

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148995.8 + phase: 0
(325 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
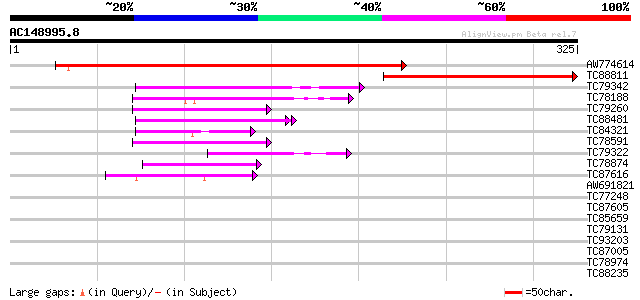
Score E
Sequences producing significant alignments: (bits) Value
AW774614 similar to PIR|T45879|T45 hypothetical protein F4P12.90... 394 e-110
TC88811 similar to PIR|T45879|T45879 hypothetical protein F4P12.... 216 7e-57
TC79342 similar to GP|10177096|dbj|BAB10430. Notchless protein h... 55 5e-08
TC78188 similar to PIR|C84870|C84870 probable splicing factor [i... 53 2e-07
TC79260 weakly similar to GP|9759350|dbj|BAB10005.1 WD-repeat pr... 52 3e-07
TC88481 similar to PIR|T00806|T02445 probable U4/U6 small nuclea... 50 9e-07
TC84321 GP|22138476|gb|AAM93460.1 hypothetical protein {Oryza sa... 45 3e-05
TC78591 similar to GP|9759350|dbj|BAB10005.1 WD-repeat protein-l... 45 4e-05
TC79322 similar to GP|13605815|gb|AAK32893.1 At2g32700/F24L7.16 ... 42 3e-04
TC78874 similar to PIR|T49098|T49098 hypothetical protein F4F15.... 41 5e-04
TC87616 similar to PIR|A96839|A96839 F23A5.2(form2) [imported] -... 41 7e-04
AW691821 similar to GP|10177570|db contains similarity to regula... 39 0.002
TC77248 similar to GP|13489182|gb|AAK27816.1 putative WD-repeat ... 39 0.004
TC87605 homologue to PIR|G96563|G96563 probable coatomer complex... 38 0.005
TC85659 SP|O24076|GBLP_MEDSA Guanine nucleotide-binding protein ... 38 0.006
TC79131 similar to GP|14031063|gb|AAK52092.1 WD-40 repeat protei... 38 0.006
TC93203 37 0.008
TC87005 similar to GP|9294061|dbj|BAB02018.1 WD40-repeat protein... 37 0.014
TC78974 similar to GP|22137238|gb|AAM91464.1 AT5g14050/MUA22_5 {... 36 0.018
TC88235 similar to GP|12656803|gb|AAK00964.1 unknown protein {Or... 36 0.023
>AW774614 similar to PIR|T45879|T45 hypothetical protein F4P12.90 -
Arabidopsis thaliana, partial (32%)
Length = 615
Score = 394 bits (1011), Expect = e-110
Identities = 190/205 (92%), Positives = 191/205 (92%), Gaps = 4/205 (1%)
Frame = +1
Query: 27 ADGNLY----NKDGAGESSFPILKDQTLFSVAHARYSKSNPIARWHICQGSINSISFSAD 82
ADGN Y NKDGAGESSFPILKDQTLFSVAHARY KSNPIARWHICQGSINSISFS D
Sbjct: 1 ADGNTYVYEKNKDGAGESSFPILKDQTLFSVAHARYRKSNPIARWHICQGSINSISFSTD 180
Query: 83 GAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQVWSM 142
GAYLATVGRDGYLRVFDYTKEHLVCGGK YYGGLLCCAWSMDGKYILTGGEDDLVQV SM
Sbjct: 181 GAYLATVGRDGYLRVFDYTKEHLVCGGKRYYGGLLCCAWSMDGKYILTGGEDDLVQVRSM 360
Query: 143 EDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQLLLWELEMDEI 202
E RKVVAWGEGHNSWVSGVA DSYW+SPNSNDNGETITYRFGSVGQDTQLLLWELEMDEI
Sbjct: 361 EHRKVVAWGEGHNSWVSGVALDSYWTSPNSNDNGETITYRFGSVGQDTQLLLWELEMDEI 540
Query: 203 VVPLRRGPPGGSPTFGSPTFSAGSQ 227
VP RRGPPGGS TFGSPTFSAGSQ
