

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148817.2 - phase: 0
(608 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
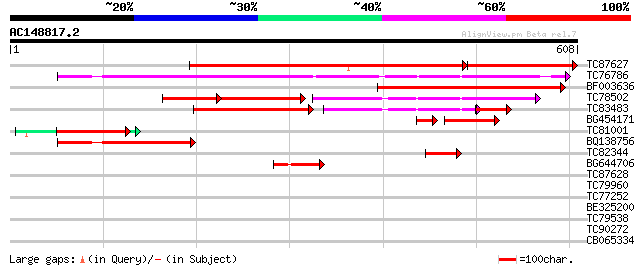
Score E
Sequences producing significant alignments: (bits) Value
TC87627 similar to SP|P37221|MAOM_SOLTU NAD-dependent malic enzy... 420 e-153
TC76786 similar to SP|P51615|MAOX_VITVI NADP-dependent malic enz... 414 e-116
BF003636 similar to GP|8894554|emb NAD-dependent malic enzyme (m... 394 e-110
TC78502 similar to GP|8118507|gb|AAF73006.1| NADP-dependent mali... 160 1e-79
TC83483 homologue to GP|510876|emb|CAA56354.1| NADP dependent ma... 135 8e-62
BG454171 homologue to GP|8894554|emb| NAD-dependent malic enzyme... 123 2e-36
TC81001 similar to SP|P37225|MAON_SOLTU NAD-dependent malic enzy... 145 6e-35
BQ138756 homologue to GP|510876|emb| NADP dependent malic enzyme... 138 7e-33
TC82344 similar to SP|P37221|MAOM_SOLTU NAD-dependent malic enzy... 62 8e-10
BG644706 PIR|E84508|E84 malate oxidoreductase (malic enzyme) [im... 60 2e-09
TC87628 similar to SP|P37224|MAOM_AMAHP NAD-dependent malic enzy... 37 0.017
TC79960 similar to PIR|T52419|T52419 CTF2A protein [imported] - ... 30 2.7
TC77252 homologue to GP|13122294|dbj|BAB32888. Dim1 homolog {Ara... 30 2.7
BE325200 similar to GP|19920182|gb| Putative Glucan 1 3-beta-glu... 30 3.5
TC79538 similar to GP|21594039|gb|AAM65957.1 carbonate dehydrata... 30 3.5
TC90272 similar to PIR|T45686|T45686 receptor-like protein kinas... 29 4.6
CB065334 weakly similar to PIR|B83279|B832 hypothetical protein ... 28 7.8
>TC87627 similar to SP|P37221|MAOM_SOLTU NAD-dependent malic enzyme 62 kDa
isoform mitochondrial precursor (EC 1.1.1.39) (NAD-ME).
[Potato], partial (67%)
Length = 1548
Score = 420 bits (1079), Expect(2) = e-153
Identities = 205/304 (67%), Positives = 255/304 (83%), Gaps = 5/304 (1%)
Frame = +2
Query: 193 DLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNNQKLLDDRLYLGLRQPRLEGEE 252
DLGVQGIGI +GKLD+YVAAAGINPQR+LP+M+DVGTNN+KLL+D LYLGL+Q RL+G++
Sbjct: 2 DLGVQGIGITIGKLDLYVAAAGINPQRVLPVMIDVGTNNKKLLEDPLYLGLQQHRLDGDD 181
Query: 253 YLSIVDEFMEAVHARWPKAIVQFEDFQMKWAFETLKRYRHKFCMFNDDIQGTAGVALAGL 312
YL+++DEFMEAV RWP IVQFEDFQ KWAF+ L+RYR + MFNDD+QGTAGVA+AGL
Sbjct: 182 YLAVIDEFMEAVFTRWPNVIVQFEDFQSKWAFKLLQRYRTTYRMFNDDVQGTAGVAIAGL 361
Query: 313 LGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQAVSRMSGCSETA---AKSQFFLI 369
LG VRAQGRP+ DF KQKIV+ GAGSAG+GVLN A + ++RM G +E A AKSQF+++
Sbjct: 362 LGAVRAQGRPMIDFPKQKIVVAGAGSAGIGVLNAARKTMARMLGNNEVAFQSAKSQFWVV 541
Query: 370 DKNGLVTTERSNLDPAAVPFAKNPR--DLEGLAEGASIIEVVKKVKPHVLLGLSGVGGVF 427
D GL+T R N+DP A+PFA+N + D +GL EGAS+ EVVK+VKP VLLGLS VGG+F
Sbjct: 542 DAKGLITEGRENIDPDALPFARNLKEMDRQGLREGASLAEVVKQVKPDVLLGLSAVGGLF 721
Query: 428 NEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGENIVFASGSPFENVDLGNGR 487
+ +VL+A+++S ST+PAIFAMSNPT NAECT +AF+ G+NI+FASGSPF NVDLGNG
Sbjct: 722 SNEVLEALKDSTSTRPAIFAMSNPTKNAECTPDEAFSILGDNIIFASGSPFSNVDLGNGN 901
Query: 488 AGHV 491
GH+
Sbjct: 902 IGHL 913
Score = 140 bits (353), Expect(2) = e-153
Identities = 67/117 (57%), Positives = 90/117 (76%)
Frame = +3
Query: 492 NQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLASYMTEEDIQKGILYPSIDCIRNV 551
NQ NNMYLFPGIGLG+LLSG+ ++DGMLQAA+E LA+YM+EE++ KGI++PSI IR++
Sbjct: 915 NQGNNMYLFPGIGLGTLLSGSRIVSDGMLQAAAERLAAYMSEEEVLKGIIFPSISRIRDI 1094
Query: 552 TAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSEDDTVEYVRGNMWYPEYCPLVHEK 608
T E+ AAV+ A+ E LAEG+ + ++EL +S D+ EYV NMW PEY LV+ K
Sbjct: 1095 TKEIAAAVIEEAVEEDLAEGYHEMDARELRKLSRDEIKEYVINNMWNPEYPTLVYRK 1265
>TC76786 similar to SP|P51615|MAOX_VITVI NADP-dependent malic enzyme (EC
1.1.1.40) (NADP-ME). [Grape] {Vitis vinifera}, complete
Length = 2039
Score = 414 bits (1063), Expect = e-116
Identities = 230/553 (41%), Positives = 332/553 (59%), Gaps = 3/553 (0%)
Frame = +1
Query: 52 FPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLH 111
F ERD LRGLLPP + S Q Q + M + R E V L K+ + L
Sbjct: 298 FTEKERDAHYLRGLLPPTVSSQQLQEKKLMHNIRQYE----------VPLQKYVAMMDLQ 447
Query: 112 DRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYN 171
+RNE L+Y+ LIDN++E PI+YTP VG CQ Y +Y+RP+G+Y S K+KG+++ ++ N
Sbjct: 448 ERNERLFYKLLIDNVEELLPIVYTPVVGEACQKYGSIYKRPQGLYISLKEKGKILEVLKN 627
Query: 172 WPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNN 231
WP + +IV+TDG RILGLGDLG QG+GIPVGKL +Y A G+ P LP+ +DVGTNN
Sbjct: 628 WPERSIQVIVVTDGERILGLGDLGCQGMGIPVGKLALYSALGGVRPSACLPVTIDVGTNN 807
Query: 232 QKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARW-PKAIVQFEDFQMKWAFETLKRY 290
+KLL+D Y+GLRQ R G+EY ++ EFM AV + K +VQFEDF AFE L +Y
Sbjct: 808 EKLLNDEFYIGLRQKRATGKEYYDLLHEFMTAVKQNYGEKVLVQFEDFANHNAFELLAKY 987
Query: 291 RHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQA 350
+FNDDIQGTA V LAG++ +++ G L + + +GAG AG G+ +
Sbjct: 988 GTTHLVFNDDIQGTAAVVLAGVVASLKLIGGTLPE---HTFLFLGAGEAGTGIAELIALE 1158
Query: 351 VSRMSGCSETAAKSQFFLIDKNGLVTTERSN-LDPAAVPFAKNPRDLEGLAEGASIIEVV 409
+S+ + ++ + +L+D GL+ + R+N L P+A + +++++ V
Sbjct: 1159MSKQTKAPIEESRKKIWLVDSKGLIVSSRANSLQHFKKPWAHEHEPV------STLLDAV 1320
Query: 410 KKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGEN 469
K +KP VL+G SGVG F ++V++AM E ++ P I A+SNPT +ECTA +A+ +
Sbjct: 1321KIIKPTVLIGSSGVGKTFTKEVVEAMTE-INKIPLILALSNPTSQSECTAEEAYTWSEGR 1497
Query: 470 IVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLAS 529
+FASGSPF+ V+ G+ + Q+NN Y+FPG GLG ++SGA + D ML AASE LA
Sbjct: 1498AIFASGSPFDPVEY-KGKTYYSGQSNNAYIFPGFGLGIVMSGAIRVHDDMLLAASEALAK 1674
Query: 530 YMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSE-DDT 588
+TEE+ +KG+ YP IR ++A + A V A GLA H+ ++
Sbjct: 1675QVTEENYKKGLTYPPFSDIRKISANIAANVAAKAYELGLA-----------THLPRPENL 1821
