

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
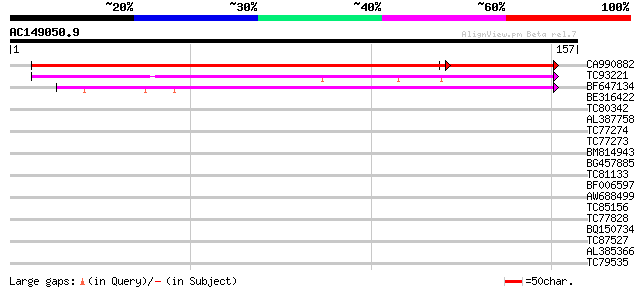
Query= AC149050.9 - phase: 0 /pseudo
(157 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CA990882 similar to GP|16648746|gb AT5g19750/T29J13_170 {Arabido... 228 1e-75
TC93221 similar to PIR|T51590|T51590 membrane protein peroxisom... 62 1e-10
BF647134 similar to GP|15450882|gb Unknown protein {Arabidopsis ... 61 2e-10
BE316422 similar to GP|9294052|dbj| gene_id:MOB24.15~ref|NP_0133... 36 0.006
TC80342 similar to GP|6692267|gb|AAF24617.1| putative phosphatid... 29 0.69
AL387758 weakly similar to GP|15146316|gb AT5g67220/K21H1_18 {Ar... 28 2.0
TC77274 similar to GP|22136956|gb|AAM91707.1 unknown protein {Ar... 27 2.6
TC77273 PIR|A26014|A26014 histone H3 - wheat, complete 27 2.6
BM814943 similar to PIR|H96569|H96 unknown protein 54928-56750 ... 27 3.4
BG457885 similar to PIR|E96622|E966 hypothetical protein F23H11.... 26 5.8
TC81133 similar to PIR|T05493|T05493 pathogenesis-related protei... 26 5.8
BF006597 similar to PIR|E96622|E966 hypothetical protein F23H11.... 26 5.8
AW688499 similar to GP|4234953|gb|A NBS-LRR-like protein cD7 {Ph... 26 7.6
TC85156 GP|1435157|emb|CAA58445.1 histone H3 variant H3.3 {Lycop... 26 7.6
TC77828 similar to GP|16648867|gb|AAL24285.1 HSP associated prot... 26 7.6
BQ150734 25 9.9
TC87527 similar to GP|21689681|gb|AAM67462.1 unknown protein {Ar... 25 9.9
AL385366 25 9.9
TC79535 weakly similar to GP|10178054|dbj|BAB11418. gene_id:MUB3... 25 9.9
>CA990882 similar to GP|16648746|gb AT5g19750/T29J13_170 {Arabidopsis
thaliana}, partial (58%)
Length = 687
Score = 228 bits (580), Expect(2) = 1e-75
Identities = 116/116 (100%), Positives = 116/116 (100%)
Frame = +2
Query: 7 YMALLAKYPVPVKALTSAILNLIGDLICQLVIDKVQTPDLKRTFLFSFLGLVLVGPTLHF 66
YMALLAKYPVPVKALTSAILNLIGDLICQLVIDKVQTPDLKRTFLFSFLGLVLVGPTLHF
Sbjct: 176 YMALLAKYPVPVKALTSAILNLIGDLICQLVIDKVQTPDLKRTFLFSFLGLVLVGPTLHF 355
Query: 67 WYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRPSQAVPKLKQ 122
WYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRPSQAVPKLKQ
Sbjct: 356 WYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRPSQAVPKLKQ 523
Score = 72.0 bits (175), Expect(2) = 1e-75
Identities = 31/33 (93%), Positives = 32/33 (96%)
Frame = +3
Query: 120 LKQEWFSAVLANWQLWIPFQFLNFRFVPQQFQV 152
L +EWFSAVLANWQLWIPFQFLNFRFVPQQFQV
Sbjct: 516 LSREWFSAVLANWQLWIPFQFLNFRFVPQQFQV 614
>TC93221 similar to PIR|T51590|T51590 membrane protein peroxisomal
[imported] - Arabidopsis thaliana, partial (88%)
Length = 694
Score = 61.6 bits (148), Expect = 1e-10
Identities = 39/149 (26%), Positives = 82/149 (54%), Gaps = 3/149 (2%)
Frame = +3
