

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149129.7 + phase: 0
(726 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
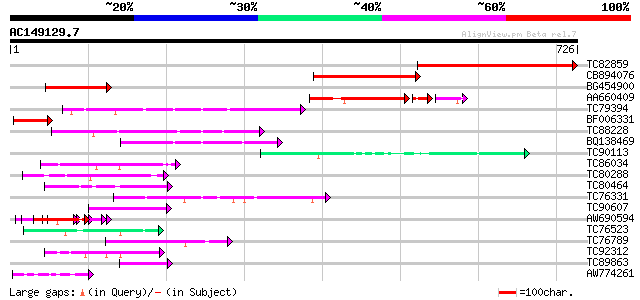
Score E
Sequences producing significant alignments: (bits) Value
TC82859 weakly similar to GP|20147226|gb|AAM10328.1 At2g44720/F1... 411 e-115
CB894076 similar to GP|20147226|gb| At2g44720/F16B22.21 {Arabido... 278 4e-75
BG454900 weakly similar to PIR|T39903|T399 serine-rich protein -... 176 2e-44
AA660409 weakly similar to GP|22655529|gb| water-stress protein ... 135 4e-38
TC79394 weakly similar to PIR|T49019|T49019 probable RNA binding... 130 2e-30
BF006331 homologue to GP|20147226|gb| At2g44720/F16B22.21 {Arabi... 96 4e-20
TC88228 similar to GP|13605916|gb|AAK32943.1 AT4g00830/A_TM018A1... 94 1e-19
BQ138469 weakly similar to PIR|T49019|T49 probable RNA binding p... 91 1e-18
TC90113 similar to PIR|C86301|C86301 arginine/serine-rich protei... 67 2e-11
TC86034 similar to GP|6815067|dbj|BAA90354.1 ESTs AU082210(C5365... 62 8e-10
TC80288 weakly similar to PIR|T06029|T06029 hypothetical protein... 60 2e-09
TC80464 weakly similar to GP|13548328|emb|CAC35875. putative pro... 60 3e-09
TC76331 similar to GP|7673355|gb|AAF66823.1| poly(A)-binding pro... 55 1e-07
TC90607 similar to GP|20466724|gb|AAM20679.1 unknown protein {Ar... 54 2e-07
AW690594 similar to GP|23498163|emb hypothetical protein {Plasmo... 53 5e-07
TC76523 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - ... 53 5e-07
TC76789 similar to GP|15294254|gb|AAK95304.1 AT4g24770/F22K18_30... 52 6e-07
TC92312 weakly similar to GP|4567275|gb|AAD23688.1| expressed pr... 52 1e-06
TC89863 similar to PIR|T03986|T03986 transformer-SR ribonucleopr... 51 1e-06
AW774261 weakly similar to EGAD|146423|156 vitellogenin {Anolis ... 51 1e-06
>TC82859 weakly similar to GP|20147226|gb|AAM10328.1 At2g44720/F16B22.21
{Arabidopsis thaliana}, partial (15%)
Length = 975
Score = 411 bits (1057), Expect = e-115
Identities = 202/205 (98%), Positives = 203/205 (98%), Gaps = 1/205 (0%)
Frame = +2
Query: 523 SQRGPSYRDYPARDSGYPDLHRDTSRTAPRRGYLDDGYDRRLERPPSPSPRLSYREGRPR 582
+Q GPSYRDYPARDSGYPDLHRDTSRTAPRRGYLDDGYDRRLERPPSPSPRLSYREGRPR
Sbjct: 14 AQLGPSYRDYPARDSGYPDLHRDTSRTAPRRGYLDDGYDRRLERPPSPSPRLSYREGRPR 193
Query: 583 DYDALSGSKRPYAAIDDISPRYADAGARQSRSRLDYDYGGSASRYREAYGD-RLERSSLG 641
DYDALSGSKRPYAAIDDISPRYADAGARQSRSRLDYDYGGSASRYREAYGD RLERSSLG
Sbjct: 194 DYDALSGSKRPYAAIDDISPRYADAGARQSRSRLDYDYGGSASRYREAYGDSRLERSSLG 373
Query: 642 YSGSRSSLSNQNSHGAYSSRQDPSYGRGSLGGSDGGIYSSSYGGDYLSRGSDVGGSSYSS 701
YSGSRSSLSNQNSHGAYSSRQDPSYGRGSLGGSDGGIYSSSYGGDYLSRGSDVGGSSYSS
Sbjct: 374 YSGSRSSLSNQNSHGAYSSRQDPSYGRGSLGGSDGGIYSSSYGGDYLSRGSDVGGSSYSS 553
Query: 702 MYSSRGMNGSSSYMSGSGGGSRSYY 726
MYSSRGMNGSSSYMSGSGGGSRSYY
Sbjct: 554 MYSSRGMNGSSSYMSGSGGGSRSYY 628
>CB894076 similar to GP|20147226|gb| At2g44720/F16B22.21 {Arabidopsis
thaliana}, partial (3%)
Length = 416
Score = 278 bits (712), Expect = 4e-75
Identities = 138/138 (100%), Positives = 138/138 (100%)
Frame = +3
Query: 389 KKAKVRARLSRPLPKVRGKHVPHRDISGRKIGRLERPSRSRPAPRSRPAPQSRPAHVFRR 448
KKAKVRARLSRPLPKVRGKHVPHRDISGRKIGRLERPSRSRPAPRSRPAPQSRPAHVFRR
Sbjct: 3 KKAKVRARLSRPLPKVRGKHVPHRDISGRKIGRLERPSRSRPAPRSRPAPQSRPAHVFRR 182
Query: 449 LGSRIPPARPSSARNRRPVTSIPVRARPASPPARSYRRLAAAYPKSSMKRDYGRPVDLPP 508
