

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
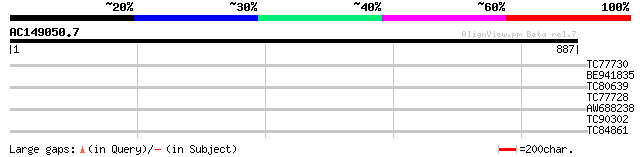
Query= AC149050.7 + phase: 0 /pseudo
(887 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC77730 weakly similar to GP|7341113|gb|AAF61210.1| unknown {Gly... 30 3.1
BE941835 weakly similar to PIR|T06404|T06 resistance complex pro... 30 3.1
TC80639 weakly similar to PIR|G96621|G96621 probable disease res... 30 3.1
TC77728 similar to GP|12056928|gb|AAG48132.1 putative resistance... 30 4.1
AW688238 weakly similar to EGAD|60190|6299 putative disease-resi... 30 4.1
TC90302 weakly similar to PIR|H96561|H96561 probable peptide tra... 29 6.9
TC84861 similar to GP|13877699|gb|AAK43927.1 protein phosphatase... 29 9.1
>TC77730 weakly similar to GP|7341113|gb|AAF61210.1| unknown {Glycine max},
partial (80%)
Length = 831
Score = 30.4 bits (67), Expect = 3.1
Identities = 16/49 (32%), Positives = 27/49 (54%)
Frame = +1
Query: 23 TMKHFKYLTQHKKITTNLEEELERLKMIKQALQTRVETERRKGYEIAPN 71
T + +YLT+H++ N E+ +ERL+ K AL + GY +P+
Sbjct: 349 TCSYGEYLTKHEEGFQNNEKNMERLRQWKMAL---TQAANLSGYHYSPH 486
>BE941835 weakly similar to PIR|T06404|T06 resistance complex protein I2C-2 -
tomato, partial (3%)
Length = 630
Score = 30.4 bits (67), Expect = 3.1
Identities = 13/30 (43%), Positives = 18/30 (59%)
Frame = +1
Query: 171 HLWDGRSGKNHNGERSHQNNRKK*AI*RSC 200
H W+GR+GKNH + S + +* * SC
Sbjct: 304 HSWNGRTGKNHYSQESF*QPKGR*TF*LSC 393
>TC80639 weakly similar to PIR|G96621|G96621 probable disease resistance
protein F23H11.10 [imported] - Arabidopsis thaliana,
partial (4%)
Length = 836
Score = 30.4 bits (67), Expect = 3.1
Identities = 13/30 (43%), Positives = 19/30 (63%)
Frame = +1
Query: 171 HLWDGRSGKNHNGERSHQNNRKK*AI*RSC 200
HLW+GR+ KNH+ E S + + +* SC
Sbjct: 715 HLWNGRTRKNHSSEESF*QPKGRKIL*LSC 804
>TC77728 similar to GP|12056928|gb|AAG48132.1 putative resistance protein
{Glycine max}, partial (12%)
Length = 765
Score = 30.0 bits (66), Expect = 4.1
Identities = 19/82 (23%), Positives = 43/82 (52%)
Frame = +1
Query: 28 KYLTQHKKITTNLEEELERLKMIKQALQTRVETERRKGYEIAPNVQKWLYDVTTIENELQ 87
++LT+H++ N E+ +ERL+ K AL ++ GY +P+ ++ + ++
Sbjct: 418 EHLTKHEERFQNNEKNMERLRQWKIAL---IQAANLSGYHYSPHGYEYKF--------IE 564
Query: 88 KWLSDDNAQI*HLIIHWERKPL 109
K + D + I H+ ++ + P+
Sbjct: 565 KIVEDISNNINHVFLNVAKYPV 630
>AW688238 weakly similar to EGAD|60190|6299 putative disease-resistance clone
pH359-3 {Oryza sativa}, partial (18%)
Length = 690
Score = 30.0 bits (66), Expect = 4.1
Identities = 12/30 (40%), Positives = 19/30 (63%)
Frame = +2
Query: 171 HLWDGRSGKNHNGERSHQNNRKK*AI*RSC 200
H W+GR+GKNH+ ++S + * +* C
Sbjct: 143 HCWNGRTGKNHSSQKSF*QPKGS*EL*LLC 232
>TC90302 weakly similar to PIR|H96561|H96561 probable peptide transporter
[imported] - Arabidopsis thaliana, partial (25%)
Length = 933
Score = 29.3 bits (64), Expect = 6.9
Identities = 13/33 (39%), Positives = 19/33 (57%)
Frame = +1
Query: 842 FHHCKKWTSKIVPIWNFFHEDLVVHLSLRVSAW 874
FHHCK T + + NF D+V +L+ S+W
Sbjct: 280 FHHCK*GTWQYSKLRNFAQHDIVFDGTLQPSSW 378
>TC84861 similar to GP|13877699|gb|AAK43927.1 protein phosphatase type
2C-like protein {Arabidopsis thaliana}, partial (74%)
Length = 914
Score = 28.9 bits (63), Expect = 9.1
Identities = 10/16 (62%), Positives = 11/16 (68%), Gaps = 2/16 (12%)
Frame = +2
Query: 337 WWLASCNCD--SWKST 350
WW+ SCNCD WK T
Sbjct: 614 WWINSCNCDINRWKKT 661
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.362 0.160 0.608
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,706,868
Number of Sequences: 36976
Number of extensions: 497093
Number of successful extensions: 5230
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 1568
Number of HSP's successfully gapped in prelim test: 210
Number of HSP's that attempted gapping in prelim test: 3610
Number of HSP's gapped (non-prelim): 1915
length of query: 887
length of database: 9,014,727
effective HSP length: 105
effective length of query: 782
effective length of database: 5,132,247
effective search space: 4013417154
effective search space used: 4013417154
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC149050.7


