

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
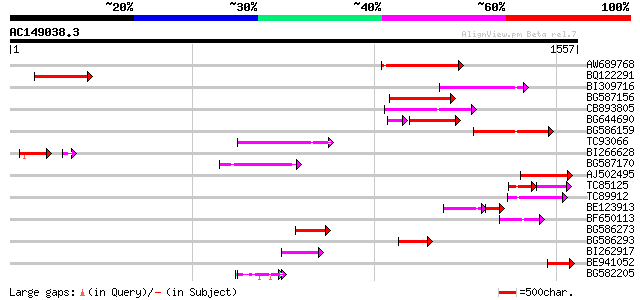
Query= AC149038.3 + phase: 0
(1557 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsi... 286 3e-77
BQ122291 260 3e-69
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 176 5e-44
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 169 1e-41
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 167 4e-41
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 132 2e-39
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 150 5e-36
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 141 2e-33
BI266628 127 7e-32
BG587170 similar to PIR|F86470|F8 probable retroelement polyprot... 133 5e-31
AJ502495 weakly similar to GP|18071369|g putative gag-pol polypr... 120 4e-27
TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-relate... 68 4e-23
TC89912 weakly similar to PIR|B84512|B84512 probable retroelemen... 107 5e-23
BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sati... 89 2e-21
BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vu... 97 5e-20
BG586273 weakly similar to PIR|F86470|F8 probable retroelement p... 85 2e-16
BG586293 weakly similar to PIR|E84473|E84 probable retroelement ... 85 2e-16
BI262917 weakly similar to GP|19920130|g Putative retroelement {... 84 4e-16
BE941052 weakly similar to PIR|B85188|B85 retrotransposon like p... 76 9e-14
BG582205 75 2e-13
>AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsis thaliana},
partial (9%)
Length = 675
Score = 286 bits (733), Expect = 3e-77
Identities = 138/228 (60%), Positives = 174/228 (75%), Gaps = 1/228 (0%)
Frame = +1
Query: 1020 TRAKHGIVQKRKHPTLLLTHIEPTGYRQA-MKQPQWLQAMQLEHEALMKNNTWTLVPLPA 1078
TR K G ++ P + LT P + P+WLQAM+ E++AL+ N TW LVPLP
Sbjct: 4 TRTKTGKLK----PKVFLTEPCPQDCENLPLSDPRWLQAMKTEYKALIDNKTWDLVPLPP 171
Query: 1079 DRQAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDYKETFSPVVKPVTVRSVLTL 1138
++A+GCKWV+R K+NPDGS+NK+KARLVAKGF Q G DY ETFSPV+KPVT+R +LT+
Sbjct: 172 HKKAIGCKWVYRVKENPDGSVNKFKARLVAKGFSQTLGCDYTETFSPVIKPVTIRLILTI 351
Query: 1139 AVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAVDPSLVCKLNKALYGLKQAPRAWF 1198
A+T KW IQQ+D+NNAFLNG+L+EEVYM+QP GFEA + SLVCKLNK+LYGLKQAPRAW+
Sbjct: 352 AITYKWEIQQIDINNAFLNGFLQEEVYMSQPQGFEAANKSLVCKLNKSLYGLKQAPRAWY 531
Query: 1199 ERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVDDILITGSSAS 1246
E L S ++ GF S+CDPSL I + N ++ +YVDDILITGSSAS
Sbjct: 532 EXLTSAQIQFGFTKSRCDPSLLIYNQNGACIYLXIYVDDILITGSSAS 675
>BQ122291
Length = 487
Score = 260 bits (664), Expect = 3e-69
Identities = 128/159 (80%), Positives = 148/159 (92%)
Frame = -2
Query: 69 DAAREAGTENPAYTEWEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQM 128
DAAREAGT NPAYTEWE++DSLLCTWILS IS SLLSRFV LR S QVWDEIH+YC+TQM
