

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
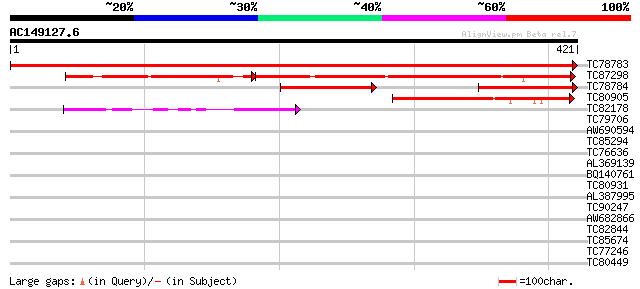
Query= AC149127.6 - phase: 0
(421 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC78783 similar to GP|21537171|gb|AAM61512.1 unknown {Arabidopsi... 862 0.0
TC87298 similar to GP|21537171|gb|AAM61512.1 unknown {Arabidopsi... 285 e-100
TC78784 similar to GP|21537171|gb|AAM61512.1 unknown {Arabidopsi... 155 4e-38
TC80905 similar to GP|16226611|gb|AAL16213.1 At1g01240/F6F3_11 {... 131 4e-31
TC82178 similar to GP|22655072|gb|AAM98127.1 expressed protein {... 70 1e-12
TC79706 similar to GP|18389230|gb|AAL67058.1 unknown protein {Ar... 36 0.024
AW690594 similar to GP|23498163|emb hypothetical protein {Plasmo... 34 0.093
TC85294 similar to GP|2961298|emb|CAA12357.1 hypothetical protei... 32 0.60
TC76636 homologue to GP|6625788|gb|AAF19398.1| dynamin homolog {... 30 1.3
AL369139 weakly similar to GP|21554368|gb| unknown {Arabidopsis ... 30 1.3
BQ140761 weakly similar to GP|18076245|emb phosphophoryn {Rattus... 29 3.0
TC80931 similar to GP|14190783|gb|AAF65166.2 putative phloem tra... 29 3.0
AL387995 29 3.9
TC90247 weakly similar to GP|21592898|gb|AAM64848.1 unknown {Ara... 28 5.1
AW682866 similar to GP|4314363|gb| unknown protein {Arabidopsis ... 28 5.1
TC82844 similar to PIR|A86198|A86198 hypothetical protein [impor... 28 6.6
TC85674 similar to PIR|E96500|E96500 probable histidine decarbox... 28 6.6
TC77246 similar to PIR|C71408|C71408 probable protein kinase - A... 28 6.6
TC80449 weakly similar to GP|15293267|gb|AAK93744.1 unknown prot... 28 6.6
>TC78783 similar to GP|21537171|gb|AAM61512.1 unknown {Arabidopsis
thaliana}, partial (21%)
Length = 1431
Score = 862 bits (2227), Expect = 0.0
Identities = 420/421 (99%), Positives = 420/421 (99%)
Frame = +2
Query: 1 MASAARARCAVNHCFTQDFMMPPTSSKQLEPDFTTSDSTDSDMKWWLHVKTNLAGDTDYT 60
MASAARARCAVNHCFTQDFMMPPTSSKQLEPDFTTSDSTDSDMKWWLHVKTNLAGDTDYT
Sbjct: 110 MASAARARCAVNHCFTQDFMMPPTSSKQLEPDFTTSDSTDSDMKWWLHVKTNLAGDTDYT 289
Query: 61 CQHLNTLESKLDAFSTRLHGDNVSIGSDQTVKNFDAVSFVGNAATAAIEQQWNVYPKYMK 120
CQHLNTLESKLDAFSTRLHGDNVSIGSDQTVKNFDAVSFVGNAATAAIEQQWNVYPKYMK
Sbjct: 290 CQHLNTLESKLDAFSTRLHGDNVSIGSDQTVKNFDAVSFVGNAATAAIEQQWNVYPKYMK 469
Query: 121 NSDTRTSKIEASLNSDLYLTPKKKNEGEFWFSDDATSFLISEHCKSTSSDFEPHWLGAEK 180
NSDTRTSKIEASLNSDLYLTPKKKNEGEFWFSDDATSFLISEHCKSTSSDFEPHWLGAEK
Sbjct: 470 NSDTRTSKIEASLNSDLYLTPKKKNEGEFWFSDDATSFLISEHCKSTSSDFEPHWLGAEK 649
Query: 181 SQPWWRTTGKDDLASLVAKKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLN 240
SQPWWRTTGKDDLASLVAKKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLN
Sbjct: 650 SQPWWRTTGKDDLASLVAKKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLN 829
