

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148819.4 - phase: 0
(1009 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
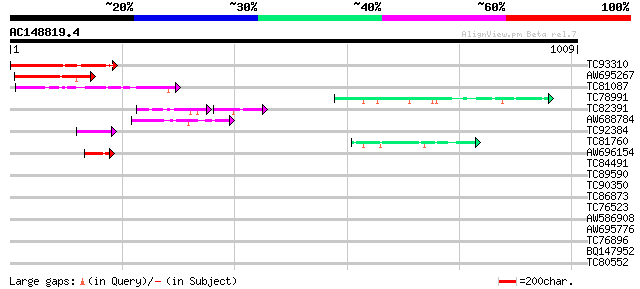
Score E
Sequences producing significant alignments: (bits) Value
TC93310 homologue to GP|9759268|dbj|BAB09589.1 101 kDa heat shoc... 212 5e-55
AW695267 similar to GP|15450731|gb At2g40130/T7M7.2 {Arabidopsis... 118 1e-26
TC81087 homologue to GP|530207|gb|AAA66338.1|| heat shock protei... 84 3e-16
TC78991 similar to GP|9759268|dbj|BAB09589.1 101 kDa heat shock ... 77 3e-14
TC82391 similar to PIR|H85354|H85354 hypothetical protein AT4g30... 54 4e-14
AW688784 weakly similar to GP|14532596|gb unknown protein {Arabi... 59 7e-09
TC92384 similar to PIR|B86207|B86207 hypothetical protein [impor... 57 3e-08
TC81760 weakly similar to PIR|T08450|T08450 hypothetical protein... 52 1e-06
AW696154 similar to GP|14532596|gb unknown protein {Arabidopsis ... 46 6e-05
TC84491 homologue to GP|1685115|gb|AAB36702.1| putative transcri... 34 0.32
TC89590 similar to GP|17473836|gb|AAL38344.1 unknown protein {Ar... 33 0.42
TC90350 weakly similar to GP|22830929|dbj|BAC15794. hypothetical... 33 0.72
TC86873 homologue to GP|530207|gb|AAA66338.1|| heat shock protei... 32 0.94
TC76523 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - ... 32 1.2
AW586908 weakly similar to GP|2104814|emb| pleiotropic effects o... 32 1.6
AW695776 weakly similar to GP|9759268|dbj| 101 kDa heat shock pr... 31 2.1
TC76896 homologue to SP|P35100|CLPA_PEA ATP-dependent clp protea... 31 2.1
BQ147952 GP|22136744|gb putative putatative protein {Arabidopsis... 31 2.1
TC80552 similar to PIR|G86472|G86472 probable hyoscyamine 6-diox... 30 3.6
>TC93310 homologue to GP|9759268|dbj|BAB09589.1 101 kDa heat shock protein;
HSP101-like protein {Arabidopsis thaliana}, partial
(15%)
Length = 817
Score = 212 bits (540), Expect = 5e-55
Identities = 119/192 (61%), Positives = 142/192 (72%)
Frame = +2
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQ 60
MR+G T+Q+TLT EAAS+L HS+ A RR H Q TPLHVAATLL+ PS LR AC+KS
Sbjct: 146 MRAGLSTIQQTLTPEAASVLNHSIAEAGRRNHGQTTPLHVAATLLASPSGYLRQACIKSH 325
Query: 61 QQNNHPLQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRRGCI 120
++HPLQCRALELCF+VAL RLPT+ + P +SNAL+AALKRAQAHQRRG
Sbjct: 326 PNSSHPLQCRALELCFSVALERLPTSQNASSTSAME-PPISNALMAALKRAQAHQRRGYP 502
Query: 121 EQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLINSSS 180