Sbjct: 486 DAAREAGTRNPAYTEWEDKDSLLCTWILSRISPSLLSRFVLLRHSWQVWDEIHSYCFTQM 307
Query: 129 RTRSRQLRSELRTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFN 188
+TRSRQLRSELR+ITKG+R+++EFIARIR+ISESL SIGDPV+HRDLIE VLEALPEEF+
Sbjct: 306 KTRSRQLRSELRSITKGSRTVSEFIARIRAISESLASIGDPVSHRDLIEVVLEALPEEFD 127
Query: 189 PIVATVNSQTEVISLDELESQLLTQEARNEKFKKALVGE 227
PIVA+VN+++EV+SLDELESQLLTQE+R EKFKKA + E
Sbjct: 126 PIVASVNAKSEVVSLDELESQLLTQESRKEKFKKAAMNE 10
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 176 bits (447), Expect = 5e-44
Identities = 98/244 (40%), Positives = 145/244 (59%)
Frame = +2
Query: 1180 VCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVDDIL 1239
VC+L K++YGLKQA R W+ +L +L+ G+ S D SLF + T +LVYVDDI+
Sbjct: 20 VCELQKSIYGLKQASRQWYSKLSESLISFGYLQSSSDFSLFTKFKDSSFTTLLVYVDDIV 199
Query: 1240 ITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKAN 1299
+ G+ S IQ + L F +KDLG L YFLG+EV S+ G +LL+Q+KY +LL +
Sbjct: 200 LAGNDISEIQHVKCFLIDRFKIKDLGSLRYFLGLEVARSKQG-ILLNQRKYTLELLEDSG 376
Query: 1300 MANANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFLS 1359
+P S KL S +D T +R ++G L Y+T TRP+IS++V ++ QF+S
Sbjct: 377 NLAVKSTLTPYDISLKLHNSDSPLYNDETQYRRLIGKLIYLTTTRPDISFAVQQLSQFVS 556
Query: 1360 APLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGAC 1419
P + H++A R+L+YLK GL + L L++F D+DWA+ P R+S +G
Sbjct: 557 KPQQVHYQAAIRVLQYLKTAPAKGLFYSAT---SNLKLSSFADSDWATCPTTRKSVTGYW 727
Query: 1420 ILLG 1423
+ LG
Sbjct: 728 VFLG 739
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 169 bits (427), Expect = 1e-41
Identities = 88/182 (48%), Positives = 117/182 (63%), Gaps = 1/182 (0%)
Frame = -1
Query: 1042 PTGYRQAMKQPQWLQAMQLEHEALMKNNTWTLVPLPADRQAVGCKWVFRTKQNPDGSINK 1101
P Y +AM+ +W +++ E A++KN+TW LP ++AV +W+F K DGSI +
Sbjct: 558 PRSYEEAMEDKEWKESVGAEAGAMIKNDTWYESELPKGKKAVSSRWIFTIKYKADGSIER 379
Query: 1102 YKARLVAKGFHQMPGFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLE 1161
K RLVA+GF G DY ETF+PV K T+R VL+LAV W + Q+DV NAFL G LE
Sbjct: 378 KKTRLVARGFTLTYGEDYIETFAPVAKLHTIRIVLSLAVNLGWGLWQMDVKNAFLQGELE 199
Query: 1162 EEVYMTQPPGFE-AVDPSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLF 1220
+EVYM PPG E V V +L KA+YGLKQ+PRAW+ +L +TL GF S+ D +LF
Sbjct: 198 DEVYMYPPPGLEHLVKRGNVLRLKKAIYGLKQSPRAWYNKLSTTLNGRGFRKSELDHTLF 19
Query: 1221 IL 1222
L
Sbjct: 18 TL 13
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 167 bits (422), Expect = 4e-41
Identities = 92/257 (35%), Positives = 147/257 (56%), Gaps = 4/257 (1%)
Frame = +3
Query: 1029 KRKHPTLLLTHIEPTGYRQAMKQPQWLQAMQLEHEALMKNNTWTLVPLPADRQAVGCKWV 1088
K ++ +L +PT + +A+K +W +M E EA +NNTW L L + + +G KW+
Sbjct: 3 KTQNLGMLTMTSDPTTFEEAVKSEKWRASMNNEMEATERNNTWELTDLRSGAKTIGLKWI 182
Query: 1089 FRTKQNPDGSINKYKARLVAKGFHQMPGFDYKETFSPVVKPVTVRSVLTLAVTNK---WC 1145
F+TK N +G I KYKARLVAKG+ Q G DY E F+PV + T+R V+ LA K C