LGSRIPPARPSSARNRRPVTSIPVRARPASPPARSYRRLAAAYPKSSMKRDYGRPVDLPP
Sbjct: 183 LGSRIPPARPSSARNRRPVTSIPVRARPASPPARSYRRLAAAYPKSSMKRDYGRPVDLPP 362
Query: 509 PRSRVSADYGSQVVSQRG 526
PRSRVSADYGSQVVSQRG
Sbjct: 363 PRSRVSADYGSQVVSQRG 416
>BG454900 weakly similar to PIR|T39903|T399 serine-rich protein - fission
yeast (Schizosaccharomyces pombe), partial (12%)
Length = 258
Score = 176 bits (447), Expect = 2e-44
Identities = 85/85 (100%), Positives = 85/85 (100%)
Frame = +3
Query: 46 DDYDERGIEQDEGQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGD 105
DDYDERGIEQDEGQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGD
Sbjct: 3 DDYDERGIEQDEGQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGD 182
Query: 106 DEHASEENVHEHLADVKEEEQVEVI 130
DEHASEENVHEHLADVKEEEQVEVI
Sbjct: 183 DEHASEENVHEHLADVKEEEQVEVI 257
>AA660409 weakly similar to GP|22655529|gb| water-stress protein {Zea mays},
partial (16%)
Length = 644
Score = 135 bits (340), Expect(3) = 4e-38
Identities = 80/135 (59%), Positives = 89/135 (65%), Gaps = 7/135 (5%)
Frame = +3
Query: 384 LGEGNKKAKVRARLSRPLPKVRGKHVPHRDISGRKIGRLERPSR-----SRPAPRSRPAP 438
LGEG+KKAKVRA LSRPL + RGKHV D RPSR SRP+ RPAP
Sbjct: 6 LGEGDKKAKVRAXLSRPLQRGRGKHVGRGDY---------RPSRGSAIMSRPSWSXRPAP 158
Query: 439 QSRPAHVFRRLGSRIPPARPSSARNRRPVTSIPVRARPASPPARSY--RRLAAAYPKSSM 496
+S + RR+GSR PP R AR+RRPV SIPVR+RP PPARSY R AAY KSS+
Sbjct: 159 RSFSSRGVRRIGSRAPPIRSIXARDRRPVMSIPVRSRPLPPPARSYDRRPADAAYSKSSV 338
Query: 497 KRDYGRPVDLPPPRS 511
K DYGR DLPPPRS
Sbjct: 339 KXDYGRRDDLPPPRS 383
Score = 34.7 bits (78), Expect(3) = 4e-38
Identities = 19/45 (42%), Positives = 26/45 (57%), Gaps = 4/45 (8%)
Frame = +1
Query: 546 TSRTAPRRGYLDDGYDRRLERPPSPS----PRLSYREGRPRDYDA 586
T+R AP + Y+DDGY R E+ S S P S RE P +Y++
Sbjct: 481 TARAAPXKSYVDDGYXPRXEKATSSSTPSHPNFSXRE-PPXEYES 612
Score = 27.3 bits (59), Expect(3) = 4e-38
Identities = 12/26 (46%), Positives = 17/26 (65%)
Frame = +2
Query: 516 DYGSQVVSQRGPSYRDYPARDSGYPD 541
+YGS++ + + P YRDYP S Y D
Sbjct: 395 NYGSRM-TLKDPXYRDYPPXGSDYSD 469
>TC79394 weakly similar to PIR|T49019|T49019 probable RNA binding protein -
Arabidopsis thaliana, partial (74%)
Length = 1909
Score = 130 bits (327), Expect = 2e-30
Identities = 96/321 (29%), Positives = 161/321 (49%), Gaps = 10/321 (3%)
Frame = +2
Query: 68 EPEENVDEEE-----GDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVK 122
EPE+ D E GD EE E E YEEVE +++V ++E EE V E +
Sbjct: 203 EPEKPTDSGEQVDLDGDNDQEESSEEEVEYEEVE-VEEEVEEEEEEEEEEEVEEESKPLD 379
Query: 123 EEEQVEVIKETQ----KQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKR 178
EE++ + K + EV++GG+ E +E DLR VGE+ EVR+ + K
Sbjct: 380 EEDEADKKKHAELLALPPHGSEVYIGGIPHETSEKDLRVFCQSVGEVAEVRVM---KGKE 550
Query: 179 NKGFALLRFETVEHVKRALAELKNPVINGKQCGITITTCQDSDTLYLDNICKSWKKEALK 238
KG+A + F+T E AL EL N G++ I + Q L++ ++ K W E +K
Sbjct: 551 AKGYAFVTFKTKELASNALKELNNSEFKGRK--IKCSPSQVKHRLFIGSVPKEWTVEDMK 724
Query: 239 EKLKHYGVESFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDVVFGVD-KP 297
+ + G + LL+D + N G AF+E+ +H+ ++ Y R + ++ F +D
Sbjct: 725 KVVAKVG-PGVISVELLKDPQSSSRNRGFAFIEYHNHACAE--YSRQKMSNSNFKLDNND 895
Query: 298 AEVSFANSFIDLGDDIMAQVKTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGAR 357
A VS+A+ + +QVK V++ LP + ++ ++ L + +G + KV L G
Sbjct: 896 AIVSWADP-RNSESSSSSQVKAVYVKNLPENITQNRLKELFEHHGKITKVALPPAKAGQE 1072