Query: 7 YMALLAKYPVPVKALTSAILNLIGDLICQLVIDKVQTPDLKRTFLFSFLGLVLVGPTLHF 66
Y+ L ++P+ K +T+ +L+ I D++ Q + +Q +KR L LG +GP H+
Sbjct: 123 YVKQLQEHPLRTKVITAGVLSGISDIVSQKLTG-IQKLQVKRLLLKVLLGAGYLGPFGHY 299
Query: 67 WYLYLSQLVTLPGTSGAIL-RLVLDQFVFSPIFLGVFLSSL-VTLEGRPSQAVP-KLKQE 123
+++ L ++ S ++ R++++Q SP+ +F+ + +EG+P V ++K+
Sbjct: 300 FHIILEKIFKGKKDSKTVIKRVLIEQLTSSPLNNLIFMIYYGLVIEGQPWVNVKARVKKG 479
Query: 124 WFSAVLANWQLWIPFQFLNFRFVPQQFQV 152
+ S A+W W ++N++F+P F+V
Sbjct: 480 YPSVQKASWTFWPVVGWINYKFMPLHFRV 566
>BF647134 similar to GP|15450882|gb Unknown protein {Arabidopsis thaliana},
partial (61%)
Length = 613
Score = 60.8 bits (146), Expect = 2e-10
Identities = 45/158 (28%), Positives = 69/158 (43%), Gaps = 19/158 (12%)
Frame = +1
Query: 14 YPVPVK-ALTSAILNLIGDLICQL-----------------VIDKVQTP-DLKRTFLFSF 54
Y P+K ALT+A L L GD I QL V+ K+ + DL R +
Sbjct: 136 YQFPLKQALTAASLTLTGDTIAQLSNRWNKAKESGENASQDVLSKLLSEHDLLRALRMTS 315
Query: 55 LGLVLVGPTLHFWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRPS 114
G + GP WY L + P +L+++L+Q V P + V + + + S
Sbjct: 316 YGFLFYGPGSFAWYQLLDHCLPKPNVQNLMLKVLLNQVVLGPCVIAVIFAWNNLWQQKLS 495
Query: 115 QAVPKLKQEWFSAVLANWQLWIPFQFLNFRFVPQQFQV 152
+ K K++ +L ++ WIP LNF VP +V
Sbjct: 496 ELPEKYKRDALPTLLYGFRFWIPVSVLNFWVVPLPXRV 609
>BE316422 similar to GP|9294052|dbj| gene_id:MOB24.15~ref|NP_013352.1~similar
to unknown protein {Arabidopsis thaliana}, partial (40%)
Length = 408
Score = 36.2 bits (82), Expect = 0.006
Identities = 26/115 (22%), Positives = 47/115 (40%), Gaps = 27/115 (23%)
Frame = +3
Query: 7 YMALLAKYPVPVKALTSAILNLIGDLICQLVIDKVQTP---------------------D 45
Y L+ +PV + TS +L +GD+ Q + D
Sbjct: 63 YQNSLSVHPVRTQVATSGVLWAVGDVTAQYITHSAAASSSSKKRLQLSATKAADDKFVID 242
Query: 46 LKRTFLFSFLGLVLVGPTLHFWYLYLSQLVT-----LPGTSGAI-LRLVLDQFVF 94
+R + S G+ VGP HFWY L + ++ +P T+ ++ ++ +D +F
Sbjct: 243 WRRVAVTSMFGVGFVGPVGHFWYEGLEKFISHKLQLMPQTARSVATKVAMDGLIF 407
>TC80342 similar to GP|6692267|gb|AAF24617.1| putative
phosphatidylinositolglycan class N short form
{Arabidopsis thaliana}, partial (18%)
Length = 1181
Score = 29.3 bits (64), Expect = 0.69
Identities = 18/49 (36%), Positives = 27/49 (54%), Gaps = 2/49 (4%)
Frame = -2
Query: 8 MALLAKYPVPVKALTSAILNLIGDLICQLVIDKVQTPDLK--RTFLFSF 54
+A+LAK+PVP KA+ + + G L + D + P L+ TFL F
Sbjct: 265 LAILAKFPVPKKAMLNKTIKANGILTSESCKDLLSYPRLR*FVTFLKEF 119
>AL387758 weakly similar to GP|15146316|gb AT5g67220/K21H1_18 {Arabidopsis
thaliana}, partial (37%)
Length = 488
Score = 27.7 bits (60), Expect = 2.0
Identities = 12/29 (41%), Positives = 17/29 (58%)
Frame = +2
Query: 3 LYYRYMALLAKYPVPVKALTSAILNLIGD 31
L Y+ L KYPVP + + S + L+GD
Sbjct: 263 LLIEYLNLCEKYPVPWRIIRSHVHKLLGD 349