E QQQQPLL+VKVEL+QLI+SILDDPSVSRVMREA FSSP+ K +E S +NS
Sbjct: 503 E----QQQQPLLAVKVELEQLIISILDDPSVSRVMREASFSSPAXKATIEQS---LNS-- 655
Query: 181 VFHSSPSPLSHN 192
+PSP++ N
Sbjct: 656 ---VAPSPVTVN 682
>AW695267 similar to GP|15450731|gb At2g40130/T7M7.2 {Arabidopsis thaliana},
partial (29%)
Length = 625
Score = 118 bits (295), Expect = 1e-26
Identities = 72/155 (46%), Positives = 102/155 (65%), Gaps = 10/155 (6%)
Frame = +2
Query: 9 QETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSS-LRNACLKSQQQNNHP- 66
++ LT EA L ++ +A+RRGHAQ T LH + LLSLPSSS LR+AC +S+ P
Sbjct: 149 RQCLTPEAIQALNDAVAVAKRRGHAQTTSLHAISALLSLPSSSILRDACSRSRNSAYSPR 328
Query: 67 LQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRR--------G 118
LQ +AL+LC +V+L+R P++ + + H P +SN+L+AA+KR+QA+QRR
Sbjct: 329 LQFKALDLCLSVSLDRSPSSHNN--VSSDHEPPVSNSLMAAIKRSQANQRRHPDNFHFYH 502
Query: 119 CIEQNQQQQQQPLLSVKVELDQLILSILDDPSVSR 153
+Q Q QQ + SVKVEL L+LS+LDDP VS+
Sbjct: 503 QQQQLQSQQTFSVSSVKVELQHLVLSVLDDPVVSK 607
>TC81087 homologue to GP|530207|gb|AAA66338.1|| heat shock protein {Glycine
max}, partial (41%)
Length = 1318
Score = 84.0 bits (206), Expect = 3e-16
Identities = 88/308 (28%), Positives = 133/308 (42%), Gaps = 14/308 (4%)
Frame = +2
Query: 10 ETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHPLQC 69
E T + L + LA GHAQITPLH+A+ L+S P+ A + +
Sbjct: 191 EKFTHKTNEALAGAHELAMTSGHAQITPLHLASILVSDPNGIFFQAISNVAGEES----A 358
Query: 70 RALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQR-RGCIEQNQQQQQ 128
RA+E AL +LP+ + P VP S ALI A++RAQA Q+ RG
Sbjct: 359 RAVERVLKQALKKLPSQSP----PPDEVPG-STALIKAIRRAQAAQKSRG---------- 493
Query: 129 QPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSPSP 188
+ +DQLIL IL+D + + +EAG + VK +E L S S
Sbjct: 494 ----DTHLAVDQLILGILEDSQIGDLFKEAGVAVSRVKTEVEK---LRGKDGKKVESASG 652
Query: 189 LSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKKKKNTVIVG 248
++ L +YG ++ + K + V+ E+I V +L R+ K N V++G
Sbjct: 653 DTNFQALKTYG-RDLVEQAGKLDPVI---------GRDEEIRRVVRILSRRTKNNPVLIG 802
Query: 249 DTVSLTEGLVSEIMKRFERGEVPDEMKTTHFV-----------KFHGLSSVSLKYMKK-- 295
+ +V + +R RG+VP + + K+ G LK + K
Sbjct: 803 EPGVGKTAVVEGLAQRIVRGDVPSNLADVRLIALDMGALVAGAKYRGEFEERLKAVLKEV 982
Query: 296 EEVEMNVI 303
EE E VI
Sbjct: 983 EEAEGKVI 1006
>TC78991 similar to GP|9759268|dbj|BAB09589.1 101 kDa heat shock protein;
HSP101-like protein {Arabidopsis thaliana}, partial
(13%)
Length = 1370
Score = 77.0 bits (188), Expect = 3e-14