Query: 241 QKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAELL 300
QKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAELL
Sbjct: 830 QKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAELL 1009
Query: 301 KALCRSQTRAREAEKAAQEACDEKEHILSLFFKQASQLFAYKQWLHVLQLENLCLQYKSK 360
KALCRSQTRAREAEKAAQEACDEKEHILSLFFKQASQLFAYKQWLHVLQLENLCLQYKSK
Sbjct: 1010KALCRSQTRAREAEKAAQEACDEKEHILSLFFKQASQLFAYKQWLHVLQLENLCLQYKSK 1189
Query: 361 NQPLLNNLFPYEGKKHRKNRRKVKNSRRGIGKCIFAFAVGLALAGAGLLLGWTIGCMFPS 420
NQPLLNNLFPYEGKKHRKNRRKVKNSRRGIGKCIFAFA GLALAGAGLLLGWTIGCMFPS
Sbjct: 1190NQPLLNNLFPYEGKKHRKNRRKVKNSRRGIGKCIFAFAGGLALAGAGLLLGWTIGCMFPS 1369
Query: 421 F 421
F
Sbjct: 1370F 1372
>TC87298 similar to GP|21537171|gb|AAM61512.1 unknown {Arabidopsis
thaliana}, partial (20%)
Length = 1773
Score = 285 bits (728), Expect(2) = e-100
Identities = 153/244 (62%), Positives = 187/244 (75%), Gaps = 6/244 (2%)
Frame = +2
Query: 183 PWWRTTGKDDLASLVAKKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLNQK 242
PWWRT GKD+LAS VA++S E+VENCDLP P K F + + + D ++ SSLNQK
Sbjct: 761 PWWRTAGKDELASFVAQRSVEHVENCDLPHPQPKSFAQRFS---KGVDHDKILPSSLNQK 931
Query: 243 LEMCSSDSNGCTS-TTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAELLK 301
E S +++G S TT TS CSFQDS+ FSS +SKDS S NK+ ++N E+++ ELL+
Sbjct: 932 AETGSLNADGYASGTTPTSSCSFQDSNSRFSSGQSKDS-GSINKDCQINPENSSITELLE 1108
Query: 302 ALCRSQTRAREAEKAAQEACDEKEHILSLFFKQASQLFAYKQWLHVLQLENLCLQYKSKN 361
ALC SQTRAREAEKAAQEA +EKEHILSLFFKQASQLFAYKQW ++LQLENLCLQ ++KN
Sbjct: 1109ALCHSQTRAREAEKAAQEAYNEKEHILSLFFKQASQLFAYKQWYYMLQLENLCLQLRNKN 1288
Query: 362 QPLLNNLFPYEGKKHRKNR-----RKVKNSRRGIGKCIFAFAVGLALAGAGLLLGWTIGC 416
QPLL NLFP+ + +KNR RK+ N + GIGKC+ AFAVGL+LAGAGLLLGWT+G
Sbjct: 1289QPLL-NLFPHRRMQLKKNRNKAEKRKINNRKCGIGKCVVAFAVGLSLAGAGLLLGWTMGW 1465
Query: 417 MFPS 420
MFPS
Sbjct: 1466MFPS 1477
Score = 97.8 bits (242), Expect(2) = e-100
Identities = 60/150 (40%), Positives = 90/150 (60%), Gaps = 8/150 (5%)
Frame = +1
Query: 42 DMKWWLHVKTNLAGDTDYTCQHLNTLESKLDAFSTRLHGDNVSIGSDQTVKNFDAVSFVG 101
DMKWWLHVK+NL + +Y+C+ ++L +F NV G DQ++ +FD +S++G
Sbjct: 355 DMKWWLHVKSNLDCEPNYSCK------TELGSFYAEFLDGNVKNGEDQSIIDFDDLSYIG 516
Query: 102 NAATAAIEQQWNVYP-KYMK-NSDTRTSKIEASLNSDLYLTPKKKNEGEFWFSD------ 153
+ +++Q +V P MK N++ R KIEASLN+DL+ TPKKK++ +F FSD
Sbjct: 517 SD-NLSVDQPHHVSPITCMKDNNNARMRKIEASLNNDLHFTPKKKDQKDFGFSDVGFLDC 693
Query: 154 DATSFLISEHCKSTSSDFEPHWLGAEKSQP 183
D ++FL+SE S H +G EK P
Sbjct: 694 DVSNFLVSEKTSS-------HLMGNEKPGP 762
>TC78784 similar to GP|21537171|gb|AAM61512.1 unknown {Arabidopsis
thaliana}, partial (7%)
Length = 695
Score = 155 bits (391), Expect = 4e-38
Identities = 73/73 (100%), Positives = 73/73 (100%)