Sbjct: 541 AVP*RRGPPGGSATFGSPTFSAGSQ 615
>TC88811 similar to PIR|T45879|T45879 hypothetical protein F4P12.90 -
Arabidopsis thaliana, partial (14%)
Length = 781
Score = 216 bits (551), Expect = 7e-57
Identities = 108/111 (97%), Positives = 108/111 (97%)
Frame = +3
Query: 215 PTFGSPTFSAGSQSSHWDNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQES 274
P SPTFSAGSQSSHWDNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQES
Sbjct: 6 PNI*SPTFSAGSQSSHWDNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQES 185
Query: 275 VLTACREGHIKIWTRPGVAESQPSNSETLLATSLKEKPSLSSKISNSIYKQ 325
VLTACREGHIKIWTRPGVAESQPSNSETLLATSLKEKPSLSSKISNSIYKQ
Sbjct: 186 VLTACREGHIKIWTRPGVAESQPSNSETLLATSLKEKPSLSSKISNSIYKQ 338
>TC79342 similar to GP|10177096|dbj|BAB10430. Notchless protein homolog
{Arabidopsis thaliana}, partial (47%)
Length = 873
Score = 54.7 bits (130), Expect = 5e-08
Identities = 33/132 (25%), Positives = 62/132 (46%), Gaps = 1/132 (0%)
Frame = +3
Query: 73 SINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGG 132
++ S++FS DG LA+ D +R +D + + + +LC AWS DGKY+++G
Sbjct: 501 AVLSVAFSPDGRQLASGSGDTTVRFWDLGTQTPMYTCTGHKNWVLCIAWSPDGKYLVSGS 680
Query: 133 EDDLVQVWSMEDRKVVAWG-EGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQ 191
+ W + K + GH W++G++ W + N RF S +D
Sbjct: 681 MSGELICWDPQTGKQLGNALTGHKKWITGIS----WEPVHLN----APCRRFVSASKDGD 836
Query: 192 LLLWELEMDEIV 203
+W++ + + +
Sbjct: 837 ARIWDVSLKKCI 872
>TC78188 similar to PIR|C84870|C84870 probable splicing factor [imported] -
Arabidopsis thaliana, partial (95%)
Length = 1436
Score = 52.8 bits (125), Expect = 2e-07
Identities = 42/135 (31%), Positives = 59/135 (43%), Gaps = 8/135 (5%)
Frame = +1
Query: 71 QGSINSISFSADGAYLATVGRDGYLRVFD---YTKEH-----LVCGGKSYYGGLLCCAWS 122
Q I S+ S DG+YL T G D L ++D Y ++ L ++ LL C WS
Sbjct: 820 QDMITSMQLSPDGSYLLTNGMDCKLCIWDMRPYAPQNRCVKILEGHQHNFEKNLLKCGWS 999
Query: 123 MDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYR 182
DG + G D +V +W R+++ GHN V+ F PN E I
Sbjct: 1000PDGSKVTAGSSDRMVYIWDTTSRRILYKLPGHNGSVNECVF-----HPN-----EPIV-- 1143
Query: 183 FGSVGQDTQLLLWEL 197
GS D Q+ L E+
Sbjct: 1144-GSCSSDKQIYLGEI 1185
Score = 35.0 bits (79), Expect = 0.039
Identities = 23/85 (27%), Positives = 38/85 (44%), Gaps = 4/85 (4%)
Frame = +1
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGE 133
I ++SFS + T G D ++++D K + + + + S DG Y+LT G
Sbjct: 703 ITAVSFSDASDKIYTGGIDNDVKIWDLRKGEVTMTLQGHQDMITSMQLSPDGSYLLTNGM 882
Query: 134 DDLVQVWSME----DRKVVAWGEGH 154
D + +W M + V EGH
Sbjct: 883 DCKLCIWDMRPYAPQNRCVKILEGH 957
Score = 28.5 bits (62), Expect = 3.7
Identities = 17/70 (24%), Positives = 31/70 (44%)
Frame = +1
Query: 128 ILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVG 187