Query: 589 VEYVRGNMWYPEY 601
V+Y M+ P Y
Sbjct: 1822VKYAESCMYSPLY 1860
>BF003636 similar to GP|8894554|emb NAD-dependent malic enzyme (malate
oxidoreductase) {Cicer arietinum}, partial (66%)
Length = 608
Score = 394 bits (1012), Expect = e-110
Identities = 199/202 (98%), Positives = 200/202 (98%)
Frame = +2
Query: 395 DLEGLAEGASIIEVVKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMN 454
DL+GLAEGASIIEVVKKV PHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMN
Sbjct: 2 DLDGLAEGASIIEVVKKVLPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMN 181
Query: 455 AECTAIDAFNHAGENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHH 514
AECTAIDAFNHAGENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHH
Sbjct: 182 AECTAIDAFNHAGENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHH 361
Query: 515 ITDGMLQAASECLASYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGG 574
ITDGMLQAASECLASYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGG
Sbjct: 362 ITDGMLQAASECLASYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGG 541
Query: 575 VGSKELEHMSEDDTVEYVRGNM 596
VGSKELEHMSEDDTVEYVR NM
Sbjct: 542 VGSKELEHMSEDDTVEYVRRNM 607
>TC78502 similar to GP|8118507|gb|AAF73006.1| NADP-dependent malic protein
{Ricinus communis}, partial (65%)
Length = 1540
Score = 160 bits (406), Expect(3) = 1e-79
Identities = 89/246 (36%), Positives = 138/246 (55%), Gaps = 1/246 (0%)
Frame = +2
Query: 325 DFAKQKIVIVGAGSAGLGVLNMAVQAVSRMSGCSETAAKSQFFLIDKNGLVTTERS-NLD 383
+ A K + +GAG AG G+ + S+ + + +L+D GL+ + R +L
Sbjct: 482 NLADHKFLFLGAGEAGTGIAELIALETSKQTNAPLDEVRKNIWLVDSKGLIVSSRKESLQ 661
Query: 384 PAAVPFAKNPRDLEGLAEGASIIEVVKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKP 443
P+A ++ L ++ V K+KP VL+G SG G F ++V++AM S++ KP
Sbjct: 662 HFKKPWAHEHEPVKNL------LDAVNKIKPTVLIGTSGQGSAFTKEVVEAMA-SINEKP 820
Query: 444 AIFAMSNPTMNAECTAIDAFNHAGENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGI 503
I ++SNPT +ECTA +A+ +FASGSPF V+ G+ QANN Y+FPG
Sbjct: 821 IILSLSNPTSQSECTAEEAYTWTQGRAIFASGSPFSPVEY-KGKVFVPGQANNAYIFPGF 997
Query: 504 GLGSLLSGAHHITDGMLQAASECLASYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAA 563
GLG ++SG + D +L AASE LA ++EE+ +KG+++P IR ++A + A V A
Sbjct: 998 GLGLIMSGTIRVHDDLLLAASEALAEQVSEENFEKGLIFPPFTNIRKISAHIAAKVAAKA 1177
Query: 564 IAEGLA 569
GLA
Sbjct: 1178YELGLA 1195
Score = 95.5 bits (236), Expect(3) = 1e-79
Identities = 47/96 (48%), Positives = 67/96 (68%), Gaps = 1/96 (1%)
Frame = +1
Query: 223 IMLDVGTNNQKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARW-PKAIVQFEDFQMK 281
++L GTNN++LL+D LY+GL+Q R G+EY ++ EFM AV + K ++QFEDF
Sbjct: 181 LLLMSGTNNERLLNDELYIGLKQRRATGQEYSELMHEFMTAVKQTYGEKVLIQFEDFANH 360
Query: 282 WAFETLKRYRHKFCMFNDDIQGTAGVALAGLLGTVR 317
AF+ L++YR +FNDDIQGTA V LAGL+ ++
Sbjct: 361 NAFDLLEKYRSTHLVFNDDIQGTASVVLAGLVAALK 468
Score = 80.9 bits (198), Expect(3) = 1e-79