Frame = +3
Query: 349 QLENLCLQYKSKNQPLLNNLFPYEGKKHRKNRRKVKNSRRGIGKCIFAFAVGLALAGAGL 408
QLENLCLQYKSKNQPLLNNLFPYEGKKHRKNRRKVKNSRRGIGKCIFAFAVGLALAGAGL
Sbjct: 234 QLENLCLQYKSKNQPLLNNLFPYEGKKHRKNRRKVKNSRRGIGKCIFAFAVGLALAGAGL 413
Query: 409 LLGWTIGCMFPSF 421
LLGWTIGCMFPSF
Sbjct: 414 LLGWTIGCMFPSF 452
Score = 151 bits (381), Expect = 5e-37
Identities = 71/71 (100%), Positives = 71/71 (100%)
Frame = +1
Query: 202 FEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSG 261
FEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSG
Sbjct: 25 FEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSG 204
Query: 262 CSFQDSDRTFS 272
CSFQDSDRTFS
Sbjct: 205 CSFQDSDRTFS 237
>TC80905 similar to GP|16226611|gb|AAL16213.1 At1g01240/F6F3_11 {Arabidopsis
thaliana}, partial (23%)
Length = 783
Score = 131 bits (330), Expect = 4e-31
Identities = 76/150 (50%), Positives = 99/150 (65%), Gaps = 15/150 (10%)
Frame = +2
Query: 285 KNSKVNSESAAKAELLKALCRSQTRAREAEKAAQEACDEKEHILSLFFKQASQLFAYKQW 344
++ +++ +KA+LL+ALC SQTRAREAE+AA++A E+EHIL+L FKQASQLFAYKQ
Sbjct: 158 QSQQISEGDTSKAQLLEALCHSQTRAREAEEAAKQAYAEREHILTLIFKQASQLFAYKQL 337
Query: 345 LHVLQLENLCLQYKSKNQPLLNNLFP-------YEGKKHRKNRRKVKNSRR---GIGKC- 393
+ +LQLE LC Q K+ N + LFP YE +K RK ++ N RR G C
Sbjct: 338 IKLLQLETLCNQIKN-NDLSTSTLFPVDLPWMSYERRKSRKRKQTYSNGRREREGKSNCD 514
Query: 394 ----IFAFAVGLALAGAGLLLGWTIGCMFP 419
A A+GL+L GAGLLLGWT+G M P
Sbjct: 515 ITTYAVAIALGLSLVGAGLLLGWTVGWMSP 604
>TC82178 similar to GP|22655072|gb|AAM98127.1 expressed protein {Arabidopsis
thaliana}, partial (20%)
Length = 897
Score = 70.5 bits (171), Expect = 1e-12
Identities = 49/176 (27%), Positives = 80/176 (44%)
Frame = +2
Query: 41 SDMKWWLHVKTNLAGDTDYTCQHLNTLESKLDAFSTRLHGDNVSIGSDQTVKNFDAVSFV 100
SD +WWLH + N T + L+ LE +++ T + GD + S ++
Sbjct: 248 SDPRWWLHSQPNYGYQKGVTNEKLSALEEEVE---TLIAGDEIKASSKNSL--------- 391
Query: 101 GNAATAAIEQQWNVYPKYMKNSDTRTSKIEASLNSDLYLTPKKKNEGEFWFSDDATSFLI 160
A + +V K+ T +I++ S +T K+ N+
Sbjct: 392 ------AFPELMDVMAKH------ETMEIDSIACS---VTSKQTND-------------- 484
Query: 161 SEHCKSTSSDFEPHWLGAEKSQPWWRTTGKDDLASLVAKKSFEYVENCDLPEPNIK 216
SS+ + W+ ++K++PWWR T KD+LAS V+ KS ++ENCDLP P K
Sbjct: 485 ------FSSEPDYSWIESDKAEPWWRMTDKDELASFVSHKSLNHIENCDLPPPKKK 634
>TC79706 similar to GP|18389230|gb|AAL67058.1 unknown protein {Arabidopsis
thaliana}, partial (40%)
Length = 676
Score = 36.2 bits (82), Expect = 0.024
Identities = 27/96 (28%), Positives = 47/96 (48%), Gaps = 10/96 (10%)
Frame = +3
Query: 240 NQKLEMCSSDSNGCT----STTLTSGCSFQDSDRTFS------SSESKDSDSSCNKNSKV 289
N L +CSS + GC +T+ F+ + ++ SS+ + S+ N NS
Sbjct: 300 NAILLLCSSYNKGCRPYMCATSRRYSNCFEQYKKAYTKATSVQSSQQETDYSNFNSNSGD 479