>TC77274 similar to GP|22136956|gb|AAM91707.1 unknown protein {Arabidopsis
thaliana}, partial (80%)
Length = 2013
Score = 27.3 bits (59), Expect = 2.6
Identities = 13/34 (38%), Positives = 22/34 (64%)
Frame = -3
Query: 46 LKRTFLFSFLGLVLVGPTLHFWYLYLSQLVTLPG 79
+ RT L++ G L+ ++ FWYL +S+ T+PG
Sbjct: 298 ISRTSLWN--GSFLIRSSVLFWYLRISRSATVPG 203
>TC77273 PIR|A26014|A26014 histone H3 - wheat, complete
Length = 697
Score = 27.3 bits (59), Expect = 2.6
Identities = 13/34 (38%), Positives = 22/34 (64%)
Frame = -2
Query: 46 LKRTFLFSFLGLVLVGPTLHFWYLYLSQLVTLPG 79
+ RT L++ G L+ ++ FWYL +S+ T+PG
Sbjct: 294 ISRTSLWN--GSFLIRSSVLFWYLRISRSATVPG 199
>BM814943 similar to PIR|H96569|H96 unknown protein 54928-56750 [imported] -
Arabidopsis thaliana, partial (28%)
Length = 481
Score = 26.9 bits (58), Expect = 3.4
Identities = 15/58 (25%), Positives = 29/58 (49%), Gaps = 1/58 (1%)
Frame = +3
Query: 85 LRLVLDQFVFSPIFLGVFLSSLVTLE-GRPSQAVPKLKQEWFSAVLANWQLWIPFQFL 141
+++ DQ +S ++ ++ + + L P +LK +F + A W+LW PF L
Sbjct: 24 VKVAFDQTAWSAVWNSIYYTVVGILRFDSPINIFNELKARFFPMLTAGWKLW-PFAHL 194
>BG457885 similar to PIR|E96622|E966 hypothetical protein F23H11.16
[imported] - Arabidopsis thaliana, partial (22%)
Length = 677
Score = 26.2 bits (56), Expect = 5.8
Identities = 21/52 (40%), Positives = 26/52 (49%), Gaps = 6/52 (11%)
Frame = +1
Query: 43 TPDLKRTFLFSF-LGLVLVGPTLHFWYLYLSQLVTL-----PGTSGAILRLV 88
+P LK TFLFSF + L LHF L L ++ L P T AI L+
Sbjct: 130 SPTLKLTFLFSFQIALQNQKLRLHFHLLLLLLILQLFTVSEPQTPNAIFYLI 285
>TC81133 similar to PIR|T05493|T05493 pathogenesis-related protein 19K4.140
- Arabidopsis thaliana, partial (62%)
Length = 1248
Score = 26.2 bits (56), Expect = 5.8
Identities = 8/14 (57%), Positives = 10/14 (71%)
Frame = +1
Query: 124 WFSAVLANWQLWIP 137
W SA+L W+LW P
Sbjct: 634 WGSAILLQWRLWFP 675
>BF006597 similar to PIR|E96622|E966 hypothetical protein F23H11.16
[imported] - Arabidopsis thaliana, partial (24%)
Length = 505
Score = 26.2 bits (56), Expect = 5.8
Identities = 21/52 (40%), Positives = 26/52 (49%), Gaps = 6/52 (11%)
Frame = +2
Query: 43 TPDLKRTFLFSF-LGLVLVGPTLHFWYLYLSQLVTL-----PGTSGAILRLV 88
+P LK TFLFSF + L LHF L L ++ L P T AI L+
Sbjct: 74 SPTLKLTFLFSFQIALQNQKLRLHFHLLLLLLILQLFTVSEPQTPNAIFYLI 229
>AW688499 similar to GP|4234953|gb|A NBS-LRR-like protein cD7 {Phaseolus
vulgaris}, partial (2%)
Length = 677
Score = 25.8 bits (55), Expect = 7.6
Identities = 8/17 (47%), Positives = 11/17 (64%)
Frame = +1
Query: 129 LANWQLWIPFQFLNFRF 145
+ NW WIPF+ + F F
Sbjct: 205 IPNWNEWIPFEGIKFAF 255
>TC85156 GP|1435157|emb|CAA58445.1 histone H3 variant H3.3 {Lycopersicon
esculentum}, complete
Length = 754
Score = 25.8 bits (55), Expect = 7.6
