP
Sbjct: 26 SPPPPPPNSPPPPPP-----PAPVFSPPPPVPYYYSSPPP 130
Score = 31.2 bits (69), Expect = 0.71
Identities = 20/49 (40%), Positives = 21/49 (42%), Gaps = 6/49 (12%)
Frame = +2
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAP---PPPPTS---SRRLPPPCWTP 48
P P S P PP P P PAP PPPP S PPP +P
Sbjct: 2 PTPPYCVRSPPPPPPNSPPPPPPPAPVFSPPPPVPYYYSSPPPPPAHSP 148
Score = 30.8 bits (68), Expect = 0.93
Identities = 19/48 (39%), Positives = 24/48 (49%), Gaps = 16/48 (33%)
Frame = +2
Query: 4 SSPPPSSPTESIPSPPSAV------------PLP----LSLPAPPPPP 35
+SPPPS P + PSPP V P+P +S +PPPPP
Sbjct: 446 TSPPPS-PVYAYPSPPPPVYSSPPPPPVYEGPIPPVFGISYASPPPPP 586
>BQ141932 weakly similar to GP|6523547|emb hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (30%)
Length = 1338
Score = 41.2 bits (95), Expect = 7e-04
Identities = 27/50 (54%), Positives = 27/50 (54%), Gaps = 12/50 (24%)
Frame = -3
Query: 6 PPPSSPTESIPSPP--SAVPLPL--------SLP--APPPPPTSSRRLPP 43
PPP SPT P PP SAVPLPL S P PPPPP S LPP
Sbjct: 415 PPPRSPTVIDPRPPAASAVPLPLRPLYPLRDSPPRRPPPPPPHSPSILPP 266
Score = 38.9 bits (89), Expect = 0.003
Identities = 22/44 (50%), Positives = 23/44 (52%)
Frame = -3
Query: 5 SPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
SPP P S P P A+PLPL P PPPPP P P TP
Sbjct: 253 SPPTPLPLSSSP*P--ALPLPLPPPTPPPPP------PAPTTTP 146
Score = 37.4 bits (85), Expect = 0.010
Identities = 17/39 (43%), Positives = 20/39 (50%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPP+ P S P+ S P P P PP P+ S PPP
Sbjct: 297 PPPTPPPSSPPALVSPPPRPFLFPPPPNQPSPSPSPPPP 181
Score = 37.4 bits (85), Expect = 0.010
Identities = 19/49 (38%), Positives = 25/49 (50%), Gaps = 2/49 (4%)
Frame = -1
Query: 5 SPPPSSPTESIPSPPSAV--PLPLSLPAPPPPPTSSRRLPPPCWTPEET 51
+PPPSSP + PP P P + P+P P P LPPP P ++
Sbjct: 288 TPPPSSPPALVSPPPRPFLFPPPPNQPSPSPSPPPPPPLPPPPPQPPQS 142
Score = 37.0 bits (84), Expect = 0.013
Identities = 20/48 (41%), Positives = 22/48 (45%), Gaps = 9/48 (18%)
Frame = -2
Query: 5 SPPPSSPTESIPSP---------PSAVPLPLSLPAPPPPPTSSRRLPP 43
SPPP P P P PS+ LPL+ P PPPPP PP
Sbjct: 308 SPPPPPPLPLHPPPQLSSLPPHAPSSFLLPLTSPPPPPPPPHPPPSPP 165
Score = 36.6 bits (83), Expect = 0.017
Identities = 20/42 (47%), Positives = 23/42 (54%)
Frame = -2
Query: 2 ATSSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPP 43
+++ PPP P P PPS P P LP PPP SS LPP
Sbjct: 362 SSTPPPPFVPIAGFPPPPSPPP-PPPLPLHPPPQLSS--LPP 246
Score = 35.0 bits (79), Expect = 0.049
Identities = 18/34 (52%), Positives = 20/34 (57%)
Frame = -1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSR 39
PPP+ P+ S PSPP PLP PPP P SR
Sbjct: 225 PPPNQPSPS-PSPPPPPPLP----PPPPQPPQSR 139
Score = 32.3 bits (72), Expect = 0.32
Identities = 19/40 (47%), Positives = 21/40 (52%)
Frame = -3
Query: 4 SSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPP 43
S+PPP T + PPSA P P LP PP PT PP
Sbjct: 481 SAPPP---TRAFALPPSAPP-PSDLPPPPRSPTVIDPRPP 374