Query: 290 NSESAAKAELLKALCRSQTRAREAEKAAQEACDEKE 325
S++A ELL LCR Q + +AA+++ + K+
Sbjct: 480 RSDNAKVPELLCPLCRRQVKGWTVVEAARKSLNGKK 587
>AW690594 similar to GP|23498163|emb hypothetical protein {Plasmodium
falciparum 3D7}, partial (10%)
Length = 633
Score = 34.3 bits (77), Expect = 0.093
Identities = 22/70 (31%), Positives = 35/70 (49%)
Frame = -3
Query: 247 SSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAELLKALCRS 306
SS S+ +S++ +S S S + SSS S S SS + +S +S S+ +LC S
Sbjct: 223 SSSSSSSSSSSSSSSSSCSSSSSSSSSSSSSSSCSSSSSSSCSSSSSSTSPSSSSSLCSS 44
Query: 307 QTRAREAEKA 316
R + + A
Sbjct: 43 TFRRFQLDSA 14
Score = 31.2 bits (69), Expect = 0.78
Identities = 22/99 (22%), Positives = 46/99 (46%), Gaps = 3/99 (3%)
Frame = -3
Query: 239 LNQKLEMCSSDSNG---CTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAA 295
+ +K +C++ +G S + +S S S + SSS S S SSC+ +S +S S++
Sbjct: 313 VGRKESICNNSGSGRVLILSISSSSPSSSSSSSSSSSSSSSSSSSSSCSSSSSSSSSSSS 134
Query: 296 KAELLKALCRSQTRAREAEKAAQEACDEKEHILSLFFKQ 334
+ C S + + + ++ + + S F++
Sbjct: 133 SSS-----CSSSSSSSCSSSSSSTSPSSSSSLCSSTFRR 32
Score = 30.8 bits (68), Expect = 1.0
Identities = 17/46 (36%), Positives = 23/46 (49%)
Frame = -3
Query: 237 SSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSS 282
SS + CSS S+ +S++ +S CS S SSS S SS
Sbjct: 199 SSSSSSSSSCSSSSSSSSSSSSSSSCSSSSSSSCSSSSSSTSPSSS 62
>TC85294 similar to GP|2961298|emb|CAA12357.1 hypothetical protein {Cicer
arietinum}, partial (61%)
Length = 1353
Score = 31.6 bits (70), Expect = 0.60
Identities = 33/125 (26%), Positives = 54/125 (42%), Gaps = 30/125 (24%)
Frame = +1
Query: 216 KPFRKIHTL----QPRETDKEEN------------QVSSLNQKLEMCSSDSNGCTSTTLT 259
+P R H + + R + K+EN S+L Q L + GCTSTT
Sbjct: 124 RPIRSRHVMMIQTRGRRSTKQENIYMQPRLHCSFTHPSTLFQSLSFPNGQC*GCTSTTTN 303
Query: 260 SGC-------------SFQDSDRTFSSSESKDSDSSCNKN-SKVNSESAAKAELLKALCR 305
+ S +SDR+ S S + S SSC+++ ++ N+ +A + + CR
Sbjct: 304 NSARNNTGGDQDRGNNSRTNSDRS-SCSRTTGS*SSCHRSTNRRNNRTANRNNRRNSSCR 480
Query: 306 SQTRA 310
TR+
Sbjct: 481 RNTRS 495
>TC76636 homologue to GP|6625788|gb|AAF19398.1| dynamin homolog {Astragalus
sinicus}, partial (69%)
Length = 2420
Score = 30.4 bits (67), Expect = 1.3
Identities = 28/114 (24%), Positives = 49/114 (42%), Gaps = 5/114 (4%)
Frame = +3
Query: 191 DDLASLVAKKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDS 250
+ L S V+ +S +E L + N+K R+ + Q K Q+S + + S+ S
Sbjct: 1590 NQLYSSVSGQSTAKIEELLLEDQNVKRRRERYQKQSSLLSKLTRQLSIHDNRAAAASNWS 1769
Query: 251 NGCTSTTLTSGCSFQDSDRTFSSSES-----KDSDSSCNKNSKVNSESAAKAEL 299
NG ++ S D F ++ + S S N +S+ NS+ A +L
Sbjct: 1770 NGSAESSPRSSGPGDDWRTAFDAASNGSVSRSGSRSGSNGHSRHNSDPAQNGDL 1931