Identities = 35/63 (55%), Positives = 45/63 (70%)
Frame = +3
Query: 165 MMSMIYNWPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIM 224
+ ++ NWP + +IV+TDG RILGLGDLG QG+GIPVGKL +Y A G+ P LPI
Sbjct: 6 IQEVLRNWPEKNIQVIVVTDGERILGLGDLGCQGMGIPVGKLSLYTALGGVRPSACLPIT 185
Query: 225 LDV 227
+DV
Sbjct: 186 IDV 194
>TC83483 homologue to GP|510876|emb|CAA56354.1| NADP dependent malic enzyme
{Phaseolus vulgaris}, partial (56%)
Length = 1003
Score = 135 bits (339), Expect(3) = 8e-62
Identities = 67/129 (51%), Positives = 87/129 (66%), Gaps = 1/129 (0%)
Frame = +2
Query: 198 GIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNNQKLLDDRLYLGLRQPRLEGEEYLSIV 257
G+GIPVGKL +Y A G+ P LPI +DVGTNN+KLL+D Y+GL Q R G+EY ++
Sbjct: 2 GMGIPVGKLSLYTALGGVRPSSCLPITIDVGTNNEKLLNDEFYIGLXQKRATGKEYAELL 181
Query: 258 DEFMEAVHARW-PKAIVQFEDFQMKWAFETLKRYRHKFCMFNDDIQGTAGVALAGLLGTV 316
+EFM AV + K +VQFEDF AF+ L +Y +FNDDIQGTA V LAGLL ++
Sbjct: 182 EEFMHAVKQNYGEKVLVQFEDFANHNAFDLLDKYSSSHLVFNDDIQGTASVVLAGLLASL 361
Query: 317 RAQGRPLSD 325
+ G L+D
Sbjct: 362 KLVGGTLAD 388
Score = 109 bits (272), Expect(3) = 8e-62
Identities = 62/171 (36%), Positives = 97/171 (56%), Gaps = 1/171 (0%)
Frame = +3
Query: 337 GSAGLGVLNMAVQAVSRMSGCSETAAKSQFFLIDKNGLVTTER-SNLDPAAVPFAKNPRD 395
G AG G+ + +S+ + + + +L+D GL+ + R +L P+A
Sbjct: 414 GKAGTGIAELIALEISKQTKAPVEETRKKIWLVDSKGLIVSSRLQSLQHFKKPWAHEHEP 593
Query: 396 LEGLAEGASIIEVVKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNA 455
++ L + VK +KP VL+G SGVG F ++V++AM S++ KP I A+SNPT +
Sbjct: 594 VKEL------LNAVKAIKPTVLIGSSGVGKTFTKEVVEAMA-SLNEKPLILALSNPTSQS 752
Query: 456 ECTAIDAFNHAGENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLG 506
ECTA +A+ + +FASGSPF+ V+ G+ Q+NN Y+FPG GLG
Sbjct: 753 ECTAEEAYTWSKGKAIFASGSPFDPVEY-EGKVFVPGQSNNAYIFPGFGLG 902
Score = 32.7 bits (73), Expect(3) = 8e-62
Identities = 15/39 (38%), Positives = 25/39 (63%)
Frame = +1
Query: 500 FPGIGLGSLLSGAHHITDGMLQAASECLASYMTEEDIQK 538
FP + G ++SGA + D ML A SE LA+ +++E+ +
Sbjct: 883 FPDLVWGLIISGAIRVRDEMLLAXSEALAAQVSQENYDR 999
>BG454171 homologue to GP|8894554|emb| NAD-dependent malic enzyme (malate
oxidoreductase) {Cicer arietinum}, partial (26%)
Length = 487
Score = 123 bits (308), Expect(2) = 2e-36
Identities = 59/59 (100%), Positives = 59/59 (100%)
Frame = +3
Query: 467 GENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASE 525
GENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASE
Sbjct: 93 GENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASE 269
Score = 48.1 bits (113), Expect(2) = 2e-36
Identities = 22/22 (100%), Positives = 22/22 (100%)
Frame = +1
Query: 437 ESVSTKPAIFAMSNPTMNAECT 458
ESVSTKPAIFAMSNPTMNAECT
Sbjct: 1 ESVSTKPAIFAMSNPTMNAECT 66
>TC81001 similar to SP|P37225|MAON_SOLTU NAD-dependent malic enzyme 59 kDa
isoform mitochondrial precursor (EC 1.1.1.39) (NAD-ME).
