

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149080.7 - phase: 0
(165 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
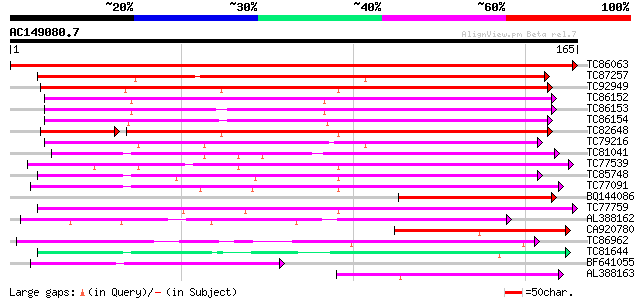
Sequences producing significant alignments: (bits) Value
TC86063 weakly similar to GP|16323382|gb|AAL15185.1 unknown prot... 335 4e-93
TC87257 similar to GP|16323382|gb|AAL15185.1 unknown protein {Ar... 166 5e-42
TC92949 similar to GP|10998131|dbj|BAB03102. gene_id:MEC18.3~unk... 114 2e-26
TC86152 similar to GP|15451080|gb|AAK96811.1 Unknown protein {Ar... 112 7e-26
TC86153 similar to GP|15451080|gb|AAK96811.1 Unknown protein {Ar... 109 6e-25
TC86154 similar to GP|15451080|gb|AAK96811.1 Unknown protein {Ar... 107 2e-24
TC82648 similar to GP|10998131|dbj|BAB03102. gene_id:MEC18.3~unk... 91 2e-23
TC79216 similar to GP|5669654|gb|AAD46412.1| ER6 protein {Lycope... 102 5e-23
TC81041 similar to GP|6714413|gb|AAF26101.1| unknown protein {Ar... 77 2e-15
TC77539 similar to GP|16118856|gb|AAL14629.1 growth-on protein G... 72 8e-14
TC85748 similar to GP|11692860|gb|AAG40033.1 AT3g53990 {Arabidop... 71 2e-13
TC77091 weakly similar to GP|11602751|emb|CAC18558. ENOD18 prote... 65 2e-11
BQ144086 similar to GP|20067187|gb| putative universal stress pr... 64 3e-11
TC77759 homologue to GP|11602747|emb|CAC18556. early nodulin ENO... 61 2e-10
AL388162 56 6e-09
CA920780 similar to GP|21689829|gb| unknown protein {Arabidopsis... 53 5e-08
TC86962 similar to GP|15320408|dbj|BAB63912. RD2 protein {Arabid... 49 7e-07
TC81644 similar to GP|19386852|dbj|BAB86230. hypothetical protei... 44 4e-05
BF641055 similar to GP|20804872|dbj contains EST C19329(E10264)~... 42 1e-04
AL388163 weakly similar to GP|2160182|gb|A ESTs gb|ATTS1236 gb|T... 40 4e-04
>TC86063 weakly similar to GP|16323382|gb|AAL15185.1 unknown protein
{Arabidopsis thaliana}, partial (46%)
Length = 984
Score = 335 bits (860), Expect = 4e-93
Identities = 165/165 (100%), Positives = 165/165 (100%)
Frame = +2
Query: 1 MAGITENGRRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTG 60
MAGITENGRRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTG
Sbjct: 254 MAGITENGRRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTG 433
Query: 61 YLFSSDITATMEKYSQQVADCVLEKAKIVCNDVQNVETRIENGDPRDVICQAVQKMGVDI 120
YLFSSDITATMEKYSQQVADCVLEKAKIVCNDVQNVETRIENGDPRDVICQAVQKMGVDI
Sbjct: 434 YLFSSDITATMEKYSQQVADCVLEKAKIVCNDVQNVETRIENGDPRDVICQAVQKMGVDI 613
Query: 121 LVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKSTTGGDN 165
LVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKSTTGGDN
Sbjct: 614 LVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKSTTGGDN 748
>TC87257 similar to GP|16323382|gb|AAL15185.1 unknown protein {Arabidopsis
thaliana}, partial (64%)
Length = 747
Score = 166 bits (419), Expect = 5e-42
Identities = 82/154 (53%), Positives = 111/154 (71%), Gaps = 5/154 (3%)
Frame = +3
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQ-NSKDHLILLYVKPPRVVYSAFDGTGYLFSSDI 67
R+IMVAIDE EES+YAL+W + NL+ N+ + L+LLYVKPP VYS D GY+FS+D
Sbjct: 168 RKIMVAIDESEESMYALSWSISNLIADTNNNNKLVLLYVKPPSAVYS-LDSAGYIFSNDT 344
Query: 68 TATMEKYSQQVADCVLEKAKIVCNDVQNVETRIEN----GDPRDVICQAVQKMGVDILVM 123
T+E YS Q+A V+++A+ + + + + IE GD ++VIC A +K+G D LVM
Sbjct: 345 IDTLENYSSQLAKSVMKRAEAIYRNFDDTDINIEKVVGTGDAKNVICNAAKKLGADTLVM 524
Query: 124 GSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKP 157
GSHGYG IKRA LGSVS++C +N KCPV+IVK+P
Sbjct: 525 GSHGYGFIKRALLGSVSDYCVKNAKCPVVIVKQP 626
>TC92949 similar to GP|10998131|dbj|BAB03102. gene_id:MEC18.3~unknown
protein {Arabidopsis thaliana}, partial (45%)
Length = 727
Score = 114 bits (284), Expect = 2e-26
Identities = 66/162 (40%), Positives = 99/162 (60%), Gaps = 13/162 (8%)
Frame = +3
Query: 10 RIMVAIDEGEESIYALTWCLKNL---------VFQNSKDHLILLYVKPPRVVYSAFDGTG 60
++MVAIDE + S YAL W L NL ++ + L++V+P Y G G
Sbjct: 93 KVMVAIDESDGSFYALKWALDNLFNVMTTMEGTSSENEGMVFLVHVEPIFHNYVHPIGPG 272
Query: 61 ---YLFSSDITATMEKYSQQVADCVLEKAKIVCNDVQ-NVETRIENGDPRDVICQAVQKM 116
+ +S + +++K Q+ + +L +A +C D Q E+ I NGDPR++ICQA ++M
Sbjct: 273 GAAFYPASVVVDSVKKGQQERSAVILSRALQMCKDKQVKAESVILNGDPREMICQASEQM 452
Query: 117 GVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPK 158
VD+L+MGS G G +KRAFLGSVS++CA + K P+LIVK P+
Sbjct: 453 QVDLLIMGSRGLGTLKRAFLGSVSDYCAHHAKAPILIVKPPE 578
>TC86152 similar to GP|15451080|gb|AAK96811.1 Unknown protein {Arabidopsis
thaliana}, partial (51%)
Length = 834
Score = 112 bits (280), Expect = 7e-26
Identities = 61/152 (40%), Positives = 90/152 (59%), Gaps = 3/152 (1%)
Frame = +3
Query: 11 IMVAIDEGEESIYALTWCLKNLVF-QNSKDHLILLYVKPPRVVYSAFDGTGYLFSSDITA 69
++VAIDE + S YAL W L + NS L+L++ +P G Y ++++
Sbjct: 93 MLVAIDESDHSAYALKWTLDHFFSTNNSVFKLVLVHARPAATSSVGLAGPVYAGAAEVLP 272
Query: 70 TMEKYSQQVADCVLEKAKIVC--NDVQNVETRIENGDPRDVICQAVQKMGVDILVMGSHG 127
++ +++A V E AK +C V +V + GD R+V+C V+K ILV+GSHG
Sbjct: 273 IVDSDLRKIAARVAENAKQLCIKKSVNDVIVEVVEGDARNVLCDTVEKYRASILVVGSHG 452
