

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149039.3 + phase: 0 /pseudo
(2185 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
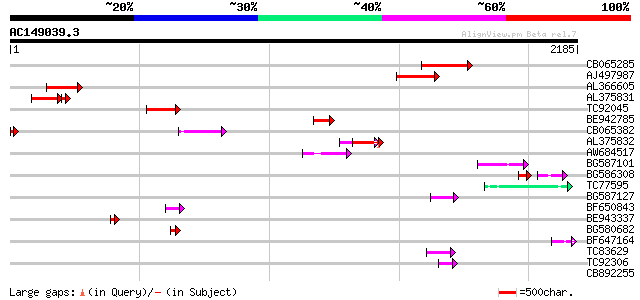
Score E
Sequences producing significant alignments: (bits) Value
CB065285 weakly similar to PIR|A84500|A84 probable retroelement ... 402 e-111
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 297 3e-80
AL366605 279 1e-74
AL375831 170 3e-58
TC92045 201 2e-51
BE942785 149 1e-35
CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fra... 134 3e-31
AL375832 132 2e-30
AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thalia... 126 1e-28
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 111 4e-24
BG586308 weakly similar to PIR|F84528|F8 probable retroelement p... 60 8e-18
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 76 1e-13
BG587127 weakly similar to PIR|H84506|H84 probable retroelement ... 66 1e-10
BF650843 similar to EGAD|117930|126 hypothetical protein F11A5.h... 64 5e-10
BE943337 61 5e-09
BG580682 53 1e-06
BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis t... 50 1e-05
TC83629 weakly similar to GP|6692261|gb|AAF24611.1| putative RNa... 48 4e-05
TC92306 46 1e-04
CB892255 37 0.065
>CB065285 weakly similar to PIR|A84500|A84 probable retroelement gag/pol
polyprotein [imported] - Arabidopsis thaliana, partial
(2%)
Length = 592
Score = 402 bits (1032), Expect = e-111
Identities = 191/197 (96%), Positives = 194/197 (97%)
Frame = -2
Query: 1585 KSKDCEEPLIDEGPDPNSKWGLVFDGAVNAYGKGIGAVIVSPQGHHIPFTARILFECTNN 1644
KSKDCEEPLI EGPDPNSKWGLVFDGAVNAYGKGIGAVIVSPQGH+IPFTARILFECTNN
Sbjct: 591 KSKDCEEPLIGEGPDPNSKWGLVFDGAVNAYGKGIGAVIVSPQGHYIPFTARILFECTNN 412
Query: 1645 MAEYEACIFGIEEAIDMRIKHLDIYGDSALVINQIKGEWETHHAKLIPYRDYARRLLTYF 1704
MAEYEACIFGIEEAIDMRIKHLDIYGDSALVINQIKGEWETHHA LIPYRDYARRLLTYF
Sbjct: 411 MAEYEACIFGIEEAIDMRIKHLDIYGDSALVINQIKGEWETHHANLIPYRDYARRLLTYF 232
Query: 1705 TKVELHHIPRDENQMADALATLSSMFRVNHWNDVPIIKVQRLERPSHVFAIGDVIDQAGE 1764
TKVELHHIPRDENQMADALATLSSMFRVNHWNDVPIIKVQRLERPSHVFAIG+VIDQ GE
Sbjct: 231 TKVELHHIPRDENQMADALATLSSMFRVNHWNDVPIIKVQRLERPSHVFAIGNVIDQTGE 52
Query: 1765 NVVDYKPWYYDIKQFLL 1781
NVVDYKPWYYDIKQFL+
Sbjct: 51 NVVDYKPWYYDIKQFLV 1
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 297 bits (761), Expect = 3e-80
Identities = 131/167 (78%), Positives = 155/167 (92%)
Frame = -2
Query: 1489 KTCCALAWAAKRLRHYLVNHTTWLISRMDPIKYIFEKAVVTGKIARWQVLLSEYDIVFKT 1548