I TGG D+ V++W + +V +GH ++ + S D +T G
Sbjct: 739 IYTGGIDNDVKIWDLRKGEVTMTLQGHQDMITSMQL--------SPDGSYLLTN-----G 879
Query: 188 QDTQLLLWEL 197
D +L +W++
Sbjct: 880 MDCKLCIWDM 909
Score = 28.1 bits (61), Expect = 4.8
Identities = 28/145 (19%), Positives = 55/145 (37%), Gaps = 8/145 (5%)
Frame = +1
Query: 71 QGSINSISFSADGAYLATVGRDGYLRVFDYTK--------EHLVCGGKSYYGGLLCCAWS 122
+ ++ + +++DG + + D LR++D EHL SY CC
Sbjct: 442 KNAVLDLHWTSDGTQIISASPDKTLRLWDTETGKQIKKMVEHL-----SYVNS--CCPTR 600
Query: 123 MDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYR 182
+++G +D ++W M R + D Y + S + Y
Sbjct: 601 RGPPLVVSGSDDGTAKLWDMRQRGSIQ-----------TFPDKYQITAVSFSDASDKIY- 744
Query: 183 FGSVGQDTQLLLWELEMDEIVVPLR 207
+ G D + +W+L E+ + L+
Sbjct: 745 --TGGIDNDVKIWDLRKGEVTMTLQ 813
>TC79260 weakly similar to GP|9759350|dbj|BAB10005.1 WD-repeat protein-like
{Arabidopsis thaliana}, partial (50%)
Length = 1227
Score = 52.0 bits (123), Expect = 3e-07
Identities = 26/80 (32%), Positives = 46/80 (57%)
Frame = +2
Query: 71 QGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILT 130
Q S++S+S+S +G L T G + +R +D + + + GL+ CAW GKYIL+
Sbjct: 680 QKSVSSVSWSPNGQELLTCGVEEAVRRWDVSTGKCLQVYEKNGSGLISCAWFPCGKYILS 859
Query: 131 GGEDDLVQVWSMEDRKVVAW 150
G D + +W ++ ++V +W
Sbjct: 860 GLSDKSICMWELDGKEVESW 919
>TC88481 similar to PIR|T00806|T02445 probable U4/U6 small nuclear
ribonucleoprotein [imported] - Arabidopsis thaliana,
partial (60%)
Length = 1836
Score = 50.4 bits (119), Expect = 9e-07
Identities = 25/92 (27%), Positives = 48/92 (52%)
Frame = +1
Query: 73 SINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGG 132
S+ + F DG+ A+ G D RV+D V + + +L ++S +G ++ TGG
Sbjct: 1333 SVYGLDFHHDGSLAASCGLDALARVWDLRTGRSVLALEGHVKSILGISFSPNGYHLATGG 1512
Query: 133 EDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFD 164
ED+ ++W + +K + H++ +S V F+
Sbjct: 1513 EDNTCRIWDLRKKKSLYTIPAHSNLISQVKFE 1608
Score = 40.8 bits (94), Expect = 7e-04
Identities = 27/90 (30%), Positives = 45/90 (50%), Gaps = 1/90 (1%)
Frame = +1
Query: 73 SINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWS-MDGKYILTG 131
SI ISFS +G +LAT G D R++D K+ + ++ + + +G +++T
Sbjct: 1459 SILGISFSPNGYHLATGGEDNTCRIWDLRKKKSLYTIPAHSNLISQVKFEPQEGYFLVTA 1638
Query: 132 GEDDLVQVWSMEDRKVVAWGEGHNSWVSGV 161
D +VWS D K V GH + V+ +
Sbjct: 1639 SYDMTAKVWSSRDFKPVKTLSGHEAKVTSL 1728
Score = 32.3 bits (72), Expect = 0.26
Identities = 41/182 (22%), Positives = 69/182 (37%), Gaps = 3/182 (1%)
Frame = +1
Query: 110 KSYYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSS 169
K + L A+ GKY+ T D ++W +E + + EGH+ V G+ F S
Sbjct: 1192 KGHLERLARIAFHPSGKYLGTASYDKTWRLWDVETEEELLLQEGHSRSVYGLDFHHDGSL 1371
Query: 170 PNSNDNGETITYRFGSVGQDTQLLLWELEMDEIVVPLRRGPPGGSPTFGSPTFSAGS--- 226