Query: 128 YGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKS 159
YG IKRA LGSVS++CA + C V+IVKKPK+
Sbjct: 453 YGAIKRAVLGSVSDYCAHHAHCTVMIVKKPKT 548
>TC86153 similar to GP|15451080|gb|AAK96811.1 Unknown protein {Arabidopsis
thaliana}, partial (74%)
Length = 784
Score = 109 bits (272), Expect = 6e-25
Identities = 61/152 (40%), Positives = 90/152 (59%), Gaps = 3/152 (1%)
Frame = +1
Query: 11 IMVAIDEGEESIYALTWCLKNLVF-QNSKDHLILLYVKPPRVVYSAFDGTGYLFSSDITA 69
++VAIDE + S YAL W L + NS L+L++ +P G G ++++
Sbjct: 109 MLVAIDESDHSAYALKWTLDHFFSTNNSVFKLVLVHARPAATSSVGLAGPG---AAEVLP 279
Query: 70 TMEKYSQQVADCVLEKAKIVC--NDVQNVETRIENGDPRDVICQAVQKMGVDILVMGSHG 127
++ +++A V E AK +C V +V + GD R+V+C V+K ILV+GSHG
Sbjct: 280 IVDSDLRKIAARVAENAKQLCIKKSVNDVIVEVVEGDARNVLCDTVEKYRASILVVGSHG 459
Query: 128 YGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKS 159
YG IKRA LGSVS++CA + C V+IVKKPK+
Sbjct: 460 YGAIKRAVLGSVSDYCAHHAHCTVMIVKKPKT 555
>TC86154 similar to GP|15451080|gb|AAK96811.1 Unknown protein {Arabidopsis
thaliana}, partial (75%)
Length = 1086
Score = 107 bits (267), Expect = 2e-24
Identities = 62/152 (40%), Positives = 91/152 (59%), Gaps = 4/152 (2%)
Frame = +1
Query: 11 IMVAIDEGEESIYALTWCLKNLV--FQNSKDHLILLYVKPPRVVYSAFDGTGYLFSSDIT 68
++V +D+ E S YAL W L +LV N L+L++ KP F G +++I
Sbjct: 325 MVVGVDDSEFSTYALEWTLDHLVTTLPNPIFKLVLVFAKPSPSTNVGFVGPAG--AAEIL 498
Query: 69 ATMEKYSQQVADCVLEKAKIVCN--DVQNVETRIENGDPRDVICQAVQKMGVDILVMGSH 126
+E ++ A V+E+A+ +C V++V + +GD +V+C AV K ILV+GSH
Sbjct: 499 PIVEADLKRTATIVIERAQEICTKRSVKDVIMEVVDGDATNVLCDAVDKHHASILVVGSH 678
Query: 127 GYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPK 158
GYG IKRA LGSVS++CA + C V+IVKKPK
Sbjct: 679 GYGAIKRAVLGSVSDYCAHHAHCTVMIVKKPK 774
>TC82648 similar to GP|10998131|dbj|BAB03102. gene_id:MEC18.3~unknown
protein {Arabidopsis thaliana}, partial (48%)
Length = 814
Score = 90.5 bits (223), Expect(2) = 2e-23
Identities = 50/131 (38%), Positives = 80/131 (60%), Gaps = 7/131 (5%)
Frame = +2
Query: 35 QNSKDHLILLYVKPPRVVYSAFDGTG------YLFSSDITATMEKYSQQVADCVLEKAKI 88
+N + + L++V+P Y+ G Y ++ +++K Q+ + + +A
Sbjct: 221 ENEEGIVFLVHVEPNFHTYAYPIPMGPAGVAFYPSIPEVGDSLKKAQQEKSAAIASQALQ 400
Query: 89 VCNDVQ-NVETRIENGDPRDVICQAVQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNV 147
+C D Q E+ I NGD R++IC+A +KM VD+L+MGS G G +KRAFLGSVS++CA +
Sbjct: 401 MCKDKQVKAESVILNGDAREMICEATEKMQVDLLIMGSRGLGKLKRAFLGSVSDYCAHHA 580
Query: 148 KCPVLIVKKPK 158