KTCCALAWAAKRLRHY++NHTTWL+S+MDPIKYIFEK +TG+IARWQ+LLSEYDI +++
Sbjct: 635 KTCCALAWAAKRLRHYMINHTTWLVSKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRS 456
Query: 1549 QKAIKGSILADHLAYQPLDDYQPIEFDFPDEEIMYLKSKDCEEPLIDEGPDPNSKWGLVF 1608
QKAIKGSILADHLA+QPL+DY+PI+FDFPDEEIMYLK KDC+EPL EGPDP+S WGL+F
Sbjct: 455 QKAIKGSILADHLAHQPLEDYRPIKFDFPDEEIMYLKMKDCDEPLFGEGPDPDSVWGLIF 276
Query: 1609 DGAVNAYGKGIGAVIVSPQGHHIPFTARILFECTNNMAEYEACIFGI 1655
DGAVN YG GIGAV+++P+G HIPFTAR+ F+CTNN+AEYEACI GI
Sbjct: 275 DGAVNVYGNGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGI 135
>AL366605
Length = 422
Score = 279 bits (713), Expect = 1e-74
Identities = 130/139 (93%), Positives = 136/139 (97%)
Frame = -3
Query: 143 EIKANRGNGDSIKTQDLCLVSKVDVPKKFKVPEFDKYNGLTCPQNHIVKYVRKMSNYKDN 202
EIKANRGN DS KTQDLCLVSKVDVPKKFK+P+FD+YNGLTCPQNHI+KYVRKM NYKDN
Sbjct: 417 EIKANRGNADSFKTQDLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKDN 238
Query: 203 DSLMIHCFQDSLMEDAAEWYTSLSKDDVHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQ 262
DSLMIHCFQDSLMEDAAEWYTSLSK+D+HTFDELAAAFKSHYGFNTRLKPNREFLRSLSQ
Sbjct: 237 DSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQ 58
Query: 263 KKEESFREYAQRWRGAAAR 281
KKEESFREYAQRWRGAAAR
Sbjct: 57 KKEESFREYAQRWRGAAAR 1
>AL375831
Length = 467
Score = 170 bits (431), Expect(2) = 3e-58
Identities = 80/119 (67%), Positives = 92/119 (77%)
Frame = +2
Query: 83 ISGANLTPVSTALTQAATTVTEPIVNTVPQFVHANAHRGSIATTGTMEERMEELAKELRR 142
I N + L Q VTEP+V+T+PQ ++ N SI T TMEE MEELAKELR
Sbjct: 8 IPATNAASIIATLPQTTAAVTEPLVHTLPQGININTQHRSIPVTKTMEEMMEELAKELRH 187
Query: 143 EIKANRGNGDSIKTQDLCLVSKVDVPKKFKVPEFDKYNGLTCPQNHIVKYVRKMSNYKD 201
EI+ANRGN DS+KTQDLCLVSKVDVPKKFK+P+FD+YNGLTCPQNHI+KYVRKM NYKD
Sbjct: 188 EIQANRGNADSVKTQDLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKD 364
Score = 75.5 bits (184), Expect(2) = 3e-58
Identities = 33/36 (91%), Positives = 35/36 (96%)
Frame = +3
Query: 200 KDNDSLMIHCFQDSLMEDAAEWYTSLSKDDVHTFDE 235
K NDSLMIHCFQDSLMEDAAEWYTSLSK+D+HTFDE
Sbjct: 360 KTNDSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDE 467
>TC92045
Length = 873
Score = 201 bits (512), Expect = 2e-51
Identities = 105/131 (80%), Positives = 111/131 (84%), Gaps = 1/131 (0%)
Frame = -3
Query: 528 KGPXGIFVQNFRTLQSQSRWHFLIGPQIHXAGYK*MCQIQIKKKISHITKTEISFSLQYQ 587
KGP GIFVQNFRTLQSQSRWHFLIGPQIH AGYK*MCQIQIKKK + +F L +
Sbjct: 868 KGPRGIFVQNFRTLQSQSRWHFLIGPQIHMAGYK*MCQIQIKKKKNQSHHKNRNFFLSPK 689
Query: 588 -LDPKQKITKKFSKNTKNKSSQNQQIRTGFTQNQLKSSQNQQIRTGIIQNHRESPHQHQI 646