>AL369139 weakly similar to GP|21554368|gb| unknown {Arabidopsis thaliana},
partial (11%)
Length = 388
Score = 30.4 bits (67), Expect = 1.3
Identities = 32/116 (27%), Positives = 50/116 (42%), Gaps = 6/116 (5%)
Frame = +3
Query: 206 ENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCT-----STTLTS 260
E D P+ + RK +T+ + +SS + E SDS+ S S
Sbjct: 36 EKDDGEPPSKRDSRKTRENSDIDTETDRKHISSKKKSRESSESDSDKRRRRKRKSRRKYS 215
Query: 261 GCSFQDSDR-TFSSSESKDSDSSCNKNSKVNSESAAKAELLKALCRSQTRAREAEK 315
S +SD + S SES S+ S +++ SES + + K + + R RE EK
Sbjct: 216 SESDSESDSGSDSESESDKSERSGSESEYSGSESEGERKERKRKEKRRRREREEEK 383
>BQ140761 weakly similar to GP|18076245|emb phosphophoryn {Rattus
norvegicus}, partial (9%)
Length = 620
Score = 29.3 bits (64), Expect = 3.0
Identities = 24/79 (30%), Positives = 39/79 (48%), Gaps = 5/79 (6%)
Frame = +1
Query: 225 QPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTS-----GCSFQDSDRTFSSSESKDS 279
+P K+E + K SSDS+ +S+ S +SD SSS S DS
Sbjct: 367 EPVPAPKKEKAKKAKKAKSVSSSSDSSDDSSSESESEPEPXNVPLPESDD--SSSASSDS 540
Query: 280 DSSCNKNSKVNSESAAKAE 298
DSS + +S +S+S++ ++
Sbjct: 541 DSSSDSSSDSSSDSSSXSD 597
>TC80931 similar to GP|14190783|gb|AAF65166.2 putative phloem transcription
factor M1 {Apium graveolens}, partial (67%)
Length = 1193
Score = 29.3 bits (64), Expect = 3.0
Identities = 26/98 (26%), Positives = 45/98 (45%), Gaps = 1/98 (1%)
Frame = +2
Query: 272 SSSESKDSDSSCNKNSKVNSESAAKAELLKALCRSQTRAREAEKAAQEACDEKEHILSLF 331
SSS ++ + + CN S+ N S+ +L Q + ++ ++ Q ++ LS
Sbjct: 485 SSSSNRCNSNKCNNTSRCNRNSSIFPQL-------QQQMQQQQQQQQLPQLQQNQQLSQI 643
Query: 332 FKQASQLFAYKQWLHVLQLENLC-LQYKSKNQPLLNNL 368
+Q QL +Q L LQ + L LQ + + P L L
Sbjct: 644 QQQIPQLQQQQQQLPQLQQQQLSQLQQQQQQLPQLQQL 757
>AL387995
Length = 486
Score = 28.9 bits (63), Expect = 3.9
Identities = 26/111 (23%), Positives = 45/111 (40%), Gaps = 5/111 (4%)
Frame = +2
Query: 214 NIKPFRKIHTLQPRETDKEE---NQVSSLNQKLEMCSS--DSNGCTSTTLTSGCSFQDSD 268
NIK RK+ +E + E+ +S ++K++ S+ D N TS + + +
Sbjct: 119 NIKYLRKLLEQIEKEVNSEQIDKTAISEYSRKIQQLSNIVDENKLTSPVTRTFSQARFVN 298
Query: 269 RTFSSSESKDSDSSCNKNSKVNSESAAKAELLKALCRSQTRAREAEKAAQE 319
+ K+ ++ +E K ELL+ RAR E QE
Sbjct: 299 HANQTQAEKNKENQLELKMVRQAEKEQKEELLQPPTTFTXRARRYENLFQE 451
>TC90247 weakly similar to GP|21592898|gb|AAM64848.1 unknown {Arabidopsis
thaliana}, partial (60%)
Length = 1041
Score = 28.5 bits (62), Expect = 5.1
Identities = 29/95 (30%), Positives = 43/95 (44%), Gaps = 7/95 (7%)
Frame = -2
Query: 217 PFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSES 276
P K+HTLQ R +D E + +L+ L + +S + S L S F S R SS
Sbjct: 704 PLDKLHTLQVRSSDAE--TIIALHPPLPITTSVTRMVCSEKLYS*RFF--SSRLLKSSNK 537
Query: 277 KDSD-------SSCNKNSKVNSESAAKAELLKALC 304