Score = 31.6 bits (70), Expect = 0.54
Identities = 45/170 (26%), Positives = 59/170 (34%), Gaps = 11/170 (6%)
Frame = -2
Query: 7 PPSS----PTESIPSPPSAVPLPLSLPAPP---PPPTSSR----RLPPPCWTPEETSALI 55
PPS+ P P PP P LPAPP PPT +R L PP P + +
Sbjct: 578 PPSTTTLTPQHPNPPPP*TDPPATPLPAPPHPVRPPTHARLRSPPLRPPTLRPPPPAPVA 399
Query: 56 DSYRDKWYSLGRTNLKATHWQEVADAVAVRCPNSSPVAKTAVQCRHKMEKLRKRYRSEIQ 115
+R S R +T P S P
Sbjct: 398 HGHRP---SPPRRIGSSTPPPPFVPIAGFPPPPSPPPPPPLP-------------LHPPP 267
Query: 116 KLRSLPVPRSRSSSSWVLFKAMDSMEKGPSPPSPPHKPENPNHNANHNHI 165
+L SLP + SS++L + P PP PPH P +P NH ++
Sbjct: 266 QLSSLP---PHAPSSFLL-----PLTSPPPPPPPPHPPPSPPRPHNHPNL 141
Score = 31.6 bits (70), Expect = 0.54
Identities = 18/49 (36%), Positives = 21/49 (42%), Gaps = 4/49 (8%)
Frame = -3
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPP----PPPTSSRRLPPPCWTPEE 50
PPP P+ IP P P PP PPPT + LPP P +
Sbjct: 568 PPP*PPSIPIPRHPEPTHQPHPCLPPPIPSAPPPTRAFALPPSAPPPSD 422
Score = 31.2 bits (69), Expect = 0.71
Identities = 22/60 (36%), Positives = 28/60 (46%), Gaps = 8/60 (13%)
Frame = -2
Query: 3 TSSPPPSSPTE--SIPSPP----SAVPLPLSLPAP--PPPPTSSRRLPPPCWTPEETSAL 54
T PPP +P PSPP S+ P P +P PPPP+ P P P + S+L
Sbjct: 431 TLRPPPPAPVAHGHRPSPPRRIGSSTPPPPFVPIAGFPPPPSPPPPPPLPLHPPPQLSSL 252
Score = 31.2 bits (69), Expect = 0.71
Identities = 14/34 (41%), Positives = 16/34 (46%)
Frame = -1
Query: 11 PTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
P ++P PP P P S PA PP PPP
Sbjct: 321 PPPAVPPPPPPTPPPSSPPALVSPPPRPFLFPPP 220
Score = 31.2 bits (69), Expect = 0.71
Identities = 23/53 (43%), Positives = 26/53 (48%), Gaps = 10/53 (18%)
Frame = -1
Query: 2 ATSSPPPSSPTESIPS--------PPSAVPLPLSLPAPPPPPTSSRRL--PPP 44
+T +PPP S PS PP AVP P P P PPP+S L PPP
Sbjct: 390 STLAPPPHRQFHS-PSALCTHCGIPPPAVPPP---PPPTPPPSSPPALVSPPP 244
Score = 30.8 bits (68), Expect = 0.93
Identities = 17/41 (41%), Positives = 19/41 (45%)
Frame = -1
Query: 4 SSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
S PPP S+ PP P P + P P P SS PPP
Sbjct: 486 SRPPPHPRAPSLSPPPP--PHPPTSPPRPGRPRSSTLAPPP 370
Score = 30.0 bits (66), Expect = 1.6
Identities = 20/58 (34%), Positives = 24/58 (40%), Gaps = 12/58 (20%)
Frame = -1
Query: 3 TSSPPPSSPTESIPSPP--------SAVPLPLSLPA----PPPPPTSSRRLPPPCWTP 48
TS P P P S +PP SA+ +P PPPPPT PP +P
Sbjct: 423 TSPPRPGRPRSSTLAPPPHRQFHSPSALCTHCGIPPPAVPPPPPPTPPPSSPPALVSP 250
Score = 27.7 bits (60), Expect = 7.9
Identities = 19/51 (37%), Positives = 22/51 (42%)
Frame = -2
Query: 8 PSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPEETSALIDSY 58
PSS S + P LP P PPPP + L P TP T L S+
Sbjct: 1178 PSSRR*SSSAQPRTRSLPYH-PRPPPPLATDHTLLSPRPTPRATHPLSPSH 1029
Score = 27.7 bits (60), Expect = 7.9
Identities = 13/29 (44%), Positives = 13/29 (44%)
Frame = -2
Query: 16 PSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
P PPS L P PPPP T P P