+ K + +KFSKNTKNKSSQNQQIRTGFTQNQLKSSQNQQIRTGIIQNHRES HQHQI
Sbjct: 688 SVRSKTENHQKFSKNTKNKSSQNQQIRTGFTQNQLKSSQNQQIRTGIIQNHRESSHQHQI 509
Query: 647 RTGFSSKSGTT 657
RTGFSSKSGTT
Sbjct: 508 RTGFSSKSGTT 476
>BE942785
Length = 460
Score = 149 bits (376), Expect = 1e-35
Identities = 79/84 (94%), Positives = 79/84 (94%)
Frame = -3
Query: 1169 VESLLPFMERKHT*LASYHLSLA**QGLQRGPLFKD*LSKVRSPRGMELQWLL*RMHREP 1228
V SLLPFMERKHT*LASYHLSLA* QGLQRGPLFKD*LSKVRSPRGMELQWLL*RMHREP
Sbjct: 377 VXSLLPFMERKHT*LASYHLSLA*KQGLQRGPLFKD*LSKVRSPRGMELQWLL*RMHREP 198
Query: 1229 SKRAKLPAGAG*YSFVRTSIRKVL 1252
SKR KLPAGAG*YSFVRTSIR L
Sbjct: 197 SKRVKLPAGAG*YSFVRTSIRIAL 126
>CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fragment),
partial (9%)
Length = 624
Score = 134 bits (338), Expect = 3e-31
Identities = 94/186 (50%), Positives = 108/186 (57%)
Frame = -3
Query: 651 SSKSGTTLNRCNINRQFNSSSSNMLDQLFLLYQCCMQSFFQLYSREDTARLGRASLHRTL 710
+S+ GT ++R N+ SSN LD FL YQC M S +LY E TA+ R SLH
Sbjct: 544 ASRPGTIIHRFT-----NNDSSNRLDLPFL*YQCYMLSSSRLYFSEGTAQSDRGSLHLIR 380
Query: 711 YLQGSALILSVTSIKAL*GMMLRGAML*STSSRSL*IKGS*LLRTMCHMSSIIRSQMMLP 770
YL GS L +V SIK L MM + AML*+T RS K S*LLRT MSS Q+ L
Sbjct: 379 YLPGSGLTSNVISIKVLWVMMSKDAML*NTL*RSSLTKES*LLRTTSLMSSTTLFQITLL 200
Query: 771 *I*LRYVKRFPCWMSAMLQLLWCRSISSCAKLYCLNMIMLSAKNVPAILWAAMSSKTMSR 830
* *LRY+++ P S MLQLLWC SS AK CL MIM AKN ILW A K SR
Sbjct: 199 *T*LRYMRKLPDLTSVMLQLLWCHYTSSYAKPLCLTMIMPIAKNDSIILWVAALFKMTSR 20
Query: 831 A*WTAI 836
A* T I
Sbjct: 19 A**TTI 2
Score = 53.9 bits (128), Expect = 7e-07
Identities = 27/31 (87%), Positives = 29/31 (93%)
Frame = -1
Query: 1 NTQLRTELASLREELAKAHDAMTALLAAQEQ 31
N QLRTELAS+REELAKA+D MTALLAAQEQ
Sbjct: 618 NAQLRTELASMREELAKANDTMTALLAAQEQ 526
>AL375832
Length = 535
Score = 132 bits (331), Expect = 2e-30
Identities = 72/132 (54%), Positives = 81/132 (60%), Gaps = 12/132 (9%)
Frame = +3
Query: 1321 FQNNNPLALPKARVIFSVRACLFRHCR*LIKNCRFFHDCFFLVPFFSWK*W*CQKKPKYL 1380
F+NNNPLALPKARVIFSVRACLFRHCR + KNCRFFHDCFF FF K W*CQKKPK L
Sbjct: 156 FKNNNPLALPKARVIFSVRACLFRHCRLMNKNCRFFHDCFFFSAFFLEK-W*CQKKPKRL 332
Query: 1381 YFLINFKNLHKKPSF*NHPTILRRLNYNEPVGHSNPAFP------------PNFEFPVHD 1428
YFLINFKNLH+K + + + H P FPV++
Sbjct: 333 YFLINFKNLHEKKKIQKTKKLFLKSS-----NHYMQVEP**TR*TQ*FYASSKL*FPVYE 497
Query: 1429 AENEEDDDIPYE 1440
+ E DDIPYE
Sbjct: 498 PKMREGDDIPYE 533
Score = 118 bits (296), Expect = 2e-26
Identities = 80/162 (49%), Positives = 87/162 (53%), Gaps = 7/162 (4%)