Identities = 97/438 (22%), Positives = 175/438 (39%), Gaps = 49/438 (11%)
Frame = +2
Query: 579 QKNQVTATLELDSLKGMEGTKEVKTTLALGN--STFSVSDQKRMENLTLQR-------DH 629
+K+ VT L L K + E + + S+ S Q + + L ++
Sbjct: 155 RKSTVTTELVLGQTKPSDTIPEESHRERINDFLSSLSSESQDKFDELHSKKLFDTDSFKR 334
Query: 630 IYKVLQENIPWHCETVSSIAEALV-----DSKSSKECATWLFLQGNDSVGKKRLALAIAE 684
+ K L E + W + S+IA A+ + K + TWL G D +GKKR+A A++E
Sbjct: 335 LLKTLTEKVWWQQDAASAIATAVTQCKLGNGKRRSKGDTWLMFTGPDRIGKKRMAAALSE 514
Query: 685 SVFGSVEMFSHVDMMKRENSETPFS-------EKVVGPLKNNEKFVVLVENADFGDTLIR 737
V GS + + + + +++V ++ N V+++E+ D +TL+R
Sbjct: 515 LVSGSNPIVISLAQRRGDGDSNAHQFRGKTVLDRIVETIRRNPHSVIMLEDIDEANTLLR 694
Query: 738 KMLADEFEIAKFG-------TLGQKIFI------------LSNGGSMVSEDQKKDSVMKL 778
+ E +F +LG +FI LSNG + E + +
Sbjct: 695 GNIKRAMEQGRFPDSHGREISLGNVMFILTSNWLPEDLSYLSNGAPLDDEKLENLASGGW 874
Query: 779 VLKISETEKKPTFELSPSSSSSSKSPCLGNKRSAELDLFSKIKIPRIEENEGNKKREFSF 838
L++S T+K + +R + L N++R
Sbjct: 875 QLRLSVTKK------------------VSKRRPSWL---------------SNEERSLKP 955
Query: 839 SRQSSFNNTLDLNMKADEEDNEDYDEGENSPISSDLT------RETLGEHLISNESLDSI 892
++ + + DLN AD E+ D + S SSD T G E LDS+
Sbjct: 956 RKELNLGLSFDLNEAADVEE----DRADGSHNSSDFTVDHEENNHNGGSPSKPRELLDSV 1123
Query: 893 ENLFEFNQSPAKNKEMTQMFMSRVKESFEEVLGN-VKFSVQDKVIEEI--GVGCGSFTNN 949
++ F P + Q F + + + F V+GN + VQ++ +++I GV G T
Sbjct: 1124DDAIVF--KPLNFDLIRQNFSASIAKRFSAVVGNGISIEVQEEALDKITSGVWLGQTT-- 1291
Query: 950 MFEKWLKGIFQTSLERVN 967
++W++ + S ++N
Sbjct: 1292-IDEWMEKVLVPSFHQLN 1342
>TC82391 similar to PIR|H85354|H85354 hypothetical protein AT4g30350
[imported] - Arabidopsis thaliana, partial (13%)
Length = 1065
Score = 53.5 bits (127), Expect(2) = 4e-14
Identities = 34/112 (30%), Positives = 58/112 (51%), Gaps = 16/112 (14%)
Frame = +1
Query: 363 GKIWLVATASYQSYMRCQMRVPTFENQWCLQAVPV--------------PSGGLGLSLHS 408
GK+WL+ TA+ ++Y+RCQ+ P+ EN W LQAVP+ +G LG +L S
Sbjct: 508 GKLWLLGTATCETYLRCQVYHPSMENDWDLQAVPITTRSPLPGMFPRLGTNGILGTTLES 687
Query: 409 SSVHDSKMSISQNP-SPMLESKFFSNKEEHEKLNCCEECVSNYEKE-AQLFK 458
S +++ P +P+ + + CC +C+ + E+E A +F+
Sbjct: 688 LS---PLKTLTPTPITPLTRASENVDPAAAAAPTCCPQCMRSCEQEIADMFE 834
Score = 43.5 bits (101), Expect(2) = 4e-14
Identities = 39/151 (25%), Positives = 74/151 (48%), Gaps = 18/151 (11%)
Frame = +3
Query: 227 EDINLVFDVLLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHGLS 286
+++ V ++L+R KK+N V+VG+ S E + E++K+ E E+ + + F H +
