

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148819.3 + phase: 0 /pseudo/partial
(880 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
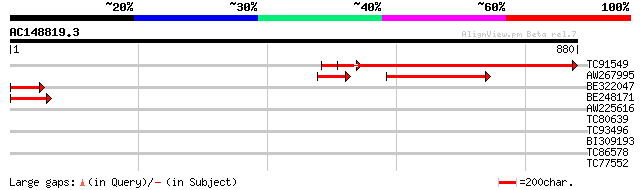
TC91549 similar to GP|17063189|gb|AAL32989.1 unknown protein {Ar... 733 0.0
AW267995 weakly similar to GP|17063189|gb unknown protein {Arabi... 203 3e-52
BE322047 similar to GP|17063189|gb| unknown protein {Arabidopsis... 91 1e-18
BE248171 74 2e-13
AW225616 similar to GP|21740506|em OSJNBa0029H02.1 {Oryza sativa... 31 1.8
TC80639 weakly similar to PIR|G96621|G96621 probable disease res... 30 4.0
TC93496 similar to GP|9757837|dbj|BAB08274.1 contains similarity... 29 6.9
BI309193 similar to GP|1223582|emb| PAC {Arabidopsis thaliana}, ... 29 9.0
TC86578 similar to PIR|C85440|C85440 myb-related protein [import... 29 9.0
TC77552 similar to GP|6984224|gb|AAF34800.1| 60S ribosomal prote... 29 9.0
>TC91549 similar to GP|17063189|gb|AAL32989.1 unknown protein {Arabidopsis
thaliana}, partial (31%)
Length = 1499
Score = 733 bits (1893), Expect = 0.0
Identities = 361/372 (97%), Positives = 361/372 (97%)
Frame = +3
Query: 509 LRQLLPYSESEFNNHHLELLKSVVLPDI*LAKTCFFLYEMIEIWCYGVPRSHIG*LNGFV 568
LRQLLPYSESEFNNHHLELLK L K FFLYEMIEIWCYGVPRSHIG*LNGFV
Sbjct: 75 LRQLLPYSESEFNNHHLELLKIYNLQ-----KLVFFLYEMIEIWCYGVPRSHIG*LNGFV 239
Query: 569 IPP*QPAIKEFVSKF*KISYENSASALEKEVRGFWPDLLITVLCDEWRKCKRAMESSSPP 628
IPP*QPAIKEFVSKF*KISYENSASALEKEVRGFWPDLLITVLCDEWRKCKRAMESSSPP
Sbjct: 240 IPP*QPAIKEFVSKF*KISYENSASALEKEVRGFWPDLLITVLCDEWRKCKRAMESSSPP 419
Query: 629 KEPKCMLFPPRMFFSEEDIPEGSSFTAGERMHELVKVFVLLHQLQIFTHGRALPDQPLIY 688
KEPKCMLFPPRMFFSEEDIPEGSSFTAGERMHELVKVFVLLHQLQIFTHGRALPDQPLIY
Sbjct: 420 KEPKCMLFPPRMFFSEEDIPEGSSFTAGERMHELVKVFVLLHQLQIFTHGRALPDQPLIY 599
Query: 689 HPCDHRTNSRAHTSGLAAVPKPGTEMNLVNAVPCRIAFERGKERHFCFLAISVGTSGWLV 748
HPCDHRTNSRAHTSGLAAVPKPGTEMNLVNAVPCRIAFERGKERHFCFLAISVGTSGWLV
Sbjct: 600 HPCDHRTNSRAHTSGLAAVPKPGTEMNLVNAVPCRIAFERGKERHFCFLAISVGTSGWLV 779
Query: 749 LAEELPLKKPYGVVRVAAPLAGCNPRVDDKHSKWLHLRIRPSALPFLDPVKYNRHGKLKT 808
LAEELPLKKPYGVVRVAAPLAGCNPRVDDKHSKWLHLRIRPSALPFLDPVKYNRHGKLKT
Sbjct: 780 LAEELPLKKPYGVVRVAAPLAGCNPRVDDKHSKWLHLRIRPSALPFLDPVKYNRHGKLKT 959
Query: 809 KAFVDGRWILAFRDEESCKNAFLMIREEINHLCDEVDRRIKPLLKLETVLDISSSSAPVS 868
KAFVDGRWILAFRDEESCKNAFLMIREEINHLCDEVDRRIKPLLKLETVLDISSSSAPVS
Sbjct: 960 KAFVDGRWILAFRDEESCKNAFLMIREEINHLCDEVDRRIKPLLKLETVLDISSSSAPVS 1139
Query: 869 EDSSSHTTPPDS 880
EDSSSHTTPPDS
Sbjct: 1140EDSSSHTTPPDS 1175
Score = 61.6 bits (148), Expect = 1e-09
Identities = 33/61 (54%), Positives = 40/61 (65%)
Frame = +2
Query: 485 LDALVSLFCRSNISAETLWDGGWLLRQLLPYSESEFNNHHLELLKSVVLPDI*LAKTCFF 544