Identities = 21/67 (31%), Positives = 30/67 (44%), Gaps = 10/67 (14%)
Frame = -3
Query: 46 LKRTFLFSFLGLVLVGPTLHFWYLYLSQLVTLPG----------TSGAILRLVLDQFVFS 95
+ RT L+ G L+ ++ FWY +S+ T+PG GA LR L +
Sbjct: 284 ISRTNLWK--GSFLIKSSVLFWYFLISRRATVPGR*RCGFLTPPVVGADLRAAL----VA 123
Query: 96 PIFLGVF 102
FLG F
Sbjct: 122 SCFLGAF 102
>TC77828 similar to GP|16648867|gb|AAL24285.1 HSP associated protein like
{Arabidopsis thaliana}, partial (83%)
Length = 1587
Score = 25.8 bits (55), Expect = 7.6
Identities = 15/40 (37%), Positives = 19/40 (47%)
Frame = -2
Query: 25 ILNLIGDLICQLVIDKVQTPDLKRTFLFSFLGLVLVGPTL 64
+L L G LIC L P FLFSF ++ PT+
Sbjct: 929 LLLLFGLLICCLSFSTATLPLTLSFFLFSFFAQPIILPTV 810
>BQ150734
Length = 1042
Score = 25.4 bits (54), Expect = 9.9
Identities = 13/27 (48%), Positives = 16/27 (59%)
Frame = +3
Query: 46 LKRTFLFSFLGLVLVGPTLHFWYLYLS 72
L R FLFSF + GPTL W + +S
Sbjct: 153 LCRFFLFSFYQVFQSGPTLAGWIVPMS 233
>TC87527 similar to GP|21689681|gb|AAM67462.1 unknown protein {Arabidopsis
thaliana}, partial (96%)
Length = 1582
Score = 25.4 bits (54), Expect = 9.9
Identities = 14/27 (51%), Positives = 17/27 (62%), Gaps = 1/27 (3%)
Frame = -2
Query: 81 SGAILR-LVLDQFVFSPIFLGVFLSSL 106
S A+LR L F+PIFLG+F S L
Sbjct: 762 SAALLRALAASALEFTPIFLGIFNSRL 682
>AL385366
Length = 444
Score = 25.4 bits (54), Expect = 9.9
Identities = 10/25 (40%), Positives = 13/25 (52%)
Frame = +1
Query: 120 LKQEWFSAVLANWQLWIPFQFLNFR 144
L++EW S + W WI F L R
Sbjct: 259 LREEWTSIMQ*YWSFWIEFGLLQCR 333
>TC79535 weakly similar to GP|10178054|dbj|BAB11418.
gene_id:MUB3.3~pir||T03813~similar to unknown protein
{Arabidopsis thaliana}, partial (86%)
Length = 1356
Score = 25.4 bits (54), Expect = 9.9
Identities = 21/77 (27%), Positives = 30/77 (38%)
Frame = -1
Query: 51 LFSFLGLVLVGPTLHFWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLE 110
L S G L T F + LS + P T + + L +S LV
Sbjct: 1242 LVSIFGATLSSVTTGFTCMTLSGMTLSPSTLASTSKCNLT------------VSGLVLHS 1099
Query: 111 GRPSQAVPKLKQEWFSA 127
G+PS+ +P +FSA
Sbjct: 1098 GKPSKLIPSPSFSFFSA 1048
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.332 0.146 0.456
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,362,886
Number of Sequences: 36976
Number of extensions: 77557
Number of successful extensions: 561
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 556
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 560
length of query: 157
length of database: 9,014,727
effective HSP length: 88
effective length of query: 69
effective length of database: 5,760,839
effective search space: 397497891
effective search space used: 397497891
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 54 (25.4 bits)
Medicago: description of AC149050.9