Sbjct: 183 FKTKLNENGEIEKYKARLVAKGYSQQYGVDYTEVFAPVARWDTIRMVIALAAQIKRDGVC 362
Query: 1146 IQQLDVNNAFLNGYLEEEVYMTQPPGFEAVDPSLVCKLNKALYGLKQAPRAWFERLKSTL 1205
I + ++ + + + + + V + ++ +ALYGLKQAPRAW+ R+++
Sbjct: 363 IS*M*KAHSCMEN*MRKFLLINHRVM*RRV---IS*RVKRALYGLKQAPRAWYSRIEAYF 533
Query: 1206 LKLGFCSSKCDPSLFI-LHANQHSTFMLVYVDDILITGSSASLIQQLVKKLNAEFSLKDL 1264
K GF + +LF+ L + +YVDD++ G+ ++ ++ K + EF++ DL
Sbjct: 534 TKEGFEKCPYEHTLFVKLSEGGKILIISLYVDDLIFIGNDENMFEEFKKSMKKEFNMSDL 713
Query: 1265 GKLDYFLGIEVHYSENG 1281
GK+ YFLG+EV +E G
Sbjct: 714 GKMHYFLGVEVTQNEKG 764
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 132 bits (332), Expect(2) = 2e-39
Identities = 68/141 (48%), Positives = 93/141 (65%), Gaps = 1/141 (0%)
Frame = -2
Query: 1099 INKYKARLVAKGFHQMPGFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNG 1158
I + K++LV +G++Q G DY E FSPV + +R ++ A + + Q+DV +AF+NG
Sbjct: 424 ITRNKSKLVVQGYNQKEGIDYDEAFSPVARMEAIRILIAFAAFMGFKLYQMDVKSAFING 245
Query: 1159 YLEEEVYMTQPPGFEAVD-PSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDP 1217
L+EEV++ QPPGFE + P+ V +LNK LYGLKQAPRAW+ERL LLK GF K D
Sbjct: 244 DLKEEVFVKQPPGFEDAEVPNHVFRLNKTLYGLKQAPRAWYERLSKFLLKNGFKRGKIDN 65
Query: 1218 SLFILHANQHSTFMLVYVDDI 1238
+LF+L + VYVDDI
Sbjct: 64 TLFLLKRE*ELLIIQVYVDDI 2
Score = 50.4 bits (119), Expect(2) = 2e-39
Identities = 19/56 (33%), Positives = 32/56 (56%)
Frame = -3
Query: 1037 LTHIEPTGYRQAMKQPQWLQAMQLEHEALMKNNTWTLVPLPADRQAVGCKWVFRTK 1092
++ IEP ++A++ W+ +MQ E ++ W LVP P + +G +WVFR K
Sbjct: 600 ISSIEPKNVKEALRDADWINSMQEELHQFERSKVWYLVPRPEGKTVIGTRWVFRNK 433
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 150 bits (378), Expect = 5e-36
Identities = 82/221 (37%), Positives = 134/221 (60%), Gaps = 1/221 (0%)
Frame = +1
Query: 1273 IEVHYSENGSLLLSQKKYIQDLLVKANMANANGIASPMASSTKLTKYGSNHVSDPTFFRS 1332
+EV +E G + + Q+KY+ DLL + M +N +P+A KL K + D T ++
Sbjct: 1 VEVIQNEEG-IYICQRKYVTDLLERFGMEKSNLSRNPIAPRCKLIKDENGVKVDATKYKQ 177
Query: 1333 IVGGLQYVTVTRPEISYSVNKVCQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMH 1392
IVG L Y+ TRP++ Y ++ + +F++ P E H AVKR+LRYL GTI+ G++ +
Sbjct: 178 IVGCLMYLAATRPDLMYVLSLISRFMNCPTELHMHAVKRVLRYLNGTINLGIMYK---RN 348
Query: 1393 QPLSLTAFCDADWASDPDDRRSTSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQA 1452
L A+ D+D+A D DDR+STSG +L +SW +KKQ +V S+ +AE+ + A
Sbjct: 349 GSEKLEAYTDSDYAGDLDDRKSTSGYVFMLSSGAVSWSSKKQPVVTLSTTKAEFIAAAFC 528
Query: 1453 SAEVLWIQSLLKEL-KVPTAIPQIFCDNLSTVSLAHNPVLH 1492
+ + +W++ +L++L + ++CDN ST+ L+ NPVLH
Sbjct: 529 ACQSVWMRRVLEKLGYTQSGSITMYCDNNSTIKLSKNPVLH 651
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 141 bits (355), Expect = 2e-33
Identities = 93/272 (34%), Positives = 137/272 (50%), Gaps = 8/272 (2%)
Frame = +1
Query: 626 DFCSACCLGKSHRLPSVSSKTVYNKPF-ELIFCDLWGPASVESHGGYSYFLICVDAYSRY 684