S G D +W+L V+ L G + +FS
Sbjct: 1372 A-------------ASCGLDALARVWDLRTGRSVLALE----GHVKSILGISFSPNGYHL 1500
Query: 227 QSSHWDNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKI 286
+ DN + L+ S+ +P S L++ +V EP ++TA + K+
Sbjct: 1501 ATGGEDNTCRIWDLRKKKSLYTIPAHSNLIS-QVKFEPQEGYF-----LVTASYDMTAKV 1662
Query: 287 WT 288
W+
Sbjct: 1663 WS 1668
Score = 31.2 bits (69), Expect = 0.57
Identities = 15/45 (33%), Positives = 25/45 (55%)
Frame = +1
Query: 119 CAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAF 163
C++S DGK + T ++WSM + K V+ +GH + VA+
Sbjct: 967 CSFSRDGKGLATCSFTGATKLWSMPNVKKVSTLKGHTQRATDVAY 1101
>TC84321 GP|22138476|gb|AAM93460.1 hypothetical protein {Oryza sativa
(japonica cultivar-group)}, partial (4%)
Length = 748
Score = 45.4 bits (106), Expect = 3e-05
Identities = 26/77 (33%), Positives = 41/77 (52%), Gaps = 8/77 (10%)
Frame = +2
Query: 73 SINSISFSADGAYLATVGRDGYLRVFDYTKE--------HLVCGGKSYYGGLLCCAWSMD 124
SI+S+S S DG L + G D LR + K H + G+ G+ WS D
Sbjct: 482 SISSLSLSPDGRELISAGHDASLRFWSLEKRSCTQEITSHRIMRGE----GVCSVVWSQD 649
Query: 125 GKYILTGGEDDLVQVWS 141
GK++++GG D +V+V++
Sbjct: 650 GKWVVSGGGDGVVKVFA 700
Score = 28.9 bits (63), Expect = 2.8
Identities = 13/54 (24%), Positives = 26/54 (48%)
Frame = +2
Query: 92 DGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDR 145
D ++R FD ++ + + S DG+ +++ G D ++ WS+E R
Sbjct: 413 DQFVRFFDANSGQCTYNMLAHPASISSLSLSPDGRELISAGHDASLRFWSLEKR 574
>TC78591 similar to GP|9759350|dbj|BAB10005.1 WD-repeat protein-like
{Arabidopsis thaliana}, partial (17%)
Length = 916
Score = 45.1 bits (105), Expect = 4e-05
Identities = 23/80 (28%), Positives = 43/80 (53%)
Frame = +2
Query: 71 QGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILT 130
Q I+S+S+S + L T G + +R +D + + + GL+ C W GK+IL+
Sbjct: 596 QKPISSVSWSPNDEELLTCGVEESIRRWDVSTGKCLQIYEKAGAGLVSCTWFPSGKHILS 775
Query: 131 GGEDDLVQVWSMEDRKVVAW 150
G D + +W ++ ++V +W
Sbjct: 776 GLSDKSICMWELDGKEVESW 835
>TC79322 similar to GP|13605815|gb|AAK32893.1 At2g32700/F24L7.16
{Arabidopsis thaliana}, partial (59%)
Length = 1824
Score = 42.0 bits (97), Expect = 3e-04
Identities = 25/83 (30%), Positives = 39/83 (46%)
Frame = +2
Query: 114 GGLLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSN 173
G ++CC +S DGK + + G D V +W+ME K + E H ++ V F PNS
Sbjct: 803 GKVVCCHFSSDGKLLASAGHDKKVVLWNMETLKTQSTPEEHTVIITDVRF-----RPNST 967
Query: 174 DNGETITYRFGSVGQDTQLLLWE 196
+ + DT + LW+
Sbjct: 968 --------QLATSSFDTTVRLWD 1012
>TC78874 similar to PIR|T49098|T49098 hypothetical protein F4F15.300 -
Arabidopsis thaliana, partial (74%)
Length = 1630
Score = 41.2 bits (95), Expect = 5e-04
Identities = 17/68 (25%), Positives = 36/68 (52%)
Frame = +2