Sbjct: 87 DEVKRVVEILMRTKKRNPVLVGE--SEPEAAIREVLKKIENKELGEGV----FSNAHAI- 245
Query: 287 SVSLKYMKKE-EVEMNVIRVLKRKVSDYVALGVG-------AIFYVGDLKWIVDD----N 334
Y++KE + I V +++ D + +G +GDLKW+V+
Sbjct: 246 -----YLEKELPSDRGQIPVRIKELGDLIESRLGNSGSCGGVFINLGDLKWLVEQPVGFG 410
Query: 335 DGSLNEKEVVD---YVVEEIGKL---FGEEG 359
G++ + + + V E+G+L FGE G
Sbjct: 411 LGNMQQPALAEAGRAAVAEMGRLVAKFGEGG 503
>AW688784 weakly similar to GP|14532596|gb unknown protein {Arabidopsis
thaliana}, partial (2%)
Length = 625
Score = 59.3 bits (142), Expect = 7e-09
Identities = 52/189 (27%), Positives = 85/189 (44%), Gaps = 6/189 (3%)
Frame = +3
Query: 217 PFLKSSESNKEDINLVFDVLLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKT 276
PFL E+ + ++L+R K KN +++G + +E +++ G +P E+
Sbjct: 6 PFLSGVGDVDENFRRIGEILVRSKGKNPLLLGACGNDALRSFTEAVEKRREGVLPLELD- 182
Query: 277 THFVKFHGLSSVSLKYMKKEEVEMNVIRVLKRKVSDYVAL-----GVGAIFYVGDLKWIV 331
GL + + +E+E V+ K+ A+ G G I G+LK V
Sbjct: 183 -------GLRVICIG----KELESGDCEVVSLKLKQIAAIVEECVGPGVIVSFGELKSFV 329
Query: 332 DDNDGSLNEKEVVDYVVEEIGKLFGEEGNKNGKIWLVATA-SYQSYMRCQMRVPTFENQW 390
+D+ G VEE+GKL +K WL A SY+SY++ R P+ E W
Sbjct: 330 NDDGG----------FVEELGKLLKIHYDK---FWLAGAADSYESYLKFLGRFPSVEKDW 470
Query: 391 CLQAVPVPS 399
LQ +P+ S
Sbjct: 471 DLQILPITS 497
>TC92384 similar to PIR|B86207|B86207 hypothetical protein [imported] -
Arabidopsis thaliana, partial (8%)
Length = 703
Score = 57.4 bits (137), Expect = 3e-08
Identities = 35/70 (50%), Positives = 39/70 (55%)
Frame = +1
Query: 120 IEQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLINSS 179
++Q QQ Q Q +KVEL ILSILDDP VSRV EAGF S VK L SS
Sbjct: 124 MQQQQQNQNQTASFLKVELKHFILSILDDPIVSRVFAEAGFRSYDVKFALLQPPPPPPSS 303
Query: 180 SVFHSSPSPL 189
FH S P+
Sbjct: 304 RFFHRSSPPV 333
>TC81760 weakly similar to PIR|T08450|T08450 hypothetical protein F22O6.130
- Arabidopsis thaliana, partial (23%)
Length = 826
Score = 51.6 bits (122), Expect = 1e-06
Identities = 61/255 (23%), Positives = 102/255 (39%), Gaps = 25/255 (9%)
Frame = +3
Query: 609 NSTFSVSDQKRMENLTLQRD-------HIYKVLQENIPWHCETVSSIAEALVDSKS---- 657
NST S SD ME L ++ + L++ +PW + + IA ++ +S
Sbjct: 45 NSTIS-SDLVEMEQLNNFKELNLENMRTLCNALEKKVPWQKDIIPEIASTVLQCRSGLVK 221
Query: 658 ---------SKECATWLFLQGNDSVGKKRLALAIAESVFGSVEMFSHVDMMKRENSETPF 708
+KE TWLF QG D K+++A +A+ VFGS F + + ++
Sbjct: 222 RKGKNNDHDAKE-ETWLFFQGVDLEAKEKIAKELAKLVFGSYNNFISISLSSFSSTRADS 398
Query: 709 SEKVVGPLKNNEKFVVLVENADFGDTLI----RKMLADEFE-IAKFGTLGQKIFILSNGG 763