Query: 358 RKNYGFVTFGTHAAAVECAES 378
+ YGFV F ++A++ ++
Sbjct: 1073KSRYGFVHFADRSSAMKALKN 1135
Score = 29.3 bits (64), Expect = 5.6
Identities = 18/63 (28%), Positives = 28/63 (43%)
Frame = +2
Query: 320 VFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAAAVECAESI 379
V+I +P E +R + G V +V + K G K Y FVTF T A + +
Sbjct: 446 VYIGGIPHETSEKDLRVFCQSVGEVAEVRVMK---GKEAKGYAFVTFKTKELASNALKEL 616
Query: 380 TSA 382
++
Sbjct: 617 NNS 625
>BF006331 homologue to GP|20147226|gb| At2g44720/F16B22.21 {Arabidopsis
thaliana}, partial (1%)
Length = 468
Score = 96.3 bits (238), Expect = 4e-20
Identities = 42/50 (84%), Positives = 46/50 (92%)
Frame = +1
Query: 5 IYLIADNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIE 54
+ ++ +NGEEVKESI EYEKDEHLDFEDNYHEYDPEEYVGDDDYDER IE
Sbjct: 319 VAMVENNGEEVKESIHEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERRIE 468
>TC88228 similar to GP|13605916|gb|AAK32943.1 AT4g00830/A_TM018A10_14
{Arabidopsis thaliana}, partial (67%)
Length = 1712
Score = 94.4 bits (233), Expect = 1e-19
Identities = 68/280 (24%), Positives = 143/280 (50%), Gaps = 7/280 (2%)
Frame = +1
Query: 54 EQDEGQEAGDEVEEE-PEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGD----DEH 108
E++ +E D+VEE+ ++ VD E + + VEE EY E D + D
Sbjct: 196 EENYMEEMDDDVEEQIDDDGVDGGEDENAEGSVEEHEYEETAAEAGQKDQFPEGEKSDHG 375
Query: 109 ASEENVHEHLADVKEEEQVEVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEV 168
A E+ + L D +E E+ + + EVF+GGL ++ ++ D+R++ +G+I E+
Sbjct: 376 AEEDELKPALIDEEEREKHDELLSRPPHGS-EVFIGGLPRDTSDDDVRELCEPMGDIVEI 552
Query: 169 RMTVNPQTKRNKGFALLRFETVEHVKRALAELKNPVINGKQCGITITTCQDSDTLYLDNI 228
++ + +T +KG+A + ++T E ++A+ ++ N GK ++ + L++ NI
Sbjct: 553 KLIKDRETGESKGYAFVGYKTKEVAQKAIDDIHNKEFKGKTLRCLLS--ETKHRLFIGNI 726
Query: 229 CKSWKKEALKEKLKHYGVESFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKT 288
K+W ++ ++ ++ G + + L++D N+ N G AF+ + ++++ + R + +
Sbjct: 727 PKTWTEDEFRKAVEGVG-PGVESIDLIKDPQNQSRNRGFAFVLY--YNNACADFSRQKMS 897
Query: 289 DVVFGVDK-PAEVSFANSFIDLGDDIMA-QVKTVFIDLLP 326
V F +D V++A+ A QVK +++ +P
Sbjct: 898 SVGFKLDGITPTVTWADPKTSPDQSAAASQVKALYVKNIP 1017
>BQ138469 weakly similar to PIR|T49019|T49 probable RNA binding protein -
Arabidopsis thaliana, partial (42%)
Length = 633
Score = 91.3 bits (225), Expect = 1e-18
Identities = 56/207 (27%), Positives = 108/207 (52%)
Frame = +2
Query: 143 VGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAELKN 202
VGG+ +A DL++ ++GE+ +VR+ NKGFA + + VE +A+ EL N
Sbjct: 2 VGGIPLDAKSEDLKEFCERIGEVVQVRIMKGKDASENKGFAFVTYRNVELASKAIEELNN 181
Query: 203 PVINGKQCGITITTCQDSDTLYLDNICKSWKKEALKEKLKHYGVESFKDLTLLEDDNNEG 262
G++ I +T Q + L++ NI +SW ++ LK+ + G + L++D N
Sbjct: 182 TEFKGRK--IKCSTSQAKNRLFIGNIPRSWGEKDLKKVVTDIG-PGVTAVELVKDMKNIS 352
Query: 263 TNCGCAFLEFSSHSDSKDAYKRLQKTDVVFGVDKPAEVSFANSFIDLGDDIMAQVKTVFI 322
N G AF+++ ++ ++ +++ G + P VS+A+ + +QVK V++
Sbjct: 353 NNRGFAFIDYHNNQCAEYGRQKMMSPSFKLGDNSPT-VSWADP-KNSDSSASSQVKAVYV 526
Query: 323 DLLPPSWDEDYVRALLKKYGAVEKVEL 349
LP + ++ ++ L + +G + KV L
Sbjct: 527 KNLPKNVTQEQLKKLFEHHGKITKVVL 607
>TC90113 similar to PIR|C86301|C86301 arginine/serine-rich protein
[imported] - Arabidopsis thaliana, partial (70%)
Length = 1259
Score = 67.0 bits (162), Expect = 2e-11
Identities = 96/351 (27%), Positives = 133/351 (37%), Gaps = 7/351 (1%)