Frame = +1
Query: 1269 GLSTQSQKKQLDLARGLHLLHQEALREIGMPLTFPQSCMFLSNIPCCLSILFFQNNNPLA 1328
GL S KKQLDL GLHLLHQEALR IG PLTF QSCMF SN CC S LF +
Sbjct: 1 GLLMLSLKKQLDLVSGLHLLHQEALRVIGTPLTFLQSCMFPSNASCCFSALFSKITILSL 180
Query: 1329 LPKARVIFSVRACLFRHCR*LIKNCRFFHDCFFLVPFFSWK*W*CQKKPKYLYFLINFKN 1388
PK F F * IK F FFLVP FSWK +K FL+ K
Sbjct: 181 CPKRE*YFL*GRVCFGIAV**IKIVVSFTTAFFLVP-FSWKNGNARKNLNVCIFLLISKT 357
Query: 1389 LHKKP-------SF*NHPTILRRLNYNEPVGHSNPAFPPNFE 1423
KK SF*NHP I+ RLN+N+PV HSN PNF+
Sbjct: 358 CMKKKKYKKQKNSF*NHPIIICRLNHNKPVEHSNSMPLPNFD 483
>AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thaliana chromosome
II BAC F26H6; putative retroelement pol polyprotein,
partial (1%)
Length = 488
Score = 126 bits (316), Expect = 1e-28
Identities = 88/187 (47%), Positives = 103/187 (55%)
Frame = +2
Query: 1130 SPFRSWILMPLTAVSWGDHGYTTREQ*PLLCTKS*NLQKVESLLPFMERKHT*LASYHLS 1189
S FRSWI M AVSW D GY T +*PL T+S*NL +++ L
Sbjct: 2 SRFRSWISMLRIAVSWEDLGYMTLAR*PLPYTRS*NL*RMDLL----------------- 130
Query: 1190 LA**QGLQRGPLFKD*LSKVRSPRGMELQWLL*RMHREPSKRAKLPAGAG*YSFVRTSIR 1249
R LFK L KV+SPR +ELQWLL*R R +R +LP GA * + VR S R
Sbjct: 131 --------RELLFKGCLWKVQSPRRLELQWLL*RTPRRLFRRDRLPTGAS*SNSVRISER 286
Query: 1250 KVLAFPRL*EFPPGHSIVLGLSTQSQKKQLDLARGLHLLHQEALREIGMPLTFPQSCMFL 1309
KV P+ P IV GLS K+ LDL G L +QEALR+IGMPL FPQSCM
Sbjct: 287 KVSDSPQHRVSPRELFIVPGLSIHLLKRWLDLFLGPCL*YQEALRKIGMPLMFPQSCMCP 466
Query: 1310 SNIPCCL 1316
N+ CCL
Sbjct: 467 XNVSCCL 487
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 111 bits (277), Expect = 4e-24
Identities = 63/196 (32%), Positives = 95/196 (48%)
Frame = +2
Query: 1803 RFLLDGDILYKRNYDMVLLRCVDEHEAEQLMHDVHDGTFRTHATGHTMSRKLLRAGYYWM 1862
R+L D LYK+ D + +RCV E E ++ H + H K+ +AG++W
Sbjct: 17 RYLWDEPFLYKQCADNIYIRCVAEEEIPGILFHCHGSNYAGHFAVSKTVSKIQQAGFWWP 196
Query: 1863 AMEHDCYQHARKCHKCQIYADKIHVPPHALNVISSPWPFSMWGIDMIGRIEPKASNGHHF 1922
M D + KC CQ + N I F +WGID +G +N +
Sbjct: 197 TMFKDAHSFISKCDPCQRQGNIS*RNEMPQNFILEVEVFDVWGIDFMGPFPSSYNN--KY 370
Query: 1923 ILVAIDYFTKWVEAASYTNVTKQVVAKFIKNNIICRYGVPSKIITDNGTNLNNNVVQALC 1982
ILVA+DY +KWVEA + VV K K+ I R+GVP +I+D G++ N V + L
Sbjct: 371 ILVAVDYVSKWVEAIASPTNDATVVVKMFKSVIFPRFGVPRVVISDGGSHFINKVFEKLL 550
Query: 1983 EEFKIEHHNSSPYRPQ 1998
++ + H ++ Y PQ
Sbjct: 551 KKNGVRHKVATAYHPQ 598
>BG586308 weakly similar to PIR|F84528|F8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 686
Score = 60.1 bits (144), Expect(2) = 8e-18