K P+LIVK P+
Sbjct: 581 KAPILIVKPPE 613
Score = 34.3 bits (77), Expect(2) = 2e-23
Identities = 14/23 (60%), Positives = 18/23 (77%)
Frame = +3
Query: 10 RIMVAIDEGEESIYALTWCLKNL 32
++MVAIDE + S+YAL W L NL
Sbjct: 111 KVMVAIDESDGSLYALKWALDNL 179
>TC79216 similar to GP|5669654|gb|AAD46412.1| ER6 protein {Lycopersicon
esculentum}, partial (86%)
Length = 1094
Score = 102 bits (255), Expect = 5e-23
Identities = 58/162 (35%), Positives = 92/162 (55%), Gaps = 17/162 (10%)
Frame = +3
Query: 11 IMVAIDEGEESIYALTWCLKNLVFQN------SKDHLILLYVKPPRVVYSA-------FD 57
++VA+D EES+ AL W L+NL ++ I+L+V+ P + + F
Sbjct: 102 VVVAVDGSEESMNALRWALENLKLRSPAPDSTDAGSFIILHVQSPPSIATGLNPGSIPFG 281
Query: 58 GTGYLFSSDITATMEKYSQQVADCVLEKAKIVCNDVQNVETRIEN----GDPRDVICQAV 113
G L A +E + +++ D + + A +C+ NV+T++ GDP++ IC+ V
Sbjct: 282 GPSDLEVPAFAAAIEAHQKRITDSIFDHALGICSTF-NVKTKVRTHVVVGDPKEKICETV 458
Query: 114 QKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
Q + D+LVMGS +G IKR FLGSVSN+CA + +CPV I+K
Sbjct: 459 QDLHADVLVMGSRAFGPIKRMFLGSVSNYCAHHSECPVTIIK 584
>TC81041 similar to GP|6714413|gb|AAF26101.1| unknown protein {Arabidopsis
thaliana}, partial (94%)
Length = 581
Score = 77.4 bits (189), Expect = 2e-15
Identities = 54/159 (33%), Positives = 80/159 (49%), Gaps = 11/159 (6%)
Frame = +2
Query: 13 VAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSA---FDGTGYLFSS---- 65
VA+D S AL W + NL+ N D +I++ V+PP ++ F+ TG
Sbjct: 56 VAMDFSPTSKLALRWAVDNLI--NKNDQIIMINVQPPSADHTRKELFEDTGSPLVPLEEL 229
Query: 66 -DITATME---KYSQQVADCVLEKAKIVCNDVQNVETRIENGDPRDVICQAVQKMGVDIL 121
+I T + +V D + +KI V ++ GDPR+ +C AV+ + +D L
Sbjct: 230 REINFTKQYGIAKDPEVIDILETASKI---KGAKVVAKVYWGDPREKLCNAVEDLHLDSL 400
Query: 122 VMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKST 160
V+GS G G IK LGSVS H N CPV +VK +S+
Sbjct: 401 VIGSRGLGTIKSVLLGSVSKHVVTNASCPVTVVKGMQSS 517
>TC77539 similar to GP|16118856|gb|AAL14629.1 growth-on protein GRO10
{Euphorbia esula}, partial (44%)
Length = 2020
Score = 72.4 bits (176), Expect = 8e-14
Identities = 47/171 (27%), Positives = 86/171 (49%), Gaps = 12/171 (7%)
Frame = +1
Query: 6 ENGRRIMVAIDEGEESIY---------ALTWCLKNLVFQN-SKDHLILLYVKPPRVVYSA 55
E R+M+ ++E Y A W + ++ N S HL+ L+V+ P
Sbjct: 1141 EEATRVMIGVNESSLKGYPHPSISSKGAFDWTVSKIIRNNVSAFHLLFLHVQVPDE--DG 1314
Query: 56 FDGTGYLFSS-DITATMEKYSQQVADCVLEKAKIVCNDVQ-NVETRIENGDPRDVICQAV 113
+D +++S + M++ + +LE CN++ E I+ GDP++VI V
Sbjct: 1315 YDDVDSIYASGEDFKNMKQQEKARGTHLLEYFVNRCNEIGVTCEAWIKQGDPKEVILNEV 1494