+FC + + S S+ T K + I DLWGP+ V S+GG Y + +D + R
Sbjct: 16 EFCKHLLFFGNRKKVSFSTATHRTKGILDYIHSDLWGPSKVTSYGGRRYMMTIIDDFPRK 195
Query: 685 TWIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEF--RPFTQFLTTLGITHRL 742
W++ L+ K+ T TF+ ++ +VE Q +K + TD EF F +F T GI
Sbjct: 196 VWVYFLRYKNETFPTFKKWRILVETQTGKNVKKLITDN*LEFCSSDFNEFCTNHGIARHK 375
Query: 743 TCPHTHHQNGSVERKHRHIVETGLTLLANAKL--PLHYWDHAFLTATYLINRLPSPILNN 800
T P QNG ER R ++E +L+NA L W A TA +L+NR P L+
Sbjct: 376 TIPRNPQQNGVAERMIRTLLERARCMLSNAGL*N*RDLWVEAASTACHLVNRSPHSALDF 555
Query: 801 KSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYK--CL 858
K P + + DY L+ FGC + N+ KL R+ EC+FL Y+ KGY+ C
Sbjct: 556 KVPEDIWSGNLVDYSNLRIFGCPAYALV---NDGKLAPRAGECIFLSYASESKGYRLWCS 726
Query: 859 DP-TGRMFISKDVIFNEYKFPYSELFTSGQSS 889
DP + ++ +S+DV FNE L +SG+ S
Sbjct: 727 DPKSQKLILSRDVTFNE-----DALLSSGKQS 807
>BI266628
Length = 536
Score = 127 bits (318), Expect(2) = 7e-32
Identities = 65/93 (69%), Positives = 73/93 (77%), Gaps = 7/93 (7%)
Frame = -1
Query: 28 ISVKLDETNY-------LQWKQQVEGVLRGTKMVRHVVSPQIPPVFLNDAAREAGTENPA 80
+ +K+D +N LQWKQQVEGVLR TK V+ V+SPQIPPV+L D AREAGT+ PA
Sbjct: 455 VVIKVDPSNKQVLS*VRLQWKQQVEGVLRETKTVKFVISPQIPPVYLTDEAREAGTKKPA 276
Query: 81 YTEWEEQDSLLCTWILSTISSSLLSRFVRLRFS 113
YTEWEEQDSLLCTWILSTI SLLSRFV LR S
Sbjct: 275 YTEWEEQDSLLCTWILSTI*PSLLSRFVLLRHS 177
Score = 30.4 bits (67), Expect(2) = 7e-32
Identities = 19/37 (51%), Positives = 22/37 (59%)
Frame = -2
Query: 146 TRSIAEFIARIRSISESLMSIGDPVAHRDLIETVLEA 182
+R +AE IARIR V+HRDLIE VLEA
Sbjct: 88 SRIVAEIIARIRRNFR--------VSHRDLIEVVLEA 2
Score = 29.6 bits (65), Expect = 9.5
Identities = 20/53 (37%), Positives = 28/53 (52%), Gaps = 5/53 (9%)
Frame = -3
Query: 116 VWDEIHNYCYTQMRTRSRQ-----LRSELRTITKGTRSIAEFIARIRSISESL 163
VWDEIH+YC+ Q + R+ L ++ +S+ EF A ISESL
Sbjct: 174 VWDEIHSYCFMQSKDSCSSHS*CFTRTRLHALS--LKSLLEFGA----ISESL 34
>BG587170 similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 718
Score = 133 bits (335), Expect = 5e-31
Identities = 80/229 (34%), Positives = 119/229 (51%), Gaps = 3/229 (1%)
Frame = -3
Query: 576 PIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVMKLCNQQLPNKSFTDF-CSACCLG 634
P+ S SS+SL+ +WH RLGHPH + N LP F + C AC LG
Sbjct: 641 PVSNYKCSFTSSSSLNKDALWHARLGHPHGRAL-------NLMLPGVVFENKNCEACILG 483
Query: 635 KSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLICVDAYSRYTWIFPLKLKS 694
K + + TVY F+LI+ DLW S+ S + YF+ +D S+YTW+ + K
Sbjct: 482 KHCKNVFPRTSTVYENCFDLIYTDLWTAPSL-SRDNHKYFVTFIDEKSKYTWLTLIPSKD 306
Query: 695 HTLITFQNFKTMVELQYNLPIKSVQTDGGGEFR--PFTQFLTTLGITHRLTCPHTHHQNG 752
+ F+NF+ V Y+ IK +++D GGE+ F L GI H+ +CP+T QNG
Sbjct: 305 RVIDAFKNFQAYVTNHYHAKIKILRSDNGGEYTSYAFKSHLDHHGILHQTSCPYTPQQNG 126
Query: 753 SVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLINRLPSPILNNK 801
+RK++H++E +L+ A + TA YLIN +P+ +L K