LDALVSLFCRSNISAETLWDGGW + + ++ ++ +I*LAKTCFF
Sbjct: 2 LDALVSLFCRSNISAETLWDGGWXFTPV-----ASLQ*VRIQQPPPGIVENI*LAKTCFF 166
Query: 545 L 545
L
Sbjct: 167 L 169
>AW267995 weakly similar to GP|17063189|gb unknown protein {Arabidopsis
thaliana}, partial (17%)
Length = 644
Score = 203 bits (516), Expect = 3e-52
Identities = 105/164 (64%), Positives = 122/164 (74%), Gaps = 3/164 (1%)
Frame = +2
Query: 585 KISYENSASALEKEVRGFWPDLLITVLCDEWRKCKRAMESSSPPKEPKCMLF--PPRMFF 642
K+SY+N AS L +EV+G W D LI+V+CDEWRKCKRAMESSSPPKEP C+LF P F
Sbjct: 152 KVSYKNCASDLVEEVKGIWSDFLISVICDEWRKCKRAMESSSPPKEPSCVLFLPHPHKFS 331
Query: 643 SEEDIPEGSSFTAGERMHELVKVFVLLHQLQIFTHGRALPDQPLIYHPCDHRTNSRAHTS 702
E++ GSSF AGERM ELVKVFVLLHQLQ+FT GRA P+QP I P D N RA S
Sbjct: 332 LEDNTSTGSSFDAGERMQELVKVFVLLHQLQLFTLGRASPEQPSIDPPGDLPANCRAQIS 511
Query: 703 GL-AAVPKPGTEMNLVNAVPCRIAFERGKERHFCFLAISVGTSG 745
G+ + PK GTE++LVNAVPCR+AF+ GKE H F AIS G SG
Sbjct: 512 GIDVSGPKAGTEISLVNAVPCRVAFKSGKEHHLYFQAISSGISG 643
Score = 99.8 bits (247), Expect = 4e-21
Identities = 46/51 (90%), Positives = 50/51 (97%)
Frame = +2
Query: 479 VPRFQVLDALVSLFCRSNISAETLWDGGWLLRQLLPYSESEFNNHHLELLK 529
V RFQV+DALVSLFCRS+ISAETLWDGGWLLRQLLPYSESEFN+HHLE+LK
Sbjct: 2 VNRFQVIDALVSLFCRSSISAETLWDGGWLLRQLLPYSESEFNSHHLEVLK 154
>BE322047 similar to GP|17063189|gb| unknown protein {Arabidopsis thaliana},
partial (9%)
Length = 410
Score = 91.3 bits (225), Expect = 1e-18
Identities = 42/54 (77%), Positives = 52/54 (95%)
Frame = +2
Query: 1 DFVIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLL 54
DFVIEALRSIAEL+TYGDQ+DP++FEFFMEKQV+G+FVRILKLS+TIS+P ++
Sbjct: 233 DFVIEALRSIAELMTYGDQNDPSYFEFFMEKQVMGEFVRILKLSKTISVPSSIV 394
>BE248171
Length = 405
Score = 73.9 bits (180), Expect = 2e-13
Identities = 36/64 (56%), Positives = 47/64 (73%)
Frame = +3
Query: 1 DFVIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLLQTVSIM 60
+FV++ L+ I E+VTYGD+ DP FE FME QV+ +FVRILK S +I LLQ +SIM
Sbjct: 186 EFVVDLLQCIVEIVTYGDKQDPLIFECFMEHQVLAEFVRILKTSEISNIEAPLLQYLSIM 365
Query: 61 IQNL 64
IQN+
Sbjct: 366 IQNM 377
>AW225616 similar to GP|21740506|em OSJNBa0029H02.1 {Oryza sativa}, partial
(6%)
Length = 448
Score = 31.2 bits (69), Expect = 1.8
Identities = 17/67 (25%), Positives = 31/67 (45%), Gaps = 8/67 (11%)
Frame = -3
Query: 134 AIRFAFHEESMIRAAVRAVTLNVYHVGDDSVNRYITSEPHTD--------YFSNLVSFFR 185
AIR+ H+E + + + TL +++ G + + D YF N++SFF
Sbjct: 401 AIRWELHKEKYLEQ-IHSSTLEIHNTGKGEKGQALVRPQEADCQPLYSTIYFGNMISFFC 225
Query: 186 KQCMDLN 192
C +L+
Sbjct: 224 GHCQNLH 204
>TC80639 weakly similar to PIR|G96621|G96621 probable disease resistance
protein F23H11.10 [imported] - Arabidopsis thaliana,
partial (4%)
Length = 836