Frame = +1
Query: 322 IDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAAAVECAESITS 381
++ L + +E +++ + +G V VEL + K YG+V F T A + +
Sbjct: 274 VEKLSRNVNEGHLKEIFSNFGEVVSVELVMDRAVNLPKGYGYVHFKTRGEAEKALLYMDG 453
Query: 382 AGLGEGNKKAKV----RARLSRPLPKVRGKH-VPHRDISGRKIGR--LERPSRSRPAPRS 434
A + KA+ R + S P V K P D +G ++ + +RP S P +
Sbjct: 454 AQIDGNVVKARFTLPPRQKASPPPKAVAPKRDAPRTDNAGAEVEKDGPKRPRESSPRRKP 633
Query: 435 RPAPQSRPAHVFRRLGSRIPPARPSSARNRRPVTSIPVRARPASPPARSYRRLAAAYPKS 494
P+P+ R + V RR GS P RP S R R PVR R SP YRR A
Sbjct: 634 PPSPRRR-SPVPRRAGS---PRRPESPRRR---ADSPVRRRLDSP----YRRGAG----- 765
Query: 495 SMKRDYGRPVDLPPPRSRVSADYGSQVVSQRGPSYRDYPARDSGYPDLHRDTSRTAPRRG 554
D PP R VS G R P S P R +R +PRR
Sbjct: 766 ----------DTPPRRRPVSPGRG------RSP---------SPPPRRLRSPARVSPRRM 870
Query: 555 YLDDGYDRRLERPPSPSPRLSYREGRPRDYDALSGSKRPYAAIDDISPRYADAGARQSRS 614
G RR PP P R R R + G +R + I S R R
Sbjct: 871 RGSPGPGRRRSPPPPPRRRSPPRRARSPPRRSPVGRRRSRSPIRR-SARSRSRSFSPRRG 1047
Query: 615 RLDYDYGGSASRYREAYGDRLERSSLGYSGSRSSLSNQNSHGAYSSRQDPS 665
R G S+S ++ R S S R +S+ + SS P+
Sbjct: 1048RPPMRRGRSSSYSDSPSPRKVSRRSKSRSPRRPLKGRASSNSSSSSSPPPA 1200
Score = 37.7 bits (86), Expect = 0.016
Identities = 19/77 (24%), Positives = 38/77 (48%)
Frame = +1
Query: 131 KETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETV 190
K + ++ L + V L + E L+++FS GE+ V + ++ KG+ + F+T
Sbjct: 238 KPSPVRESLVLHVEKLSRNVNEGHLKEIFSNFGEVVSVELVMDRAVNLPKGYGYVHFKTR 417
Query: 191 EHVKRALAELKNPVING 207
++AL + I+G
Sbjct: 418 GEAEKALLYMDGAQIDG 468
>TC86034 similar to GP|6815067|dbj|BAA90354.1 ESTs AU082210(C53655)
AU056546(S20671) AU056545(S20671) correspond to a
region of the predicted, partial (42%)
Length = 2412
Score = 62.0 bits (149), Expect = 8e-10
Identities = 62/217 (28%), Positives = 92/217 (41%), Gaps = 38/217 (17%)
Frame = +3
Query: 40 EEYVGDDDYDERGIEQDEGQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDD 99
EE + + IE+D+ Q G E EEEPEE V EEE D E E +E ++ G+
Sbjct: 183 EEEPTTQEEENTPIEEDQPQNEG-EGEEEPEEEVAEEEEDDQQGEDETLENQQQQDGGEA 359
Query: 100 DDVAGDDEHA--------SEENVHEHLADVK-EEEQVEVIKETQKQKEL----------- 139
+ A E E N E D++ E+E VE + E +++L
Sbjct: 360 SNNAAVSEQTVTVEANGGGENNNGEEEEDLELEDEPVEKLLEPFTKEQLHALVKQAVEKY 539
Query: 140 ------------------EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKG 181
++FV GL +AT L FSK GEI + + + + ++KG
Sbjct: 540 PDLSENVRQLADVDPAHRKIFVHGLGWDATAETLTTTFSKYGEIEDCKAVTDKASGKSKG 719
Query: 182 FALLRFETVEHVKRALAELKNPVINGKQCGITITTCQ 218
+A + F+ +RA LK P K G T+CQ
Sbjct: 720 YAFILFKHRTGCRRA---LKQP---QKVIGNRTTSCQ 812
Score = 40.8 bits (94), Expect = 0.002
Identities = 22/68 (32%), Positives = 37/68 (54%)
Frame = +3
Query: 140 EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAE 199
++FV + + L + F + GE+ + + ++ T + KGFAL ++TVE K+AL E
Sbjct: 882 KIFVSNVSSDIDPQKLLEFFKQFGEVDDGPLGLDKATGKPKGFALFVYKTVESAKKALEE 1061
Query: 200 LKNPVING 207
N V G
Sbjct: 1062-PNKVFEG 1082
>TC80288 weakly similar to PIR|T06029|T06029 hypothetical protein T28I19.100
- Arabidopsis thaliana, partial (14%)
Length = 1460
Score = 60.5 bits (145), Expect = 2e-09
Identities = 53/192 (27%), Positives = 86/192 (44%), Gaps = 5/192 (2%)
Frame = +3
Query: 17 ESIDEYEKDEHLDFE-DNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEENVDE 75
E +E E+DEH+ + N E+D ++ G E G + + +E+ D V E EE DE