Identities = 39/122 (31%), Positives = 68/122 (54%), Gaps = 4/122 (3%)
Frame = -3
Query: 2032 LHGYRTTVRSSTGATPFSLVYGMEAVLPLEVEIPSL---RVIMEAKLSEAEWCQSRYDQL 2088
L +RT R +T +TPFS+ + +EA+ P EV + SL R+ +L+ ++ L
Sbjct: 462 LWSHRTNPRGATKSTPFSMAHRVEAMAPAEVNVTSL*RSRMPQNIELNN----DRLFNAL 295
Query: 2089 NLIEEKRMDAVARGQSYQARMKTAFDKKVHPREFKVGELVLKRRISQQPD-PRGKWTPNY 2147
IEE+R A+ R Q+YQ ++++ ++K V + K+G++VL + + GK N+
Sbjct: 294 ETIEERRDQALLRIQNYQHQIESYYNKTVKSQPLKLGDIVLCKVFENTKELNAGKLGTNW 115
Query: 2148 EG 2149
EG
Sbjct: 114 EG 109
Score = 50.4 bits (119), Expect(2) = 8e-18
Identities = 22/52 (42%), Positives = 35/52 (67%)
Frame = -2
Query: 1959 YGVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEATNKNI 2010
+G+P +I+TDNG++ +N + CE ++I + +SP PQ NG EA+NK I
Sbjct: 685 HGLPYEIVTDNGSHFISNKFREFCERWRIRLNTASPRYPQSNGQAEASNKII 530
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial (14%)
Length = 1708
Score = 76.3 bits (186), Expect = 1e-13
Identities = 75/353 (21%), Positives = 139/353 (39%), Gaps = 15/353 (4%)
Frame = +2
Query: 1831 QLMHDVHDGTFRTHATGHTMSRKLLRAGYYWMAMEHDCYQHARKCHKCQIYADKIHVPPH 1890
+L+ + HD T H G + +++ ++W + R C C IH+
Sbjct: 176 KLVQESHDSTAAGHP-GRNGTLEIVSRKFFWPGQSQTVRRFVRNCDVC----GGIHIWRQ 340
Query: 1891 ALNVISSPWPF-----SMWGIDMIGRIEPKASNGHHFILVAIDYFTKWVEAASYTNVTKQ 1945
A P P S +D I + P G ++ V +D +K V + +
Sbjct: 341 AKRGFLKPLPVPNRLHSDLSMDFITSLPPTRGRGSQYLWVIVDRLSKSVTLEEMDTMEAE 520
Query: 1946 VVAKFIKNNIICRYGVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEA 2005
A+ + +G+P I++D G+N + C + S+ Y PQ +G E
Sbjct: 521 ACAQRFLSCHYRFHGMPQSIVSDRGSNWVGRFWREFCRLTGVTQLLSTSYHPQTDGGTER 700
Query: 2006 TNKNIKRIVQKMVTTYKD-WHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVEI 2064
N+ I+ +++ V +D W ++LP R SS GATPF + +G V P+
Sbjct: 701 WNQEIQAVLRAYVCWSQDNWGDLLPTVQLALRNRHNSSIGATPFFVEHGYH-VDPIPTVE 877
Query: 2065 PSLRVIMEAKLSEAEWCQSRYDQLNLIEEKRMDAVARGQSYQARMKTAFDKKVHPREFKV 2124
+ V+ E + + + D I+ + + A R ++ + + D+ ++V
Sbjct: 878 DTGGVVSEGEAAAQLLVKRMKDVTGFIQAEIVAAQQRSEASANKRRCPADR------YQV 1039
Query: 2125 GELV---LKRRISQQPDPRGKW------TPNYEGPYVVKKAFSGGALILTHMD 2168
G+ V + S +P + W + P+VV+ G H+D
Sbjct: 1040 GDKVWLNVSNYKSPRPSKKLDWLHHKYEVTRFVTPHVVELNVPGTVYPKFHVD 1198
>BG587127 weakly similar to PIR|H84506|H84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(13%)
Length = 415
Score = 66.2 bits (160), Expect = 1e-10
Identities = 38/109 (34%), Positives = 55/109 (49%)
Frame = +3