Score = 30.0 bits (66), Expect = 4.0
Identities = 23/94 (24%), Positives = 40/94 (42%), Gaps = 3/94 (3%)
Frame = -2
Query: 176 YFSNLVSFFRKQCMDLNRLISETLKNPGPDSNSTVTTAVDEIEDNLYYFSDIISAGIPDV 235
+FS+ + +F D+ L+S ++ I L+ DII I V
Sbjct: 517 FFSSCIIYFTYSGFDVLNLVSYLETESHSFDTVCSNSSTTRIRSMLFLKKDIIINDIFHV 338
Query: 236 ERL---ITDSILMLLIFPVLLPSLRTVADNDMQS 266
+R + D + L+F ++ PS+R + D D S
Sbjct: 337 KRCFYQLLDHVFDFLVFYIISPSIR-IVDEDFNS 239
>TC93496 similar to GP|9757837|dbj|BAB08274.1 contains similarity to
chloroplast nucleoid DNA-binding
protein~gene_id:MMG4.12, partial (28%)
Length = 988
Score = 29.3 bits (64), Expect = 6.9
Identities = 14/27 (51%), Positives = 19/27 (69%), Gaps = 1/27 (3%)
Frame = +3
Query: 154 LNVYHVGDDSVNRY-ITSEPHTDYFSN 179
LN G+DS++R+ IT P+ DYFSN
Sbjct: 801 LNFTSSGNDSLSRWAITPRPYADYFSN 881
>BI309193 similar to GP|1223582|emb| PAC {Arabidopsis thaliana}, partial
(31%)
Length = 631
Score = 28.9 bits (63), Expect = 9.0
Identities = 12/27 (44%), Positives = 18/27 (66%), Gaps = 1/27 (3%)
Frame = -2
Query: 613 DEW-RKCKRAMESSSPPKEPKCMLFPP 638
DEW +KCK+ M + P ++P +L PP
Sbjct: 570 DEWLKKCKKIMIDTFPREDPYSILVPP 490
>TC86578 similar to PIR|C85440|C85440 myb-related protein [imported] -
Arabidopsis thaliana, partial (42%)
Length = 1283
Score = 28.9 bits (63), Expect = 9.0
Identities = 13/36 (36%), Positives = 21/36 (58%)
Frame = -2
Query: 43 LSRTISIPLQLLQTVSIMIQNLQSEHAIYYMFSNEH 78
LSR IS P+ L+T SI + ++ +Y++ N H
Sbjct: 841 LSRNISKPINSLKTHSISLFHITPHFILYHLLHNRH 734
>TC77552 similar to GP|6984224|gb|AAF34800.1| 60S ribosomal protein L35
{Euphorbia esula}, partial (93%)
Length = 443
Score = 28.9 bits (63), Expect = 9.0
Identities = 25/95 (26%), Positives = 43/95 (44%)
Frame = +2
Query: 42 KLSRTISIPLQLLQTVSIMIQNLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISF 101
K+S+ + L + Q ++++ QN ++ Y +H YL D R ++
Sbjct: 188 KISKIKVVRLNIAQVLTVISQNQKTALRAAY----KHKKYL---PLDLRPKKT------- 325
Query: 102 LRAIGGKLNKNTISLLVKTRNDEVVSFPLYVEAIR 136
RAI +L KN +SL + + FPL AI+
Sbjct: 326 -RAIRRRLTKNQLSLKTDREKKKEIYFPLRKYAIK 427
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.139 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,363,766
Number of Sequences: 36976
Number of extensions: 447766
Number of successful extensions: 2166
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 2141
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2162
length of query: 880
length of database: 9,014,727
effective HSP length: 105
effective length of query: 775
effective length of database: 5,132,247
effective search space: 3977491425
effective search space used: 3977491425
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC148819.3