Sbjct: 393 EGHEEEEEDEHIVYNMQNKREHDEQQQEG-----EEGNKHETEEESEDNVHERREE-QDE 554
Query: 76 EEGDTGDEEVEEVEYVYEEVEGDDDDV---AGDDEHASEENVHE-HLADVKEEEQVEVIK 131
EE G E EE E EEVE + DV D E + +N E + D +++++ E
Sbjct: 555 EENKHGAEVQEENESKSEEVEDEGGDVEIDENDHEKSEADNDREDEVVDEEKDKEEEGDD 734
Query: 132 ETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVE 191
ET+ + + + GGL + H+ R+ K + + +T G +E
Sbjct: 735 ETENEDKEDEEKGGLVENHENHEAREEHYKADDASSAVAHDTHETSTETG-------NLE 893
Query: 192 HVKRALAELKNP 203
H +L + P
Sbjct: 894 HSDLSLQNTRKP 929
Score = 40.0 bits (92), Expect = 0.003
Identities = 37/144 (25%), Positives = 62/144 (42%), Gaps = 1/144 (0%)
Frame = +3
Query: 13 EEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEEN 72
+E ES E +DE D E + ++++ E D + + E+D+ +E DE E E +E
Sbjct: 582 QEENESKSEEVEDEGGDVEIDENDHEKSEADNDREDEVVDEEKDKEEEGDDETENEDKE- 758
Query: 73 VDEEEGD-TGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEVIK 131
DEE+G + E E + + + VA D S E + +D+ + +
Sbjct: 759 -DEEKGGLVENHENHEAREEHYKADDASSAVAHDTHETSTETGNLEHSDLSLQNTRKPEN 935
Query: 132 ETQKQKELEVFVGGLDKEATEHDL 155
ET E D + TE +L
Sbjct: 936 ETNHSDESYGSQNVSDLKVTEGEL 1007
>TC80464 weakly similar to GP|13548328|emb|CAC35875. putative protein
{Arabidopsis thaliana}, partial (19%)
Length = 1437
Score = 60.1 bits (144), Expect = 3e-09
Identities = 41/164 (25%), Positives = 70/164 (42%)
Frame = +3
Query: 45 DDDYDERGIEQDEGQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAG 104
DDD + D+ ++ E E + T +E++ + ++ DVA
Sbjct: 744 DDDSSDSASSDDDDKDKHSHATEHEENRCNNPSETTPRSGAQELD-LEDQKNTVGKDVAN 920
Query: 105 DDEHASEENVHEHLADVKEEEQVEVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGE 164
D S+ N E + E + + E + +FV L TE +L + FS+ G
Sbjct: 921 DK---SQVNATEEEGQLSNPEDKKGVSEPCR-----LFVRNLPYSTTEEELEEHFSQFGS 1076
Query: 165 ITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAELKNPVINGK 208
+++ + V+ TKR+KG A + F E RAL E N + G+
Sbjct: 1077VSQAHLVVDKDTKRSKGIAYIHFSAPEFAARALEESDNSIFQGR 1208
>TC76331 similar to GP|7673355|gb|AAF66823.1| poly(A)-binding protein
{Nicotiana tabacum}, partial (97%)
Length = 2581
Score = 54.7 bits (130), Expect = 1e-07
Identities = 63/306 (20%), Positives = 125/306 (40%), Gaps = 29/306 (9%)
Frame = +3
Query: 134 QKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHV 193
+K + +F+ LDK L FS G I ++ V+ + ++KG+ ++F+T E
Sbjct: 693 RKSGQGNIFIKNLDKAIDHKALHDTFSSFGNILSCKVAVDG-SGQSKGYGFVQFDTEEAA 869
Query: 194 KRALAELKNPVINGKQCGI-TITTCQDSDT---------LYLDNICKSWKKEALKEKLKH 243
++A+ +L ++N KQ + Q+ ++ +++ N+ +S + LK+
Sbjct: 870 QKAIEKLNGMLLNDKQVYVGPFLRKQERESTGDRAKFNNVFVKNLSESTTDDELKKTFGE 1049
Query: 244 YGVESFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRL------QKTDVVFGVDKP 297
+G + ++ D + + G F+ F S D+ A + L K V K
Sbjct: 1050FG--TITSAVVMRDGDGKSKCFG--FVNFESTDDAARAVEALNGKKIDDKEWYVGKAQKK 1217
Query: 298 AE------VSFANSFIDLGDDIMAQVKTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAK 351
+E + F S + D Q +++ L S ++ ++ L YG + ++ +
Sbjct: 1218SEREHELKIKFEQSMKEAADKY--QGANLYVKNLDDSIADEKLKELFSSYGTITSCKVMR 1391
Query: 352 NMPGARRKNYGFVTFGTHAAAVECAESITSAGLGE-------GNKKAKVRARLSRPLPKV 404
+ G R + GFV F T A + + +K RARL ++
Sbjct: 1392DPNGVSRGS-GFVAFSTPEEASRALLEMNGKMVASKPLYVTLAQRKEDRRARLQAQFAQM 1568
Query: 405 RGKHVP 410
R +P
Sbjct: 1569RPVSMP 1586
>TC90607 similar to GP|20466724|gb|AAM20679.1 unknown protein {Arabidopsis