Query: 114 QKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKSTTGGD 164
+++ D+LV+GS G G ++ F+G+VS C ++ +CPV+ +K+ T D
Sbjct: 1495 KRVRPDLLVVGSRGLGPFQKVFVGTVSEFCWKHAECPVMTIKRNADETPRD 1647
>TC85748 similar to GP|11692860|gb|AAG40033.1 AT3g53990 {Arabidopsis
thaliana}, complete
Length = 823
Score = 70.9 bits (172), Expect = 2e-13
Identities = 53/155 (34%), Positives = 78/155 (50%), Gaps = 8/155 (5%)
Frame = +1
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVK-----PPRVVYSAFDGTGYLF 63
R I VA+D + S AL W L+NL + D++ ++++ R A DG+ +
Sbjct: 70 RTIGVALDFSKSSKNALKWALENLA--DKGDNIYIIHISHDSLDEARNQLWAKDGSPLIP 243
Query: 64 SSDITAT--MEKYSQQVADCVLEKAKIVCNDVQ-NVETRIENGDPRDVICQAVQKMGVDI 120
+ M+KY Q+ VL+ + NV T++ GD R+ + AV+ + +D
Sbjct: 244 LKEFREPEIMKKYGVQIDIEVLDLLDTFSRQKEVNVVTKVYWGDAREKLMDAVEDLKLDS 423
Query: 121 LVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
LVMGS G I+R LGSVSN N CPV IVK
Sbjct: 424 LVMGSRGLSTIQRILLGSVSNFVMTNAPCPVTIVK 528
>TC77091 weakly similar to GP|11602751|emb|CAC18558. ENOD18 protein {Vicia
faba}, partial (67%)
Length = 896
Score = 64.7 bits (156), Expect = 2e-11
Identities = 52/163 (31%), Positives = 78/163 (46%), Gaps = 8/163 (4%)
Frame = +3
Query: 7 NGRRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYS-----AFDGTGY 61
+GR+I VA+D + S AL W + NL+ + D L +++V S A G+
Sbjct: 111 SGRQIGVALDFSKGSKIALKWAIDNLL--RTGDTLYIVHVNHSHPTESRNLLWATTGSPL 284
Query: 62 LFSSDITA--TMEKYSQQVADCVLEKAKIVCNDVQ-NVETRIENGDPRDVICQAVQKMGV 118
+ S+ + +Y VL+ Q V ++ GD R+ I +V + +
Sbjct: 285 IPLSEFREKNVVHQYEVDPDAEVLDILDTASRQKQVTVVGKVYWGDAREKIVDSVGDLKL 464
Query: 119 DILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKSTT 161
D LVMGS G G I+R LGSVS + N CPV IVK+ + T
Sbjct: 465 DALVMGSRGLGAIQRVLLGSVSTYVTSNASCPVTIVKESVAPT 593
>BQ144086 similar to GP|20067187|gb| putative universal stress protein USP1
{Oryza sativa (indica cultivar-group)}, partial (27%)
Length = 842
Score = 63.9 bits (154), Expect = 3e-11
Identities = 29/46 (63%), Positives = 36/46 (78%)
Frame = +1
Query: 114 QKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKS 159
+K ILV+GSHGYG IKRA LGSVS++CA + C V+IVKKPK+
Sbjct: 1 EKYRASILVVGSHGYGAIKRAVLGSVSDYCAHHAHCTVMIVKKPKT 138
>TC77759 homologue to GP|11602747|emb|CAC18556. early nodulin ENOD18 {Vicia
faba}, complete
Length = 848
Score = 61.2 bits (147), Expect = 2e-10
Identities = 48/163 (29%), Positives = 76/163 (46%), Gaps = 6/163 (3%)
Frame = +1
