

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148996.15 + phase: 0
(153 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
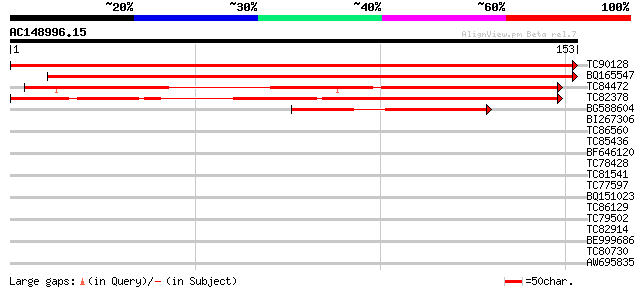
Score E
Sequences producing significant alignments: (bits) Value
TC90128 similar to GP|7770334|gb|AAF69704.1| F27J15.21 {Arabidop... 314 1e-86
BQ165547 similar to GP|7770334|gb|A F27J15.21 {Arabidopsis thali... 293 2e-80
TC84472 similar to GP|7770334|gb|AAF69704.1| F27J15.21 {Arabidop... 145 6e-36
TC82378 weakly similar to GP|7770334|gb|AAF69704.1| F27J15.21 {A... 119 5e-28
BG588604 similar to PIR|G96739|G967 F14O23.12 [imported] - Arabi... 50 4e-07
BI267306 similar to PIR|T05452|T05 hypothetical protein F7K2.160... 29 0.84
TC86560 similar to GP|8843737|dbj|BAA97285.1 myosin heavy chain-... 29 0.84
TC85436 GP|2522320|gb|AAB81011.1| asparagine synthetase {Medicag... 28 1.4
BF646120 similar to GP|17064156|gb| WRKY transcription factor 22... 28 1.4
TC78428 similar to PIR|F96595|F96595 unknown protein 25817-2483... 28 1.4
TC81541 similar to GP|22597168|gb|AAN03471.1 unknown protein {Gl... 27 2.5
TC77597 similar to GP|6041823|gb|AAF02138.1| lysyl-tRNA syntheta... 27 2.5
BQ151023 27 3.2
TC86129 similar to GP|8571476|gb|AAF76898.1| apetala2 domain-con... 27 3.2
TC79502 similar to GP|19881712|gb|AAM01113.1 Unknown protein {Or... 27 4.2
TC82914 similar to PIR|A84828|A84828 hypothetical protein At2g40... 27 4.2
BE999686 similar to SP|O70133|DDX9 ATP-dependent RNA helicase A ... 26 5.5
TC80730 weakly similar to GP|20198248|gb|AAD26886.2 expressed pr... 26 5.5
AW695835 similar to GP|14582465|gb| putative transcription facto... 26 5.5
>TC90128 similar to GP|7770334|gb|AAF69704.1| F27J15.21 {Arabidopsis
thaliana}, partial (32%)
Length = 716
Score = 314 bits (804), Expect = 1e-86
Identities = 153/153 (100%), Positives = 153/153 (100%)
Frame = +3
Query: 1 MVKITSPESPNRHHSNGGFFVTISAILALLSRKANRLKEKAKSSSTTKPIRDEEWRFDLK 60
MVKITSPESPNRHHSNGGFFVTISAILALLSRKANRLKEKAKSSSTTKPIRDEEWRFDLK
Sbjct: 45 MVKITSPESPNRHHSNGGFFVTISAILALLSRKANRLKEKAKSSSTTKPIRDEEWRFDLK 224
Query: 61 TPPKSPMAKPKKLLSNISNKALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDF 120
TPPKSPMAKPKKLLSNISNKALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDF
Sbjct: 225 TPPKSPMAKPKKLLSNISNKALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDF 404
Query: 121 SGVIYYDINGKQTREVPIRSPRASPLPGYLTRG 153
SGVIYYDINGKQTREVPIRSPRASPLPGYLTRG
Sbjct: 405 SGVIYYDINGKQTREVPIRSPRASPLPGYLTRG 503
>BQ165547 similar to GP|7770334|gb|A F27J15.21 {Arabidopsis thaliana},
partial (32%)
Length = 649
Score = 293 bits (749), Expect = 2e-80
Identities = 142/143 (99%), Positives = 143/143 (99%)
Frame = -1
Query: 11 NRHHSNGGFFVTISAILALLSRKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKSPMAKP 70
NRHHSNGGFFVTISAILALLSRKA+RLKEKAKSSSTTKPIRDEEWRFDLKTPPKSPMAKP