Query: 77 ISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDL 136
I+FS DG+ A G DG+LR+ ++ ++ + + +S+D +++ + D
Sbjct: 536 ITFSVDGSKFAAGGSDGHLRIMEWPSMRIILDEPRAHKSVRDMDFSLDSEFLASTSTDGS 715
Query: 137 VQVWSMED 144
++W +ED
Sbjct: 716 ARIWKVED 739
>TC87616 similar to PIR|A96839|A96839 F23A5.2(form2) [imported] -
Arabidopsis thaliana, partial (96%)
Length = 1471
Score = 40.8 bits (94), Expect = 7e-04
Identities = 26/95 (27%), Positives = 43/95 (44%), Gaps = 8/95 (8%)
Frame = +3
Query: 56 ARYSKSNPIARWHICQ---GSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGK-- 110
A + +NP + + Q SI+S+SFS +L D +R ++ K V
Sbjct: 144 AANTNTNPNKSYEVSQPPTDSISSLSFSPKANFLVATSWDNQVRCWEIAKNGTVVTSTPK 323
Query: 111 ---SYYGGLLCCAWSMDGKYILTGGEDDLVQVWSM 142
S+ +LC AW DG + +GG D ++W +
Sbjct: 324 ASISHDQPVLCSAWKDDGTTVFSGGCDKQAKMWPL 428
>AW691821 similar to GP|10177570|db contains similarity to regulatory protein
Nedd1~gene_id:K18J17.16 {Arabidopsis thaliana}, partial
(19%)
Length = 648
Score = 39.3 bits (90), Expect = 0.002
Identities = 11/39 (28%), Positives = 27/39 (69%)
Frame = +2
Query: 126 KYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFD 164
+Y+ +GG +V++W ++ ++ + W +GH + V+GV ++
Sbjct: 446 RYVCSGGSGQVVKIWDLQKKRCIKWLKGHTNTVTGVMYN 562
>TC77248 similar to GP|13489182|gb|AAK27816.1 putative WD-repeat containing
protein {Oryza sativa}, partial (92%)
Length = 2005
Score = 38.5 bits (88), Expect = 0.004
Identities = 32/143 (22%), Positives = 62/143 (42%), Gaps = 4/143 (2%)
Frame = +1
Query: 36 GAGESSFPILKDQT--LFSVAHARYSKSNPIARWHICQGSINSISFSADGAYLATVGRDG 93
G GES D+T L+ + + I + H + + +++ A Y T DG
Sbjct: 916 GQGESIITGSADKTVRLWQGSDDGHYNCKQILKDHSAE--VEAVTVHATNNYFVTASLDG 1089
Query: 94 YLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEG 153
++ + + G A+ DG + TG D LV++W ++ + VA +G
Sbjct: 1090 SWCFYELSSGTCLTQVSDSSEGYTSAAFHPDGLILGTGTTDSLVKIWDVKSQANVAKFDG 1269
Query: 154 HNSWVSGVAF--DSYWSSPNSND 174
H V+ ++F + Y+ + ++D
Sbjct: 1270 HVGHVTAISFSENGYYLATAAHD 1338
Score = 30.0 bits (66), Expect = 1.3
Identities = 21/65 (32%), Positives = 30/65 (45%), Gaps = 2/65 (3%)
Frame = +1
Query: 72 GSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMD--GKYIL 129
G + +ISFS +G YLAT DG ++++D K Y + D G Y+
Sbjct: 1276 GHVTAISFSENGYYLATAAHDG-VKLWDLRKLKNFRDYGPYDSATPTNSVEFDHSGSYLA 1452
Query: 130 TGGED 134
GG D
Sbjct: 1453 VGGSD 1467
>TC87605 homologue to PIR|G96563|G96563 probable coatomer complex subunit
33791-27676 [imported] - Arabidopsis thaliana, partial
(23%)
Length = 765
Score = 38.1 bits (87), Expect = 0.005
Identities = 31/141 (21%), Positives = 56/141 (38%), Gaps = 1/141 (0%)
Frame = +3
Query: 58 YSKSNPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLL 117
Y + + + + S F A ++ D ++RV++Y V +++ +