thaliana}, partial (49%)
Length = 1143
Score = 54.3 bits (129), Expect = 2e-07
Identities = 33/108 (30%), Positives = 52/108 (47%), Gaps = 2/108 (1%)
Frame = +1
Query: 102 VAGDDEHASEENVHE--HLADVKEEEQVEVIKETQKQKELEVFVGGLDKEATEHDLRKVF 159
V DD SE +E +L + + K ++FV GL +E LR F
Sbjct: 460 VQPDDNFESENKDYEGRNLENSLNMPNSSEASQEASVKTKKLFVTGLSFYTSEKTLRAAF 639
Query: 160 SKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAELKNPVING 207
GE+ EV++ ++ +KR+KG+A + + T E AL E+ +ING
Sbjct: 640 EGFGELVEVKVIIDKISKRSKGYAFIEYTTEEAASAALKEMNGKIING 783
>AW690594 similar to GP|23498163|emb hypothetical protein {Plasmodium
falciparum 3D7}, partial (10%)
Length = 633
Score = 52.8 bits (125), Expect = 5e-07
Identities = 32/78 (41%), Positives = 44/78 (56%), Gaps = 1/78 (1%)
Frame = +3
Query: 31 EDNYHEYDPEEYVGDDDYDERGIEQDE-GQEAGDEVEEEPEENVDEEEGDTGDEEVEEVE 89
E ++ EE+ ++D E E+DE +E DE +EE EE +EEE D D+E EE E
Sbjct: 18 ESSWKRRKVEEHNEEEDEGEVEEEEDEHDEEEEDEHDEEEEEEEEEEEEDEHDDEEEEEE 197
Query: 90 YVYEEVEGDDDDVAGDDE 107
EE E DDD+ +DE
Sbjct: 198 EEEEEEEEDDDEEGEEDE 251
Score = 51.6 bits (122), Expect = 1e-06
Identities = 32/88 (36%), Positives = 49/88 (55%)
Frame = +3
Query: 43 VGDDDYDERGIEQDEGQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDV 102
+ + + R +E+ +E EVEEE +E+ DEEE D DEE EE E EE E DD++
Sbjct: 12 MAESSWKRRKVEEHNEEEDEGEVEEEEDEH-DEEEEDEHDEEEEEEEEEEEEDEHDDEE- 185
Query: 103 AGDDEHASEENVHEHLADVKEEEQVEVI 130
++E EE E + EE++++ I
Sbjct: 186 --EEEEEEEEEEEEDDDEEGEEDEIDRI 263
Score = 48.9 bits (115), Expect = 7e-06
Identities = 25/77 (32%), Positives = 45/77 (57%), Gaps = 1/77 (1%)
Frame = +3
Query: 16 KESIDEY-EKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEENVD 74
+ ++E+ E+++ + E+ E+D EE D+ +E E++E E DE EEE EE +
Sbjct: 33 RRKVEEHNEEEDEGEVEEEEDEHDEEEEDEHDEEEEEEEEEEEEDEHDDEEEEEEEEEEE 212
Query: 75 EEEGDTGDEEVEEVEYV 91
EEE D + E +E++ +
Sbjct: 213 EEEDDDEEGEEDEIDRI 263
Score = 46.2 bits (108), Expect = 4e-05
Identities = 24/72 (33%), Positives = 44/72 (60%)
Frame = +3
Query: 31 EDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEY 90
E N E + E +D++DE ++ + +E +E EEE +E+ DEEE + +EE EE +
Sbjct: 48 EHNEEEDEGEVEEEEDEHDEEEEDEHDEEEEEEEEEEEEDEHDDEEEEEEEEEEEEEED- 224
Query: 91 VYEEVEGDDDDV 102
++ EG++D++
Sbjct: 225 --DDEEGEEDEI 254
Score = 45.4 bits (106), Expect = 8e-05
Identities = 23/81 (28%), Positives = 46/81 (56%)
Frame = +3
Query: 8 IADNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEE 67
+ ++ EE E E E+DEH + E++ H+ + EE +E E+DE + +E EE
Sbjct: 42 VEEHNEEEDEGEVEEEEDEHDEEEEDEHDEEEEE-------EEEEEEEDEHDDEEEEEEE 200
Query: 68 EPEENVDEEEGDTGDEEVEEV 88
E EE ++++ + ++E++ +
Sbjct: 201 EEEEEEEDDDEEGEEDEIDRI 263
Score = 42.7 bits (99), Expect = 5e-04
Identities = 23/78 (29%), Positives = 45/78 (57%), Gaps = 3/78 (3%)
Frame = +3
Query: 49 DERGIEQDEGQ---EAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGD 105
+E E+DEG+ E + EEE +E+ +EEE + +EE +E + EE E ++++ D
Sbjct: 45 EEHNEEEDEGEVEEEEDEHDEEEEDEHDEEEEEEEEEEEEDEHDDEEEEEEEEEEEEEED 224
Query: 106 DEHASEENVHEHLADVKE 123
D+ EE+ + ++ + +
Sbjct: 225 DDEEGEEDEIDRISTLPD 278
>TC76523 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - alfalfa,
partial (69%)
Length = 1615
Score = 52.8 bits (125), Expect = 5e-07
Identities = 44/194 (22%), Positives = 78/194 (39%), Gaps = 14/194 (7%)