Sbjct: 125 VAKRKNKHLMEVARSLMFQANVS---------TACYLINWIPTKVLRIK 6
>AJ502495 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (9%)
Length = 542
Score = 120 bits (301), Expect = 4e-27
Identities = 55/144 (38%), Positives = 91/144 (63%), Gaps = 1/144 (0%)
Frame = +2
Query: 1403 ADWASDPDDRRSTSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSL 1462
+DWA D + R+STSG LG ISW +KKQ +VA S+AEAEY + + + +W++ +
Sbjct: 2 SDWAGDTETRKSTSGYAFHLGTGAISWSSKKQPVVAFSTAEAEYIASTSCATQTVWLRRI 181
Query: 1463 LKELKVPTAIP-QIFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPA 1521
L+ + P +I+CDN S ++L+ NPV H R+KH+++ +RE + K++++ + P
Sbjct: 182 LEVMHHEQNTPTKIYCDNKSAIALSKNPVFHGRSKHIDIQFHKIRELIAEKEVVIEYCPT 361
Query: 1522 QYQVADILTKPLSASRFLELRNKL 1545
+ ++ADI TKPL F +L+ L
Sbjct: 362 EEKIADIFTKPLKIESFYKLKKML 433
>TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
(EC 3.4.23.-);, partial (7%)
Length = 705
Score = 68.2 bits (165), Expect(2) = 4e-23
Identities = 34/96 (35%), Positives = 54/96 (55%)
Frame = +3
Query: 1448 SLAQASAEVLWIQSLLKELKVPTAIPQIFCDNLSTVSLAHNPVLHSRTKHMELDIFFVRE 1507
SL QA E +W+Q L++EL ++CD+ S + +A NP HSRTKH+ + FVRE
Sbjct: 228 SLPQACKEAIWMQRLMEELGHKQEQITVYCDSQSALHIARNPAFHSRTKHIGIQYHFVRE 407
Query: 1508 KVISKDLIVSHIPAQYQVADILTKPLSASRFLELRN 1543
V + + I +AD +TK ++ +F+ R+
Sbjct: 408 VVEEGSVDMQKIHTNDNLADAMTKSINTDKFIWCRS 515
Score = 59.7 bits (143), Expect(2) = 4e-23
Identities = 32/78 (41%), Positives = 47/78 (60%)
Frame = +1
Query: 1369 VKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGACILLGPNLIS 1428
VKRI+RY+KGT + + L++ + D+D+A D D R+ST+G L +S
Sbjct: 1 VKRIMRYIKGTSGVAVCFGGSE----LTVRGYVDSDFAGDHDKRKSTTGYVFTLAGGAVS 168
Query: 1429 WWAKKQTLVARSSAEAEY 1446
W +K QT+VA S+ EAEY
Sbjct: 169 WLSKLQTVVALSTTEAEY 222
>TC89912 weakly similar to PIR|B84512|B84512 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(10%)
Length = 814
Score = 107 bits (266), Expect = 5e-23
Identities = 52/165 (31%), Positives = 97/165 (58%)
Frame = +1
Query: 1367 KAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGACILLGPNL 1426
+A+K +L+YL ++ L A + +L + DAD+A + D R+S SG L
Sbjct: 1 QALKWVLKYLNESLKSSLKYTKAAQEED-ALEGYVDADYAGNVDTRKSLSGFVFTLYGTT 177
Query: 1427 ISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVPTAIPQIFCDNLSTVSLA 1486
ISW A +Q++V S+ +AEY + + + +W++ ++ EL + +I CD+ S + LA
Sbjct: 178 ISWKANQQSVVTLSTTQAEYIAFVEGVKDAIWLKGMIGELGITQEYVKIHCDSQSAIHLA 357
Query: 1487 HNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTK 1531
++ V H RTKH+++ + F+R+ + SK+++V + ++ AD+ TK
Sbjct: 358 NHQVYHERTKHIDIRLHFIRDMIESKEIVVEKMASEENPADVFTK 492
>BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 503
Score = 88.6 bits (218), Expect(2) = 2e-21
Identities = 48/119 (40%), Positives = 69/119 (57%), Gaps = 1/119 (0%)
Frame = +1