Sbjct: 255 YQSQTMAKSFEVTELPVRSAKFVARKQWVVAGADDMFIRVYNYNTMDKVKVFEAHTDYIR 434
Query: 118 CCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAW-GEGHNSWVSGVAFDSYWSSPNSNDNG 176
C A Y+L+ +D L+++W E + EGH+ +V V F N D
Sbjct: 435 CVAVHPTLPYVLSSSDDMLIKLWDWEKGWICTQIFEGHSHYVMQVTF-------NPKD-- 587
Query: 177 ETITYRFGSVGQDTQLLLWEL 197
T F S D + +W L
Sbjct: 588 ---TNTFASASLDRTIKIWNL 641
>TC85659 SP|O24076|GBLP_MEDSA Guanine nucleotide-binding protein beta
subunit-like protein. [Alfalfa] {Medicago sativa},
complete
Length = 1261
Score = 37.7 bits (86), Expect = 0.006
Identities = 19/81 (23%), Positives = 40/81 (48%), Gaps = 4/81 (4%)
Frame = +2
Query: 72 GSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYY----GGLLCCAWSMDGKY 127
G +N+++ S DG+ A+ G+DG + ++D + GK Y G ++ +Y
Sbjct: 641 GYVNTVAVSPDGSLCASGGKDGVILLWDLAE------GKRLYSLDAGSIIHALCFSPNRY 802
Query: 128 ILTGGEDDLVQVWSMEDRKVV 148
L + +++W +E + +V
Sbjct: 803 WLCAATESSIKIWDLESKSIV 865
Score = 34.3 bits (77), Expect = 0.067
Identities = 30/126 (23%), Positives = 60/126 (46%), Gaps = 4/126 (3%)
Frame = +2
Query: 76 SISFSADGAYLATVGRDGYLRVFDYTKE--HLVCGGKSYYGGLLCCAWSMDGKY--ILTG 131
S++FS D + + RD +++++ E + + G ++ + C +S I++
Sbjct: 389 SVAFSIDNRQIVSASRDRTIKLWNTLGECKYTIQDGDAHSDWVSCVRFSPSTLQPTIVSA 568
Query: 132 GEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQ 191
D V+VW++ + K+ GH+ +V+ VA SP+ + S G+D
Sbjct: 569 SWDRTVKVWNLTNCKLRNTLAGHSGYVNTVAV-----SPDGS--------LCASGGKDGV 709
Query: 192 LLLWEL 197
+LLW+L
Sbjct: 710 ILLWDL 727
Score = 29.6 bits (65), Expect = 1.7
Identities = 38/166 (22%), Positives = 61/166 (35%), Gaps = 11/166 (6%)
Frame = +2
Query: 86 LATVGRDGYLRVFDYTKEHLVCG-------GKSYYGGLLCCAWSMDGKYILTGGEDDLVQ 138
+ T RD + ++ TKE G G S++ + S DG++ L+G D ++
Sbjct: 152 IVTASRDKSIILWHLTKEDKTYGVPRRRLTGHSHF--VQDVVLSSDGQFALSGSWDGELR 325
Query: 139 VWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQLLLWELE 198
+W + GH V VAF S DN + + S +D + LW
Sbjct: 326 LWDLNAGTSARRFVGHTKDVLSVAF--------SIDNRQIV-----SASRDRTIKLWN-T 463
Query: 199 MDEIVVPLRRGPPGGS----PTFGSPTFSAGSQSSHWDNAVPLGTL 240
+ E ++ G F T S+ WD V + L
Sbjct: 464 LGECKYTIQDGDAHSDWVSCVRFSPSTLQPTIVSASWDRTVKVWNL 601
>TC79131 similar to GP|14031063|gb|AAK52092.1 WD-40 repeat protein
{Lycopersicon esculentum}, partial (71%)
Length = 1224
Score = 37.7 bits (86), Expect = 0.006
Identities = 23/79 (29%), Positives = 38/79 (47%), Gaps = 8/79 (10%)
Frame = +2
Query: 79 FSADGAYLATVGRDGYLRVFDY----TKEHLVCGGKSYY----GGLLCCAWSMDGKYILT 130
FS DG YL + DG++ V+DY K+ L + + +LC +S D + I +
Sbjct: 734 FSPDGQYLVSCSIDGFIEVWDYISGKLKKDLQYQAEETFMMHDEPVLCVDFSRDSEMIAS 913
Query: 131 GGEDDLVQVWSMEDRKVVA 149
G D ++VW + + A
Sbjct: 914 GSTDGKIKVWRIRTAQCXA 970
>TC93203