SE+ +E +E FGD + R L ++ E + F LG K I G
Sbjct: 399 SEESRNKRTRDEASCTYIER--FGDAMSSNPHRVFLVEDIEQVDYFSQLGFKRAI-EKGK 569
Query: 764 SMVSEDQKKDSVMKLVLKISETEKKPTFELSPSSSSSSKSPCLGNKRSAELDLFSKIKIP 823
+ S ++ +++ LS + SS C +RS++ D I +
Sbjct: 570 VLDSNGEEVCFCDAIII------------LSCENFSSRSRVCSPKQRSSQEDKDDDINVA 713
Query: 824 RIEENEGNKKREFSF 838
+EE + F
Sbjct: 714 TLEETSSYVSLDLKF 758
>AW696154 similar to GP|14532596|gb unknown protein {Arabidopsis thaliana},
partial (5%)
Length = 557
Score = 46.2 bits (108), Expect = 6e-05
Identities = 29/53 (54%), Positives = 33/53 (61%)
Frame = +3
Query: 134 VKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSP 186
+KVEL +LSILDDP V+RV EAGF S VK L + SSS F SSP
Sbjct: 123 LKVELKHFVLSILDDPIVNRVFSEAGFRSCDVK--LALLQPPVQSSSRFLSSP 275
>TC84491 homologue to GP|1685115|gb|AAB36702.1| putative transcription
factor {Dictyostelium discoideum}, partial (3%)
Length = 675
Score = 33.9 bits (76), Expect = 0.32
Identities = 25/104 (24%), Positives = 45/104 (43%), Gaps = 8/104 (7%)
Frame = +3
Query: 418 ISQNPSPMLESKF-----FSNKEEHEKLNCCEECVSNYEKEAQLFKPD--QKNLLPSWLQ 470
+SQ P+P L F +EH + C + + N EKE K D +K P ++
Sbjct: 324 LSQRPAPTLFDTHDKDIEFERFKEHREPRCASKQIVNLEKECLKLKNDRCKKEDQPKFVS 503
Query: 471 SHSTEARQKDELTQLNKKWNRLC-QCLHQNKQPQNHWSNNHSSN 513
S + E + + ++ +++W C +P H+ + SN
Sbjct: 504 SENEEDNARAKRSKKHREWQGFWNSCSLTLLKPLQHFKVEYLSN 635
>TC89590 similar to GP|17473836|gb|AAL38344.1 unknown protein {Arabidopsis
thaliana}, partial (30%)
Length = 1754
Score = 33.5 bits (75), Expect = 0.42
Identities = 23/88 (26%), Positives = 38/88 (43%), Gaps = 1/88 (1%)
Frame = +3
Query: 165 VKNNLENSSTLINSSSVFHSSPSPLS-HNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSE 223
+ N LE T + S + + HNH L L KE++ YH LKS+
Sbjct: 138 ISNVLEQLETNVLSPGIGNELADAYDIHNHILD-------LKDMLLKEKMDYHSLLKSAN 296
Query: 224 SNKEDINLVFDVLLRKKKKNTVIVGDTV 251
E N+ D+L + + ++++G V
Sbjct: 297 ETAEPGNMTLDILELNRLRRSLLIGSHV 380
>TC90350 weakly similar to GP|22830929|dbj|BAC15794. hypothetical
protein~similar to Arabidopsis thaliana chromosome 1
T6L1.12, partial (32%)
Length = 984
Score = 32.7 bits (73), Expect = 0.72
Identities = 27/118 (22%), Positives = 56/118 (46%), Gaps = 1/118 (0%)
Frame = +2
Query: 61 QQNNHPLQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRRGCI 120
++NN ++C+ L L + L++ T + L H+ ++ N + ++ ++ G +
Sbjct: 161 KENNIKIRCKVLNLLYT--LSKDMTDELTEYLDESHIFNIVNIVSSSTSDSEKAAAVGIL 334