Sbjct: 649 NRHHSNGGFFVTISAILALLSRKASRLKEKAKSSSTTKPIRDEEWRFDLKTPPKSPMAKP 470
Query: 71 KKLLSNISNKALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDFSGVIYYDING 130
KKLLSNISNKALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDFSGVIYYDING
Sbjct: 469 KKLLSNISNKALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDFSGVIYYDING 290
Query: 131 KQTREVPIRSPRASPLPGYLTRG 153
KQTREVPIRSPRASPLPGYLTRG
Sbjct: 289 KQTREVPIRSPRASPLPGYLTRG 221
>TC84472 similar to GP|7770334|gb|AAF69704.1| F27J15.21 {Arabidopsis
thaliana}, partial (25%)
Length = 751
Score = 145 bits (366), Expect = 6e-36
Identities = 85/147 (57%), Positives = 100/147 (67%), Gaps = 2/147 (1%)
Frame = +1
Query: 5 TSPESPN-RHHSNGGFFVTISAILALLSRKANRLKEKAKSSSTTKPIRDEEWRFDLKTPP 63
+SPES + + SN FF+TISAILA+LSR+A RLK KAK+
Sbjct: 82 SSPESLSPKRQSNSSFFITISAILAMLSRQARRLKGKAKT-------------------- 201
Query: 64 KSPMAKPKKLLSNISNKALSQFGK-KKQREEREKEGWGNGGVWQKEILMGGKCEPLDFSG 122
KKLLSN+SNKALSQFGK KK +++R+ E N GVWQKEILMGGKCEPLDFSG
Sbjct: 202 -------KKLLSNMSNKALSQFGKMKKTKKQRDNEL--NDGVWQKEILMGGKCEPLDFSG 354
Query: 123 VIYYDINGKQTREVPIRSPRASPLPGY 149
VI+YDING+QT EVP+RSPRA P Y
Sbjct: 355 VIHYDINGRQTGEVPLRSPRAIPFSRY 435
>TC82378 weakly similar to GP|7770334|gb|AAF69704.1| F27J15.21 {Arabidopsis
thaliana}, partial (37%)
Length = 568
Score = 119 bits (298), Expect = 5e-28
Identities = 67/149 (44%), Positives = 91/149 (60%)
Frame = +2
Query: 1 MVKITSPESPNRHHSNGGFFVTISAILALLSRKANRLKEKAKSSSTTKPIRDEEWRFDLK 60
MV I + ES R + F + ++ ++ ++ ++A LK KA
Sbjct: 2 MVDIITRESSKRRANR--FIINVTTLMTVMVKRAT-LKLKA------------------- 115
Query: 61 TPPKSPMAKPKKLLSNISNKALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDF 120
P S + PKKLL NISNKA+ F +K +++++ K WG+GGVWQK ILMG KCEPLDF
Sbjct: 116 APMTSEVKSPKKLLKNISNKAMP-FIEKMKKKKKGKLEWGDGGVWQKTILMGDKCEPLDF 292
Query: 121 SGVIYYDINGKQTREVPIRSPRASPLPGY 149
SGVIYYD GKQ E P+RSPRA+P+PG+
Sbjct: 293 SGVIYYDNKGKQVNEFPLRSPRATPVPGF 379
>BG588604 similar to PIR|G96739|G967 F14O23.12 [imported] - Arabidopsis
thaliana, partial (32%)
Length = 568
Score = 50.1 bits (118), Expect = 4e-07
Identities = 25/54 (46%), Positives = 34/54 (62%)
Frame = +3
Query: 77 ISNKALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDFSGVIYYDING 130
++N ++ G ++QREE +WQK ILMGGKC+ DFSGVI YD +G
Sbjct: 135 LNNDVVTMNGCEQQREE--------DSIWQKNILMGGKCQLPDFSGVIIYDSDG 272
>BI267306 similar to PIR|T05452|T05 hypothetical protein F7K2.160 -
Arabidopsis thaliana, partial (24%)
Length = 539
Score = 28.9 bits (63), Expect = 0.84
Identities = 21/72 (29%), Positives = 34/72 (47%)
Frame = +2
Query: 7 PESPNRHHSNGGFFVTISAILALLSRKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKSP 66
P +P++ H + +++I I LL + K S+TT+ ++ W K PPK
Sbjct: 68 PMAPSKPHIS--IYISIITIFLLLLNPTKSEENKNLKSTTTENECEQRWIHIRKLPPKFN 241
Query: 67 MAKPKKLLSNIS 78
+ LLSN S
Sbjct: 242 L----DLLSNCS 265
>TC86560 similar to GP|8843737|dbj|BAA97285.1 myosin heavy chain-like