Frame = +3
Query: 18 SIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEP---EENVD 74
S + E DE D +++ V + DE ++ + +++P ++ V
Sbjct: 162 SSSDEESDEESDEDEDAKPVSKPAAVAKKSKKDSSDSDDEDDDSSSDEDKKPVAAKKEVS 341
Query: 75 EEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEVIKETQ 134
E E D+ D+E + V+ D D +E + +E +K+ E V+ K +
Sbjct: 342 ESESDSSDDEDKM------NVDKDGSDSDESEEESEDEPSKTPQKKIKDVEMVDAGKSGK 503
Query: 135 KQKELE-----------VFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFA 183
K +FVG L D+ K F GE+ +VR + + + R KGF
Sbjct: 504 KAPNTPATPIENSGSKTLFVGNLSFSVQRSDIEKFFQDCGEVVDVRFS-SDEEGRFKGFG 680
Query: 184 LLRFETVEHVKRAL 197
+ F + E + AL
Sbjct: 681 HVEFASAEAAQSAL 722
>TC76789 similar to GP|15294254|gb|AAK95304.1 AT4g24770/F22K18_30
{Arabidopsis thaliana}, partial (62%)
Length = 1294
Score = 52.4 bits (124), Expect = 6e-07
Identities = 40/176 (22%), Positives = 77/176 (43%), Gaps = 13/176 (7%)
Frame = +3
Query: 123 EEEQVEVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGF 182
E E+VEV + ++L++FVG L + L ++F + G + + N T R++GF
Sbjct: 405 ENEEVEVASFEEPSEDLKIFVGNLPFDVDSEKLAQLFEQSGTVEIAEVIYNRDTDRSRGF 584
Query: 183 ALLRFETVEHVKRALAELKNPVINGKQCGITITTCQDSDTL------------YLDNICK 230
+ T E V+RA+ + ++G+ + + + L Y+ N+
Sbjct: 585 GFVTMSTSEEVERAVNKFSGFELDGRLLTVNKAAPRGTPRLRQPRTFNSGLRAYVGNLPW 764
Query: 231 SWKKEALKEKLKHYG-VESFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRL 285
+L++ +G VES + + D G + G F+ S+ ++ DA L
Sbjct: 765 DVDNSSLEQLFSEHGKVESAQ----VVYDRETGRSRGFGFVTMSNEAEMNDAIAAL 920
Score = 37.7 bits (86), Expect = 0.016
Identities = 17/76 (22%), Positives = 36/76 (47%)
Frame = +3
Query: 139 LEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALA 198
L +VG L + L ++FS+ G++ ++ + +T R++GF + + A+A
Sbjct: 735 LRAYVGNLPWDVDNSSLEQLFSEHGKVESAQVVYDRETGRSRGFGFVTMSNEAEMNDAIA 914
Query: 199 ELKNPVINGKQCGITI 214
L NG+ + +
Sbjct: 915 ALDGQSFNGRAIRVNV 962
>TC92312 weakly similar to GP|4567275|gb|AAD23688.1| expressed protein
{Arabidopsis thaliana}, partial (29%)
Length = 1319
Score = 51.6 bits (122), Expect = 1e-06
Identities = 43/171 (25%), Positives = 76/171 (44%), Gaps = 17/171 (9%)
Frame = +1
Query: 45 DDDYDERGIEQDEGQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEV---EGDDDD 101
+DD + + D+G ++ +++ EE+V E D D+E + V + D+
Sbjct: 601 EDDLENTDKKSDQGDDS--DIDSVVEEDVPSE--DDFDKEADIARKVLNNLITSSAKDES 768
Query: 102 VAGDDEHASEENVHEHLADVK--------EEEQVEVIKETQKQKELE------VFVGGLD 147
V D + E+N + VK E ++V I + + KE E VF+ L
Sbjct: 769 VNNDSVSSEEKNKPKSKETVKGADSKTSKESDKVSDISKPETSKETEDDLHRTVFITNLP 948
Query: 148 KEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALA 198
E +L++ FS GE+ ++ TKR +G L+F+T E A++
Sbjct: 949 FELDTEELKQRFSAFGEVEYFAPVLHQVTKRPRGTGFLKFKTAEAADNAIS 1101
>TC89863 similar to PIR|T03986|T03986 transformer-SR ribonucleoprotein -
common tobacco (fragment), partial (68%)
Length = 1157
Score = 51.2 bits (121), Expect = 1e-06
Identities = 22/68 (32%), Positives = 38/68 (55%)
Frame = +1
Query: 141 VFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAEL 200
++V GL TE DL + FSK G++ + V P+T+ ++GFA + +T E R + L
Sbjct: 472 LYVTGLSSRVTERDLEEHFSKEGKVASCFLVVEPRTRISRGFAFVTMDTAEDANRCIKYL 651
Query: 201 KNPVINGK 208
++ G+
Sbjct: 652 NQSILEGR 675
Score = 32.0 bits (71), Expect = 0.86
Identities = 61/255 (23%), Positives = 90/255 (34%), Gaps = 44/255 (17%)