Query: 1620 GAVIVSPQGHHIPFTARILFECTNNMAEYEACIFGIEEAIDMRIKHLDIYGDSALVINQI 1679
G + SP + + R+ F +NN YEA I G+ A ++I+++ Y DS LV +Q
Sbjct: 3 GIRLTSPTNEILKQSFRLEFHASNNETNYEALIAGVRLAHGLKIRNIHAYCDSQLVASQF 182
Query: 1680 KGEWETHHAKLIPYRDYARRLLTYFTKVELHHIPRDENQMADALATLSS 1728
GE+E + Y ++L L IPR EN ADALA L+S
Sbjct: 183 SGEYEARDELMDTYLKLVQKLAQKLDYFALTRIPRSENVQADALAALAS 329
>BF650843 similar to EGAD|117930|126 hypothetical protein F11A5.h
{Caenorhabditis elegans}, partial (1%)
Length = 566
Score = 64.3 bits (155), Expect = 5e-10
Identities = 35/76 (46%), Positives = 43/76 (56%), Gaps = 2/76 (2%)
Frame = -3
Query: 600 KNTKNKSSQNQQIRTGFTQNQLKSSQNQQIRTGIIQNHRESPHQHQIRTGFSSKSGTT-- 657
+ + QNQQ G QNQL+ QNQQIRTG QNHRESPHQHQ F +K+ T
Sbjct: 558 RTPRTPHXQNQQX*XGXAQNQLQPPQNQQIRTGFTQNHRESPHQHQNVPDFRAKTTPTEG 379
Query: 658 LNRCNINRQFNSSSSN 673
+ R R+ S S +
Sbjct: 378 IARAKEKRRIRSFSED 331
>BE943337
Length = 496
Score = 60.8 bits (146), Expect = 5e-09
Identities = 28/37 (75%), Positives = 31/37 (83%)
Frame = -2
Query: 387 FQKIIQKGILVISCSGTHFSPCVCL*VFSICLYKSLS 423
+QK+IQKGILVISCSGTHF PC CL*VFS C S+S
Sbjct: 435 WQKVIQKGILVISCSGTHFHPCDCL*VFSDCFLHSIS 325
Score = 33.5 bits (75), Expect = 0.93
Identities = 47/171 (27%), Positives = 68/171 (39%), Gaps = 1/171 (0%)
Frame = -3
Query: 346 LFFSESLLSF*NIFYCLCLSFLALQVI*ELSETSHYFSCILFQKIIQKGILVISCSGTHF 405
++F S+ +F+ SFLA+ I L S F + + + LV S H
Sbjct: 449 VYFKIGKKSYRRVFW----SFLAVGPIFILVTVSESFQIVFYIPSVN---LVKFISNLHL 291
Query: 406 SPCVCL*VFSICLYKSLSSLY*ENNFKNCI*VNSSQSSP*GRFGLFLLIFRQGGI-LKYL 464
S C ++ L++S+ +L G F F F + LKYL
Sbjct: 290 SQCQV----NLSLFRSILAL--------------------GAFWSFHTKFWHREVTLKYL 183
Query: 465 MSLIGSF*SFC*F*F*FHFTFVKITFFSSILFLNHNLVQNFLIFIFCPFLI 515
+ IGSF*S F F FTF +IT N + F+ FI FL+
Sbjct: 182 I*AIGSF*SVLSILFVFRFTFGQIT----------NFIFIFIAFIIFAFLV 60
>BG580682
Length = 509
Score = 52.8 bits (125), Expect = 1e-06
Identities = 26/39 (66%), Positives = 28/39 (71%)
Frame = -1
Query: 618 QNQLKSSQNQQIRTGIIQNHRESPHQHQIRTGFSSKSGT 656
QNQL+S QNQQIRTG QNHRES HQHQ F K+ T
Sbjct: 509 QNQLQSPQNQQIRTGFTQNHRESSHQHQNVPDF*EKTTT 393
>BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana},
partial (6%)
Length = 469
Score = 49.7 bits (117), Expect = 1e-05
Identities = 35/101 (34%), Positives = 52/101 (50%), Gaps = 3/101 (2%)
Frame = +1
Query: 2087 QLNLIEEKRMDAVARGQSYQARMKTAFDKKVHPREFKVGELVL--KRRISQQPDPRGKWT 2144
+LN +EE R+DA + Y+ R K D+++ REF+ GELVL R+ P GK
Sbjct: 103 ELNELEELRLDAYENAKIYKERTKKWHDRRIIRREFREGELVLLFNSRLKLFP---GKLR 273