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPP---RVVYSAFDGTGYLFSS 65
R+I VAID + S AL W + N+ + +LI + R A G+ +
Sbjct: 82 RKIGVAIDFSKNSKNALKWAIVNMADKGDTFYLIHINSNSSDESRNKQFAKTGSPLISLE 261
Query: 66 DI--TATMEKYSQQVADCVLEKAKIVCNDVQ-NVETRIENGDPRDVICQAVQKMGVDILV 122
++ M KY Q VL+ + + +V ++ GD R + +++ + +D LV
Sbjct: 262 ELKEVEVMSKYGVQTDVEVLDMLDTLATQKEVSVVAKLYWGDARQKLMDSIEDLKLDALV 441
Query: 123 MGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKSTTGGDN 165
+GS G IKR LGSVSN + CPV IVK S++ +
Sbjct: 442 LGSRGLSTIKRILLGSVSNFVMVHSPCPVTIVKYYSSSSSSSS 570
>AL388162
Length = 501
Score = 56.2 bits (134), Expect = 6e-09
Identities = 46/156 (29%), Positives = 71/156 (45%), Gaps = 13/156 (8%)
Frame = +2
Query: 4 ITENGRRIMVAID-EGEESIYALTWCLKN-LVFQNSKDHLILLYVKPPRVVYSAFD---- 57
I R+++VA+D EE+ Y + W + N L+ + + HLI + S FD
Sbjct: 59 IMSKQRKVVVALDPSSEEAPYTIDWIIDNFLIPERDEVHLISALC-----LNSDFDVTEL 223
Query: 58 GTGYLFSSDITATMEKYSQQVADCV-------LEKAKIVCNDVQNVETRIENGDPRDVIC 110
G ++++ A +EK +Q + LE A I C VE N D R+++
Sbjct: 224 GMTVNYAAEYAANLEKEVEQKTNDAMQPFAKKLETANIRCE----VELIKSNADSRNIVV 391
Query: 111 QAVQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQN 146
+ D+L+MGS KR FLGS S+ C N
Sbjct: 392 DYTENEKADVLIMGSRDLSTWKRLFLGSFSDFCQHN 499
>CA920780 similar to GP|21689829|gb| unknown protein {Arabidopsis thaliana},
partial (27%)
Length = 780
Score = 53.1 bits (126), Expect = 5e-08
Identities = 22/54 (40%), Positives = 39/54 (71%), Gaps = 3/54 (5%)
Frame = -1
Query: 113 VQKMGVDILVMGSHGYGVIKRAF---LGSVSNHCAQNVKCPVLIVKKPKSTTGG 163
V+++G+ ++MGS G+G KR+ LGSVS++C ++ CPV++V+ P+ + GG
Sbjct: 717 VERLGLSAVIMGSRGFGATKRSSNGKLGSVSDYCVRHCVCPVVVVRYPEESNGG 556
>TC86962 similar to GP|15320408|dbj|BAB63912. RD2 protein {Arabidopsis
thaliana}, partial (88%)
Length = 913
Score = 49.3 bits (116), Expect = 7e-07
Identities = 44/155 (28%), Positives = 67/155 (42%), Gaps = 3/155 (1%)
Frame = +2
Query: 3 GITENGRRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYL 62
G GR I++AID G S +A W L +L HL+ V + T Y
Sbjct: 233 GERRRGRDIVLAIDHGPNSKHAFDWALIHLCRLADTIHLV-------HAVSDVKNQTVY- 388
Query: 63 FSSDITATMEKYSQQVADCVLEKAKIVCNDVQNVET--RIENGDPRDVICQAVQKMGVDI 120
D+T + +EK + V V+T RI GD VIC+ +++
Sbjct: 389 ---DLTQGL-----------MEKLAVEAFQVSMVKTVARIVQGDAGKVICKEAERIKPAA 526
Query: 121 LVMGSHGYGVIKRAFLGSVSNHCAQNVK-CPVLIV 154
+V+G+ G + + GSV +C + K PV+IV
Sbjct: 527 VVLGTRGRSLFQSVIQGSVGEYCFHHCKAAPVVIV 631
>TC81644 similar to GP|19386852|dbj|BAB86230. hypothetical protein~similar