Query: 1192 QAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQ-HSTFMLVYVDDILITGSSASLIQQ 1250
Q+PR WF+R + K G+ + D ++FI H++ ++VYVDDI +TG I++
Sbjct: 1 QSPRDWFDRFT*VVKKFGYIQCQTDHAMFIKHSSTVKKAILIVYVDDIFLTGDHGK*IKR 180
Query: 1251 LVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKANMANANGIASP 1309
L L EF +KDLG L YFLG+EV + GS +SQ+KY+ DLL + M I P
Sbjct: 181 LKNLLAEEFEIKDLGNLKYFLGMEVARWKKGS-SISQRKYVLDLLKETRMIGCKTIRDP 354
Score = 33.9 bits (76), Expect(2) = 2e-21
Identities = 17/51 (33%), Positives = 31/51 (60%)
Frame = +2
Query: 1308 SPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFL 1358
+PM ++ KL + + D ++ +VG L Y++ TRP+IS+ V + QF+
Sbjct: 350 TPMDATVKLGTLDNGTLVDKGRYQRLVGKLIYLSHTRPDISFVVCTMSQFM 502
>BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vulgaris},
partial (13%)
Length = 494
Score = 97.1 bits (240), Expect = 5e-20
Identities = 52/124 (41%), Positives = 72/124 (57%)
Frame = +1
Query: 1344 RPEISYSVNKVCQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDA 1403
RP+I YSV+ + +F+ P + H A RILRY++GT+ +GLL + L + D+
Sbjct: 121 RPDICYSVSVISKFMHDPRKPHLIAANRILRYVRGTMEYGLLFPYGAKSEVYELICYSDS 300
Query: 1404 DWASDPDDRRSTSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLL 1463
DW DRRSTSG ISW KKQ + A SS EAEY + A+ + LW+ S++
Sbjct: 301 DWCG---DRRSTSGYVFKFNDAAISWCTKKQPITALSSYEAEYIAGTFATFQALWLDSVI 471
Query: 1464 KELK 1467
KELK
Sbjct: 472 KELK 483
>BG586273 weakly similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 705
Score = 84.7 bits (208), Expect = 3e-16
Identities = 40/97 (41%), Positives = 63/97 (64%), Gaps = 1/97 (1%)
Frame = -2
Query: 786 ATYLINRLPSPILNNKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVF 845
A YLINR+P+ +L +++PF +L+ + P +++ FGC C+ +KLE RS++ +F
Sbjct: 704 ACYLINRIPTRVLKDQAPFEVLNQRKPSLTYMRVFGCLCYVLVPGELRNKLEARSRKAMF 525
Query: 846 LGYSPSHKGYKCLDPTG-RMFISKDVIFNEYKFPYSE 881
+GYS + KGYKC DP R+ +S+DV F E + Y E
Sbjct: 524 IGYSTTQKGYKCYDPEARRVLVSRDVKFIEERGYYEE 414
>BG586293 weakly similar to PIR|E84473|E84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(7%)
Length = 763
Score = 84.7 bits (208), Expect = 3e-16
Identities = 41/93 (44%), Positives = 60/93 (64%)
Frame = +2
Query: 1067 KNNTWTLVPLPADRQAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDYKETFSPV 1126
+ T LV P + +G +W+++ K+N DG++ KYKARLVAKG+ + G D+ E F+PV
Sbjct: 47 QKQTLKLVKKPTGVKPIGLRWIYKIKRNEDGTLIKYKARLVAKGYVKQQGIDFDEVFAPV 226
Query: 1127 VKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGY 1159
V+ T+ +L LA TN I +DV AFLNG+
Sbjct: 227 VRIETI*LLLALAATNGC*IHHIDVKIAFLNGH 325
>BI262917 weakly similar to GP|19920130|g Putative retroelement {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (8%)
Length = 426
Score = 84.0 bits (206), Expect = 4e-16
Identities = 42/115 (36%), Positives = 63/115 (54%)
Frame = +1
Query: 746 HTHHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLINRLPSPILNNKSPFF 805
HT QNG ER +R ++E +L A + +W A TA Y+INR PS +++ K+P
Sbjct: 82 HTPQQNGVAERMNRTLLERTRAMLKTAGMAKSFWAEAVKTACYVINRSPSTVIDLKTPME 261
Query: 806 LLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYKCLDP 860
+ + DY L FGC + KL+ +S++C+FLGY+ + KGY DP
Sbjct: 262 MWKGKPVDYSSLHVFGCPVYVMYNSQERTKLDPKSRKCIFLGYADNVKGYXLWDP 426
>BE941052 weakly similar to PIR|B85188|B85 retrotransposon like protein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 480
Score = 76.3 bits (186), Expect = 9e-14
Identities = 40/73 (54%), Positives = 47/73 (63%)
Frame = +2
Query: 1477 CDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPLSAS 1536
CD LS L HNPV HSR KH+ +DI FVR+ V L V H+ Q+AD LTKPLS S
Sbjct: 29 CDYLSATYLTHNPVYHSRMKHISIDIHFVRDLVQQGKLKVQHVCTVDQLADCLTKPLSKS 208
Query: 1537 RFLELRNKLRVSD 1549
R LRNK+ V+D
Sbjct: 209 RHQLLRNKIGVTD 247
>BG582205
Length = 801
Score = 75.1 bits (183), Expect = 2e-13
Identities = 62/151 (41%), Positives = 79/151 (52%), Gaps = 15/151 (9%)
Frame = +3
Query: 625 TDFCSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLICVDAYSR- 683
+DFCS CCLGK+HRLPS F L+ S+ + S+F+IC D SR
Sbjct: 384 SDFCSFCCLGKAHRLPS--------------FHPLYLTLSLLN----SFFVICGDYSSRV 509
Query: 684 --------YTWIFPLKLK---SHTLITFQNFKTMVELQYNLPIKSVQTD--GGGEFRPFT 730
+++ + K SH I FQNFK +VELQY L + V ++ GGEFRPF
Sbjct: 510 FFNLCGCLFSF*LDISFKTEVSHFTI-FQNFKRIVELQYKL-LHQVCSNCLRGGEFRPFI 683
Query: 731 QFLTTLG-ITHRLTCPHTHHQNGSVERKHRH 760
+L LG H LT + GSVERKHRH
Sbjct: 684 TYLNNLG*PAHTLTT-----KMGSVERKHRH 761
Score = 43.9 bits (102), Expect = 5e-04
Identities = 47/145 (32%), Positives = 57/145 (38%), Gaps = 11/145 (7%)
Frame = +2
Query: 619 LPNKSFTDFCSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLICV 678
LPNK+ F G S + SS Y KP ELIFCDLWG +
Sbjct: 368 LPNKTL*-FLFFLLFG*ST*ITFFSSTVSYTKPLELIFCDLWG--------------LLQ 502
Query: 679 DAYSRYTWIFPLKLKSHTL----ITFQNFKTMVELQYNL-------PIKSVQTDGGGEFR 727
+ W+ L L + L +T N + +LQ N P + GG
Sbjct: 503 *SLF*PVWMPILLLIGYFL*N*SLTLYN---LSKLQKNC*ITI*TPPSSLFKLFEGG*VS 673
Query: 728 PFTQFLTTLGITHRLTCPHTHHQNG 752
PF RLTCPHTHHQNG
Sbjct: 674 PFYYIFE----*SRLTCPHTHHQNG 736
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.132 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 56,775,886
Number of Sequences: 36976
Number of extensions: 1033799
Number of successful extensions: 9797
Number of sequences better than 10.0: 392
Number of HSP's better than 10.0 without gapping: 8449
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 9425
length of query: 1557
length of database: 9,014,727
effective HSP length: 109
effective length of query: 1448
effective length of database: 4,984,343
effective search space: 7217328664
effective search space used: 7217328664
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC149038.3