{Arabidopsis thaliana}, partial (28%)
Length = 2450
Score = 28.9 bits (63), Expect = 0.84
Identities = 21/84 (25%), Positives = 41/84 (48%), Gaps = 1/84 (1%)
Frame = +2
Query: 27 LALLSRKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKSPMAKPKKLLSNISNKAL-SQF 85
L + ++ N+LKE+ K++ TTK E +T + + KL++ A+ +
Sbjct: 500 LHVAEKELNKLKEQVKNAETTKAQALVELERAQRT--VDDLTQKLKLITESRESAVKATE 673
Query: 86 GKKKQREEREKEGWGNGGVWQKEI 109
K Q +++ E G G W++E+
Sbjct: 674 AAKSQAKQKYGESDGVNGAWKEEL 745
>TC85436 GP|2522320|gb|AAB81011.1| asparagine synthetase {Medicago sativa},
complete
Length = 2147
Score = 28.1 bits (61), Expect = 1.4
Identities = 17/44 (38%), Positives = 24/44 (53%)
Frame = -1
Query: 5 TSPESPNRHHSNGGFFVTISAILALLSRKANRLKEKAKSSSTTK 48
T P SP + ++ A + LLSR +NR KEK S+ +TK
Sbjct: 557 TEPSSPQPMYKEVTPIASL-ATMKLLSRVSNRTKEKIPSNMSTK 429
>BF646120 similar to GP|17064156|gb| WRKY transcription factor 22
{Arabidopsis thaliana}, partial (21%)
Length = 615
Score = 28.1 bits (61), Expect = 1.4
Identities = 14/35 (40%), Positives = 23/35 (65%)
Frame = +1
Query: 42 KSSSTTKPIRDEEWRFDLKTPPKSPMAKPKKLLSN 76
KSSS+++P+ + + +P KSP +PK+LL N
Sbjct: 316 KSSSSSQPLSSGSFSYSSPSP-KSPHIQPKQLLVN 417
>TC78428 similar to PIR|F96595|F96595 unknown protein 25817-24837
[imported] - Arabidopsis thaliana, partial (36%)
Length = 1153
Score = 28.1 bits (61), Expect = 1.4
Identities = 22/62 (35%), Positives = 31/62 (49%)
Frame = +3
Query: 29 LLSRKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKSPMAKPKKLLSNISNKALSQFGKK 88
LL NR + + K +T + + + R +K P M K KKLL +S LSQ KK
Sbjct: 543 LLEITENRRRRRRKIDTTQQ---EGQQRITVKAPAMLSMEK-KKLLFGLSFTLLSQAKKK 710
Query: 89 KQ 90
K+
Sbjct: 711 KK 716
>TC81541 similar to GP|22597168|gb|AAN03471.1 unknown protein {Glycine max},
partial (36%)
Length = 927
Score = 27.3 bits (59), Expect = 2.5
Identities = 23/92 (25%), Positives = 42/92 (45%)
Frame = +3
Query: 7 PESPNRHHSNGGFFVTISAILALLSRKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKSP 66
P+ ++++ + VTI+ + + ++L K S + I D PPK
Sbjct: 408 PQIRDQNYDDKANIVTITVVCCSPEKIRDKLCYKGGGSIKSIEIVD---------PPKPK 560
Query: 67 MAKPKKLLSNISNKALSQFGKKKQREEREKEG 98
A+P+K + K S +KK+ E++KEG
Sbjct: 561 AAEPEK--KKEAEKPKSAEPEKKKEPEKKKEG 650
>TC77597 similar to GP|6041823|gb|AAF02138.1| lysyl-tRNA synthetase
{Arabidopsis thaliana}, partial (86%)
Length = 2157
Score = 27.3 bits (59), Expect = 2.5
Identities = 12/39 (30%), Positives = 20/39 (50%)
Frame = +1
Query: 59 LKTPPKSPMAKPKKLLSNISNKALSQFGKKKQREEREKE 97
++ PP P IS AL + KK++REE +++
Sbjct: 112 MEQPPSDPPVTAPAASETISKNALKREQKKREREEEKRK 228
>BQ151023
Length = 270
Score = 26.9 bits (58), Expect = 3.2
Identities = 13/49 (26%), Positives = 20/49 (40%)
Frame = +1
Query: 58 DLKTPPKSPMAKPKKLLSNISNKALSQFGKKKQREEREKEGWGNGGVWQ 106
D+ T P+ +P + K + + REE+ K W G WQ
Sbjct: 34 DITTINLEPLKEPPHFKA*QGEKPVLNLARTGNREEKPKGAWALKGPWQ 180