SD S + +S+ +SE+ + LK C
Sbjct: 536 LKSDQTLTYPSSEPDISSESSSENTRQVTALK*AC 432
>AW682866 similar to GP|4314363|gb| unknown protein {Arabidopsis thaliana},
partial (16%)
Length = 557
Score = 28.5 bits (62), Expect = 5.1
Identities = 22/64 (34%), Positives = 27/64 (41%), Gaps = 1/64 (1%)
Frame = +3
Query: 61 CQHLNTLESKLDAFSTRLHGDNVSIGSDQTVKNFDAVSFVGNAATAAIEQQWN-VYPKYM 119
C LN + KLD L+G + NFD VS V E+ WN +YP
Sbjct: 324 CHRLNPVRYKLDC--EELYG--------LVLDNFDVVSTVEGICGRQTEEIWNKLYPDEP 473
Query: 120 KNSD 123
NSD
Sbjct: 474 YNSD 485
>TC82844 similar to PIR|A86198|A86198 hypothetical protein [imported] -
Arabidopsis thaliana, partial (9%)
Length = 834
Score = 28.1 bits (61), Expect = 6.6
Identities = 21/61 (34%), Positives = 26/61 (42%), Gaps = 5/61 (8%)
Frame = +3
Query: 226 PRETDKEENQVSSLNQKLEMCSSDSNG-----CTSTTLTSGCSFQDSDRTFSSSESKDSD 280
PRE +E S D+NG +S++ SG S DSD SSS DS
Sbjct: 219 PRERQADERNASPSLAVQGEIRVDNNGNKTSSSSSSSSDSGSSTSDSDSDSSSSSGSDSG 398
Query: 281 S 281
S
Sbjct: 399 S 401
>TC85674 similar to PIR|E96500|E96500 probable histidine decarboxylase
[imported] - Arabidopsis thaliana, partial (85%)
Length = 1839
Score = 28.1 bits (61), Expect = 6.6
Identities = 13/26 (50%), Positives = 16/26 (61%)
Frame = -2
Query: 166 STSSDFEPHWLGAEKSQPWWRTTGKD 191
STS+ F H G EK++ W RT KD
Sbjct: 122 STSNHFVIHNRGTEKNEIWKRTVAKD 45
>TC77246 similar to PIR|C71408|C71408 probable protein kinase - Arabidopsis
thaliana, partial (65%)
Length = 1715
Score = 28.1 bits (61), Expect = 6.6
Identities = 10/24 (41%), Positives = 19/24 (78%)
Frame = -1
Query: 345 LHVLQLENLCLQYKSKNQPLLNNL 368
+++L+L++ LQ+KS Q ++NNL
Sbjct: 1238 MNILKLQSTILQFKSNQQQIINNL 1167
>TC80449 weakly similar to GP|15293267|gb|AAK93744.1 unknown protein
{Arabidopsis thaliana}, partial (56%)
Length = 1239
Score = 28.1 bits (61), Expect = 6.6
Identities = 11/44 (25%), Positives = 20/44 (45%)
Frame = -3
Query: 144 KNEGEFWFSDDATSFLISEHCKSTSSDFEPHWLGAEKSQPWWRT 187
+ ++W S + S + SD E W+G E+ + W+T
Sbjct: 352 ETRAKYWLSGSSKSAEARRRLAAEMSDGEARWVGLEEMKRLWKT 221
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.127 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,690,784
Number of Sequences: 36976
Number of extensions: 211642
Number of successful extensions: 1360
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 1317
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1343
length of query: 421
length of database: 9,014,727
effective HSP length: 99
effective length of query: 322
effective length of database: 5,354,103
effective search space: 1724021166
effective search space used: 1724021166
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC149127.6