Query: 2145 PNYEGPYVVKKAFSGGAL-ILTHMDGVELPNPVNADIVKKY 2184
++ GP+ VK GA+ + + G P VN +K Y
Sbjct: 274 SHWSGPFQVKNVMPSGAVEVWSESTG---PFTVNGQRLKHY 387
>TC83629 weakly similar to GP|6692261|gb|AAF24611.1| putative RNase H
{Arabidopsis thaliana}, partial (21%)
Length = 1071
Score = 48.1 bits (113), Expect = 4e-05
Identities = 34/113 (30%), Positives = 50/113 (44%)
Frame = +2
Query: 1605 GLVFDGAVNAYGKGIGAVIVSPQGHHIPFTARILFECTNNMAEYEACIFGIEEAIDMRIK 1664
G V+ +N G GA++++ G + + L T AEY A + G++ A K
Sbjct: 563 G*VW*SNINRGPAGAGALLLAEDGSLLYGFRQGLGHQTKESAEYRALLLGLKHASMKGFK 742
Query: 1665 HLDIYGDSALVINQIKGEWETHHAKLIPYRDYARRLLTYFTKVELHHIPRDEN 1717
++ GDS LVINQI W+ L A L F + HI R+ N
Sbjct: 743 YVTAKGDSELVINQILDPWKIKDEHLKKLCAEALELSDNFHSFRIQHISRERN 901
>TC92306
Length = 521
Score = 46.2 bits (108), Expect = 1e-04
Identities = 24/73 (32%), Positives = 36/73 (48%)
Frame = +2
Query: 1652 IFGIEEAIDMRIKHLDIYGDSALVINQIKGEWETHHAKLIPYRDYARRLLTYFTKVELHH 1711
I G+ EA + +H+ + GD V Q G W+ ++ L + A L + F V + H
Sbjct: 2 ILGLNEARNQGYEHVHVRGDFQXVCKQFXGSWKVNNPNLRNLCNXAVELKSNFKSVSVEH 181
Query: 1712 IPRDENQMADALA 1724
+PR N ADA A
Sbjct: 182 VPRGXNXAADAQA 220
>CB892255
Length = 853
Score = 37.4 bits (85), Expect = 0.065
Identities = 28/98 (28%), Positives = 46/98 (46%)
Frame = +1
Query: 2051 VYGMEAVLPLEVEIPSLRVIMEAKLSEAEWCQSRYDQLNLIEEKRMDAVARGQSYQARMK 2110
V+ ++A+ ++V++ S+ V +AK+ C R ++RG S +
Sbjct: 31 VFNVKALCHMKVKVLSMEVPKKAKID*T*KCYCRVP---------WTVISRG*SKYST-- 177
Query: 2111 TAFDKKVHPREFKVGELVLKRRISQQPDPRGKWTPNYE 2148
KK PRE + VLK + P+ RGKW PNY+
Sbjct: 178 ----KKARPRECQESGFVLKNML---PNSRGKWMPNYK 270
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.340 0.148 0.488
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 71,763,814
Number of Sequences: 36976
Number of extensions: 1151696
Number of successful extensions: 9508
Number of sequences better than 10.0: 60
Number of HSP's better than 10.0 without gapping: 5257
Number of HSP's successfully gapped in prelim test: 519
Number of HSP's that attempted gapping in prelim test: 3917
Number of HSP's gapped (non-prelim): 6368
length of query: 2185
length of database: 9,014,727
effective HSP length: 111
effective length of query: 2074
effective length of database: 4,910,391
effective search space: 10184150934
effective search space used: 10184150934
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 66 (30.0 bits)
Medicago: description of AC149039.3