Sbjct: 584 PHPPSTTTLTPQHPNPPPP*TDPPATPLP 498
>BQ751144 weakly similar to PIR|PQ0479|PQ04 pistil extensin-like protein
(clone pMG14) - common tobacco (fragment), partial (11%)
Length = 632
Score = 41.2 bits (95), Expect = 7e-04
Identities = 26/55 (47%), Positives = 29/55 (52%), Gaps = 15/55 (27%)
Frame = +1
Query: 5 SPPPSSPTESIPSPPSA----VPLPLSLP-----------APPPPPTSSRRLPPP 44
SPPP SP+ S P PPS +PLPL LP +PPP P SS L PP
Sbjct: 316 SPPPPSPSRSRPPPPSTPSPLLPLPL-LPLATEPSPFPPSSPPPRPASSPSLSPP 477
Score = 38.9 bits (89), Expect = 0.003
Identities = 18/44 (40%), Positives = 25/44 (55%)
Frame = +1
Query: 1 MATSSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
+ + + PP++P S PS+ P SL +PPPP S R PPP
Sbjct: 229 LRSPTAPPAAPLSPPASSPSSPTRPSSLFSPPPPSPSRSRPPPP 360
Score = 38.1 bits (87), Expect = 0.006
Identities = 25/60 (41%), Positives = 30/60 (49%), Gaps = 15/60 (25%)
Frame = +1
Query: 4 SSPPPSSPTESI----PSPPSAVPL--PLSLPAPP---------PPPTSSRRLPPPCWTP 48
SSPP +SP+ P+ P A PL P S P+ P PPP+ SR PPP TP
Sbjct: 190 SSPPSTSPSPHALLRSPTAPPAAPLSPPASSPSSPTRPSSLFSPPPPSPSRSRPPPPSTP 369
Score = 31.6 bits (70), Expect = 0.54
Identities = 18/49 (36%), Positives = 22/49 (44%)
Frame = +1
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPEETSAL 54
PP S P SP + P+ L P+ PPP S R PP P +L
Sbjct: 424 PPSSPPPRPASSPSLSPPVLLFPPSASPPPWSPSR-PPSALAPPSRPSL 567
Score = 31.2 bits (69), Expect = 0.71
Identities = 18/46 (39%), Positives = 23/46 (49%)
Frame = +1
Query: 3 TSSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
++SPPP SP+ PPSA+ P P PP+S R P P
Sbjct: 496 SASPPPWSPSR----PPSALAPPSRPSLPASPPSSPRSSTRPSRFP 621
Score = 30.8 bits (68), Expect = 0.93
Identities = 24/48 (50%), Positives = 28/48 (58%), Gaps = 5/48 (10%)
Frame = +1
Query: 2 ATSSPPPSSPTESIPSPPSAV--PLPLSLPA---PPPPPTSSRRLPPP 44
A SPP SSP S P+ PS++ P P S P+ PPPP T S LP P
Sbjct: 256 APLSPPASSP--SSPTRPSSLFSPPPPS-PSRSRPPPPSTPSPLLPLP 390
Score = 29.6 bits (65), Expect = 2.1
Identities = 23/51 (45%), Positives = 24/51 (46%), Gaps = 10/51 (19%)
Frame = +1
Query: 4 SSPPPS---SPTESIP---SPPSAVPLPLSLPAPP----PPPTSSRRLPPP 44
SSPPP SP+ S P PPSA P P S PP PP S PP
Sbjct: 430 SSPPPRPASSPSLSPPVLLFPPSASPPPWSPSRPPSALAPPSRPSLPASPP 582
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.310 0.129 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,264,298
Number of Sequences: 36976
Number of extensions: 315770
Number of successful extensions: 16988
Number of sequences better than 10.0: 848
Number of HSP's better than 10.0 without gapping: 5653
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 11659
length of query: 388
length of database: 9,014,727
effective HSP length: 98
effective length of query: 290
effective length of database: 5,391,079
effective search space: 1563412910
effective search space used: 1563412910
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC149129.9