Query: 121 EQNQQQQQQPLLSVK-VELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLIN 177
++ +K L QL++SIL + S+ SPS N +EN++ +IN
Sbjct: 335 SNLPASDKKVTDILKRASLLQLLISILYSSNASK--------SPSTNNLIENATGVIN 484
>TC86873 homologue to GP|530207|gb|AAA66338.1|| heat shock protein {Glycine
max}, partial (53%)
Length = 1475
Score = 32.3 bits (72), Expect = 0.94
Identities = 20/61 (32%), Positives = 34/61 (54%), Gaps = 5/61 (8%)
Frame = +2
Query: 643 ETVSSIAEALVDSKSS-----KECATWLFLQGNDSVGKKRLALAIAESVFGSVEMFSHVD 697
+ V+++AEA++ S++ + ++LFL G VGK LA A+AE +F +D
Sbjct: 629 QAVNAVAEAVLRSRAGLGRPQQPTGSFLFL-GPTGVGKTELAKALAEQLFDDENQLVRID 805
Query: 698 M 698
M
Sbjct: 806 M 808
>TC76523 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - alfalfa,
partial (69%)
Length = 1615
Score = 32.0 bits (71), Expect = 1.2
Identities = 35/128 (27%), Positives = 56/128 (43%)
Frame = +3
Query: 833 KREFSFSRQSSFNNTLDLNMKADEEDNEDYDEGENSPISSDLTRETLGEHLISNESLDSI 892
K+E S S S ++ +N+ ++D D DE E S D +T + + E +D+
Sbjct: 327 KKEVSESESDSSDDEDKMNV---DKDGSDSDESEEE--SEDEPSKTPQKKIKDVEMVDAG 491
Query: 893 ENLFEFNQSPAKNKEMTQMFMSRVKESFEEVLGNVKFSVQDKVIEEIGVGCGSFTNNMFE 952
++ A N T + S K F +GN+ FSVQ IE+ CG + F
Sbjct: 492 KS-----GKKAPNTPATPIENSGSKTLF---VGNLSFSVQRSDIEKFFQDCGEVVDVRFS 647
Query: 953 KWLKGIFQ 960
+G F+
Sbjct: 648 SDEEGRFK 671
>AW586908 weakly similar to GP|2104814|emb| pleiotropic effects on cellular
differentiation and slug behaviour {Dictyostelium
discoideum}, partial (8%)
Length = 668
Score = 31.6 bits (70), Expect = 1.6
Identities = 25/79 (31%), Positives = 33/79 (41%)
Frame = +2
Query: 53 RNACLKSQQQNNHPLQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQ 112
+ A L +QQ + LQ +L+L L R P PQ++ L K Q
Sbjct: 425 QQAGLLQRQQQQNQLQQSSLQLLQQSMLQRGPQQPQMSQTVPQNISDQQQQLQLLQKLQQ 604
Query: 113 AHQRRGCIEQNQQQQQQPL 131
+Q QQQQQQPL
Sbjct: 605 --------QQQQQQQQQPL 637
>AW695776 weakly similar to GP|9759268|dbj| 101 kDa heat shock protein;
HSP101-like protein {Arabidopsis thaliana}, partial (2%)
Length = 655
Score = 31.2 bits (69), Expect = 2.1
Identities = 18/57 (31%), Positives = 26/57 (45%), Gaps = 8/57 (14%)
Frame = +1
Query: 630 IYKVLQENIPWHCETVSSIAEALVDSKSSK--------ECATWLFLQGNDSVGKKRL 678
+YK+L E + W E + SI + +SS TW G D VGK++L
Sbjct: 466 LYKLLIEKVWWQDEAIYSIINIMTLCRSSDGKRSGSNVRADTWFSFLGPDRVGKRKL 636
>TC76896 homologue to SP|P35100|CLPA_PEA ATP-dependent clp protease
ATP-binding subunit clpA homolog chloroplast precursor.