Frame = +1
Query: 417 RKIGRLERPSRSRPAPRSRPAPQSRPAHVFRRLGSRIPPARPSSARNRRPVTSIP--VRA 474
R R P RP SR RP R L + PP++ S +RP + R
Sbjct: 220 RPRSRSRSPEMQRPRSLSRSPEMQRPRSRSRSLEKQRPPSQSMSREMQRPRSRSRSLERQ 399
Query: 475 RPASPPARSYRRLAAAYPKSSMKRDYGRPVDLPPPRSRVSADYGSQVVSQRGPSYRDYPA 534
RP S P+R+ R + P R+ G + + SRV+ + S+ G +
Sbjct: 400 RPRS-PSRNRGRSRSRSP----DRNGGDTLYVTGLSSRVTERDLEEHFSKEGKVASCFLV 564
Query: 535 RD------SGYPDLHRDTSRTAPR------RGYLDDGY----DRRLERPPSPSPRLSYRE 578
+ G+ + DT+ A R + L+ Y + +RP +P+P
Sbjct: 565 VEPRTRISRGFAFVTMDTAEDANRCIKYLNQSILEGRYITVERSKRKRPRTPTPGHYLGL 744
Query: 579 GRPRDYDALSGS--------------------KRPYAAIDDISPRYADAGA----RQSRS 614
RDY G + PY D SPR++ G R+ RS
Sbjct: 745 KNTRDYGPRGGDRGRHQGGFGRDDYPYHRSPRRSPYRGSRDYSPRHSPYGGSGRYRRDRS 924
Query: 615 RLD--YDYGGSASRY 627
R YG RY
Sbjct: 925 RSPPYAPYGSPXRRY 969
Score = 28.9 bits (63), Expect = 7.3
Identities = 26/85 (30%), Positives = 36/85 (41%), Gaps = 2/85 (2%)
Frame = +1
Query: 421 RLERPSRSRPAPRSRPAPQSRPAHVFRRLGSRIPPARPSSARNRRP--VTSIPVRARPAS 478
R + SRSR +S + RP + R + P +R S +RP ++ P RP S
Sbjct: 124 RAQSRSRSRSRSQSGSLDRHRPRSLSRSPEMQRPRSRSRSPEMQRPRSLSRSPEMQRPRS 303
Query: 479 PPARSYRRLAAAYPKSSMKRDYGRP 503
RS P SM R+ RP
Sbjct: 304 ---RSRSLEKQRPPSQSMSREMQRP 369
>AW774261 weakly similar to EGAD|146423|156 vitellogenin {Anolis pulchellus},
partial (40%)
Length = 495
Score = 51.2 bits (121), Expect = 1e-06
Identities = 42/106 (39%), Positives = 57/106 (53%), Gaps = 2/106 (1%)
Frame = +1
Query: 4 NIYLIADNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGD 63
N L+A E E E DE +D E+ E DPEE + + +Y+E +E++E E
Sbjct: 223 NCSLVAKKSEPEPEKT--VESDEKIDLEE---ENDPEEEMEEIEYEE--VEEEEEVE--- 372
Query: 64 EVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDD--DDVAGDDE 107
E+EEE EEN ++ E EE EE E EEVE DD ++ DDE
Sbjct: 373 EIEEEVEENEEDAE-----EEEEE*EKEEEEVEEDDTMQNLDDDDE 495
Score = 41.6 bits (96), Expect = 0.001
Identities = 24/69 (34%), Positives = 37/69 (52%)
Frame = +1
Query: 60 EAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLA 119
E E E +E +D EE + +EE+EE+EY E E + +++ + E E+ E
Sbjct: 250 EPEPEKTVESDEKIDLEEENDPEEEMEEIEYEEVEEEEEVEEIEEEVEENEEDAEEEEEE 429
Query: 120 DVKEEEQVE 128
KEEE+VE
Sbjct: 430 *EKEEEEVE 456
Score = 38.9 bits (89), Expect = 0.007
Identities = 30/83 (36%), Positives = 45/83 (54%), Gaps = 2/83 (2%)
Frame = +1
Query: 36 EYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEV 95
E +PE+ V D+ I+ +E + +E+EE E V+EEE EVEE+E EEV
Sbjct: 250 EPEPEKTVESDEK----IDLEEENDPEEEMEEIEYEEVEEEE------EVEEIE---EEV 390
Query: 96 EGDDDDVAGDDE--HASEENVHE 116
E +++D ++E EE V E
Sbjct: 391 EENEEDAEEEEEE*EKEEEEVEE 459
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.311 0.133 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,507,214
Number of Sequences: 36976
Number of extensions: 304303
Number of successful extensions: 3305
Number of sequences better than 10.0: 379
Number of HSP's better than 10.0 without gapping: 2307
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2837
length of query: 726
length of database: 9,014,727
effective HSP length: 103
effective length of query: 623
effective length of database: 5,206,199
effective search space: 3243461977
effective search space used: 3243461977
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC149129.7