>TC86129 similar to GP|8571476|gb|AAF76898.1| apetala2 domain-containing
protein {Atriplex hortensis}, partial (41%)
Length = 2500
Score = 26.9 bits (58), Expect = 3.2
Identities = 16/38 (42%), Positives = 22/38 (57%), Gaps = 7/38 (18%)
Frame = -2
Query: 86 GKKKQREER--EKEGWGNGGVWQ-----KEILMGGKCE 116
GK+++ EE EKEGW GGV + E + G+CE
Sbjct: 1263 GKEEEEEED*GEKEGWKKGGVEE*KGDDDEEEVEGRCE 1150
>TC79502 similar to GP|19881712|gb|AAM01113.1 Unknown protein {Oryza sativa}
[Oryza sativa (japonica cultivar-group)], partial (8%)
Length = 720
Score = 26.6 bits (57), Expect = 4.2
Identities = 14/33 (42%), Positives = 18/33 (54%)
Frame = -3
Query: 44 SSTTKPIRDEEWRFDLKTPPKSPMAKPKKLLSN 76
SST+ P W + +PPKS P+ LLSN
Sbjct: 358 SSTSSPSLKFPW-IESPSPPKSSFTLPQSLLSN 263
>TC82914 similar to PIR|A84828|A84828 hypothetical protein At2g40320
[imported] - Arabidopsis thaliana, partial (35%)
Length = 798
Score = 26.6 bits (57), Expect = 4.2
Identities = 13/33 (39%), Positives = 18/33 (54%)
Frame = +3
Query: 78 SNKALSQFGKKKQREEREKEGWGNGGVWQKEIL 110
SN + + KK++ EE EKE G W K+ L
Sbjct: 246 SNNTVQEVEKKEEEEEEEKECDLFSGTWVKDEL 344
>BE999686 similar to SP|O70133|DDX9 ATP-dependent RNA helicase A (Nuclear DNA
helicase II) (NDH II) (DEAD-box protein 9) (mHEL-5).,
partial (1%)
Length = 424
Score = 26.2 bits (56), Expect = 5.5
Identities = 18/59 (30%), Positives = 25/59 (41%)
Frame = +3
Query: 49 PIRDEEWRFDLKTPPKSPMAKPKKLLSNISNKALSQFGKKKQREEREKEGWGNGGVWQK 107
P ++W KTPPK+P P K K G K+ ++K+ G G QK
Sbjct: 240 PPPPKKWGG*KKTPPKNPGCPPPKTPPGGGKKNFWGQGPPKKFFPQKKKKGGGGAPPQK 416
>TC80730 weakly similar to GP|20198248|gb|AAD26886.2 expressed protein
{Arabidopsis thaliana}, partial (27%)
Length = 827
Score = 26.2 bits (56), Expect = 5.5
Identities = 9/16 (56%), Positives = 15/16 (93%)
Frame = -2
Query: 83 SQFGKKKQREEREKEG 98
S +GKKK++++RE+EG
Sbjct: 223 SHYGKKKKKKKREREG 176
>AW695835 similar to GP|14582465|gb| putative transcription factor {Vitis
vinifera}, partial (57%)
Length = 442
Score = 26.2 bits (56), Expect = 5.5
Identities = 9/25 (36%), Positives = 16/25 (64%)
Frame = +3
Query: 126 YDINGKQTREVPIRSPRASPLPGYL 150
+ ++G+Q ++P R P SPL Y+
Sbjct: 210 HGVHGRQNHQIPQRQPPQSPLLSYV 284
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.133 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,889,516
Number of Sequences: 36976
Number of extensions: 61898
Number of successful extensions: 370
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 364
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 365
length of query: 153
length of database: 9,014,727
effective HSP length: 88
effective length of query: 65
effective length of database: 5,760,839
effective search space: 374454535
effective search space used: 374454535
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 54 (25.4 bits)
Medicago: description of AC148996.15