Length = 681
Score = 37.4 bits (85), Expect = 0.008
Identities = 33/142 (23%), Positives = 61/142 (42%)
Frame = +2
Query: 9 CTCISWVPGGDGAFVVAHADGNLYNKDGAGESSFPILKDQTLFSVAHARYSKSNPIARWH 68
CTC + FV A A G G+G + F L +L +V + + K+ +
Sbjct: 20 CTCDIKCSPVEPIFVSAAASG------GSGSNYFDNLGFASL-TVWNMKTWKAMTVLPLG 178
Query: 69 ICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYI 128
+I S+ F+ +G LA DG + +FD + + G ++ + + D I
Sbjct: 179 EDPPAITSLCFNHNGKILAASAVDGMIHMFDMSAGLQITGWPAHDSSISSILFGPDETSI 358
Query: 129 LTGGEDDLVQVWSMEDRKVVAW 150
+ G D + WS++++ + W
Sbjct: 359 FSLGSDGKIFEWSLQNQGQILW 424
>TC87005 similar to GP|9294061|dbj|BAB02018.1 WD40-repeat protein
{Arabidopsis thaliana}, partial (74%)
Length = 1480
Score = 36.6 bits (83), Expect = 0.014
Identities = 21/77 (27%), Positives = 36/77 (46%), Gaps = 3/77 (3%)
Frame = +3
Query: 74 INSISFSADGAYLATVGRDGYLRVFD-YTKEHL--VCGGKSYYGGLLCCAWSMDGKYILT 130
+N + +S DG+ T D +FD T E + + + G + +WS DGK +LT
Sbjct: 609 VNCVRYSPDGSKFVTGSSDKKGIIFDGKTGEKIGELSSEGGHTGSIYAVSWSPDGKQVLT 788
Query: 131 GGEDDLVQVWSMEDRKV 147
D +VW + + +
Sbjct: 789 VSADKSAKVWDISEDNI 839
>TC78974 similar to GP|22137238|gb|AAM91464.1 AT5g14050/MUA22_5 {Arabidopsis
thaliana}, partial (40%)
Length = 774
Score = 36.2 bits (82), Expect = 0.018
Identities = 22/79 (27%), Positives = 36/79 (44%)
Frame = +3
Query: 73 SINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGG 132
SI S D +A VG +GY+ + TK + G G + A++ DG+ +L+ G
Sbjct: 33 SIEVFDVSPDSQLIAFVGNEGYILLVS-TKSKQLVGTLKMNGTVRSLAFTEDGQKLLSAG 209
Query: 133 EDDLVQVWSMEDRKVVAWG 151
D + W + R + G
Sbjct: 210 GDGHIYHWDLRTRTCIHKG 266
>TC88235 similar to GP|12656803|gb|AAK00964.1 unknown protein {Oryza
sativa}, partial (91%)
Length = 1080
Score = 35.8 bits (81), Expect = 0.023
Identities = 19/60 (31%), Positives = 32/60 (52%), Gaps = 3/60 (5%)
Frame = +2
Query: 85 YLATVGRDGYLRVF---DYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQVWS 141
YLAT D ++++ D+T E + G + L C +S+DG Y++T D ++WS
Sbjct: 896 YLATASSDHTVKIWNVDDFTLEKTLIG---HQRRLWDCVFSVDGAYLITASSDSTARLWS 1066
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.133 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,001,569
Number of Sequences: 36976
Number of extensions: 187376
Number of successful extensions: 990
Number of sequences better than 10.0: 91
Number of HSP's better than 10.0 without gapping: 921
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 981
length of query: 325
length of database: 9,014,727
effective HSP length: 96
effective length of query: 229
effective length of database: 5,465,031
effective search space: 1251492099
effective search space used: 1251492099
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC148995.8