to Arabidopsis thaliana chromosome 1 At1g44760, partial
(51%)
Length = 797
Score = 43.5 bits (101), Expect = 4e-05
Identities = 41/164 (25%), Positives = 63/164 (38%), Gaps = 9/164 (5%)
Frame = +2
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYLFSSDIT 68
+R+MV +D S +A+ W L ++V N D L LLY+ P+ ++ T YL
Sbjct: 350 KRVMVVVDGTSHSKHAMIWALTHVV--NKGDLLTLLYIVSPQSASDSYSST-YL------ 502
Query: 69 ATMEKYSQQVADCVLEKAKIVCNDVQNVETRIENGDPRDVICQAVQKMGVDILVMGSHGY 128
+ DC E VE + G + V+K+ V ILV+G
Sbjct: 503 --VNHLGSLCKDCKPE---------VEVEALVIQGPKLATVMSQVKKLEVSILVLGQKKP 649
Query: 129 GVIKRAFLGSVSN---------HCAQNVKCPVLIVKKPKSTTGG 163
GS S+ +C N +C + V+K G
Sbjct: 650 SSFFSCLCGSSSSCSSKEEFVEYCINNAECLTIGVRKRSQGNNG 781
>BF641055 similar to GP|20804872|dbj contains EST C19329(E10264)~similar to
Arabidopsis thaliana chromosome 1 T23J18.3~unknown
protein, partial (31%)
Length = 492
Score = 41.6 bits (96), Expect = 1e-04
Identities = 23/74 (31%), Positives = 43/74 (58%)
Frame = +2
Query: 7 NGRRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYLFSSD 66
+ R++ +A+D +ES YA+ W ++N + D +ILL+V+P V+Y A G+ ++D
Sbjct: 179 SNRKVAIAVDLSDESAYAVRWAVQN--YLRPGDTVILLHVRPTYVLYGADWGSVTSPTAD 352
Query: 67 ITATMEKYSQQVAD 80
E+ Q++ D
Sbjct: 353 GGDASEESRQKMED 394
>AL388163 weakly similar to GP|2160182|gb|A ESTs gb|ATTS1236 gb|T43334
gb|N97019 gb|AA395203 come from this gene. {Arabidopsis
thaliana}, partial (13%)
Length = 569
Score = 40.0 bits (92), Expect = 4e-04
Identities = 23/69 (33%), Positives = 34/69 (48%), Gaps = 3/69 (4%)
Frame = +1
Query: 96 VETRIENGDPRDVICQA---VQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVL 152
VE N D R+++ ++++ V KR FLGS S+ C N CPVL
Sbjct: 199 VELIKSNADSRNIVVDV*LKMRRLTVSCNKWEVDDLSTWKRLFLGSFSDFCQHNAHCPVL 378
Query: 153 IVKKPKSTT 161
IVK P+ ++
Sbjct: 379 IVKGPRKSS 405
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.137 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,657,637
Number of Sequences: 36976
Number of extensions: 53392
Number of successful extensions: 308
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 297
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 297
length of query: 165
length of database: 9,014,727
effective HSP length: 89
effective length of query: 76
effective length of database: 5,723,863
effective search space: 435013588
effective search space used: 435013588
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC149080.7