[Garden pea], partial (35%)
Length = 1348
Score = 31.2 bits (69), Expect = 2.1
Identities = 36/141 (25%), Positives = 54/141 (37%), Gaps = 4/141 (2%)
Frame = +1
Query: 647 SIAEALVDSKSSKECATWLFLQGNDSVGKKRLALAIAESVFGSVEMFSHVDMMKRENSET 706
+I A V K+ G VGK LA +A FGS E +DM E E
Sbjct: 100 AIRRARVGLKNPNRPIASFIFSGPTGVGKSELAKTLASYYFGSEEAMIRLDM--SEFMER 273
Query: 707 PFSEKVVGPLKNNEKFVVLVENADFGDTLIRK----MLADEFEIAKFGTLGQKIFILSNG 762
K++G + +V E + + R+ +L DE E A + IL +G
Sbjct: 274 HTVSKLIG---SPPGYVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDG 444
Query: 763 GSMVSEDQKKDSVMKLVLKIS 783
S+ + D L++ S
Sbjct: 445 RLTDSKGRTVDFKNTLLIMTS 507
>BQ147952 GP|22136744|gb putative putatative protein {Arabidopsis thaliana},
partial (2%)
Length = 597
Score = 31.2 bits (69), Expect = 2.1
Identities = 28/110 (25%), Positives = 52/110 (46%), Gaps = 1/110 (0%)
Frame = +3
Query: 782 ISETEKKPTFELSPSSSSSSKSPCLGNKRSAELDLF-SKIKIPRIEENEGNKKREFSFSR 840
++ KKP + K L ++R A LD S+ K R++E++ KKR++ S
Sbjct: 81 LTNLSKKPK---KSKNKKHKKQKLLHSEREASLDDGESRSKKKRVKESK--KKRDYKESS 245
Query: 841 QSSFNNTLDLNMKADEEDNEDYDEGENSPISSDLTRETLGEHLISNESLD 890
S + ++ K+ + ++D+D N P +R+T E +S + D
Sbjct: 246 SSGSFASEKVHGKSRNKHSDDFDSDRNDP-----SRKTKPERSLSLKDYD 380
>TC80552 similar to PIR|G86472|G86472 probable hyoscyamine 6-dioxygenase
hydroxylase [imported] - Arabidopsis thaliana, partial
(92%)
Length = 1307
Score = 30.4 bits (67), Expect = 3.6
Identities = 17/47 (36%), Positives = 27/47 (57%), Gaps = 5/47 (10%)
Frame = +3
Query: 937 EEIGVGCGSFTNN---MFEKWLKGIFQTSLERV--NGGDKNGIVYTL 978
E++ G+F N M E+W G+F+++L RV NG ++ I Y L
Sbjct: 678 EDVAPLKGAFVVNLGDMLERWSNGVFKSTLHRVLGNGQERYSIAYFL 818
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.129 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,764,639
Number of Sequences: 36976
Number of extensions: 477240
Number of successful extensions: 2770
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 2713
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2756
length of query: 1009
length of database: 9,014,727
effective HSP length: 106
effective length of query: 903
effective length of database: 5,095,271
effective search space: 4601029713
effective search space used: 4601029713
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC148819.4