[Potato], partial (20%)
Length = 679
Score = 145 bits (365), Expect = 6e-35
Identities = 67/79 (84%), Positives = 74/79 (92%)
Frame = +1
Query: 51 GFPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRL 110
GFPLTERDRLGLRGLLPPR+ISF++QYDRF+ SYRSLEKNT GQ + +VSL+KWRILNRL
Sbjct: 409 GFPLTERDRLGLRGLLPPRVISFEQQYDRFLDSYRSLEKNTLGQPENVVSLAKWRILNRL 588
Query: 111 HDRNETLYYRALIDNIKEF 129
HDRNETLYYR LIDNIKEF
Sbjct: 589 HDRNETLYYRVLIDNIKEF 645
Score = 45.4 bits (106), Expect = 6e-05
Identities = 44/139 (31%), Positives = 53/139 (37%), Gaps = 5/139 (3%)
Frame = +2
Query: 7 LEECGRLRDS---PHHRDRGCSRRRYLHRVWFINVVLIFFMILGSIR--GFPLTERDRLG 61
L ECG RDS P+ DRG S R+ F NVVLI ILGS R GF + G
Sbjct: 263 LGECGNSRDSFQLPNSSDRGVSPPRFPVLALFTNVVLISSTILGSTRILGFL*LKGIDWG 442
Query: 62 LRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLHDRNETLYYRA 121
G P + F + + L +
Sbjct: 443 FEGFFLPVLYHLSSNMTAFWIRTDLWRRILWDNQKMLFPLQNGES*IDCMTGTKLCTTEF 622
Query: 122 LIDNIKEFAPIIYTPTVGL 140
L+ +K FAPIIYTPT GL
Sbjct: 623 LLIILKSFAPIIYTPTGGL 679
>BQ138756 homologue to GP|510876|emb| NADP dependent malic enzyme {Phaseolus
vulgaris}, partial (30%)
Length = 680
Score = 138 bits (347), Expect = 7e-33
Identities = 67/148 (45%), Positives = 95/148 (63%)
Frame = +2
Query: 52 FPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLH 111
F ERD +RGLLPP + + + Q R M + R + V L K+ L L
Sbjct: 266 FTEKERDAHYVRGLLPPAVFTQELQEKRLMHNLRQYD----------VPLHKYIALMDLQ 415
Query: 112 DRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYN 171
+RNE L+Y+ +IDN++E P++YTPTVG CQ Y +YRRP+G+Y S K+KG+++ ++ N
Sbjct: 416 ERNERLFYKLMIDNVEELLPVVYTPTVGEACQKYGSIYRRPQGLYISLKEKGKILEVLKN 595
Query: 172 WPAPQVDMIVLTDGSRILGLGDLGVQGI 199
WP + +IV+TDG RILGLGDLG QG+
Sbjct: 596 WPEKTIQVIVVTDGERILGLGDLGCQGM 679
>TC82344 similar to SP|P37221|MAOM_SOLTU NAD-dependent malic enzyme 62 kDa
isoform mitochondrial precursor (EC 1.1.1.39) (NAD-ME).
[Potato], partial (5%)
Length = 675
Score = 61.6 bits (148), Expect = 8e-10
Identities = 28/38 (73%), Positives = 32/38 (83%)
Frame = +1
Query: 447 AMSNPTMNAECTAIDAFNHAGENIVFASGSPFENVDLG 484
AMSNPT NAECT +AF+ G+NI+FASGSPF NVDLG
Sbjct: 1 AMSNPTKNAECTPDEAFSILGDNIIFASGSPFSNVDLG 114
>BG644706 PIR|E84508|E84 malate oxidoreductase (malic enzyme) [imported] -
Arabidopsis thaliana, partial (5%)
Length = 645
Score = 60.5 bits (145), Expect = 2e-09
Identities = 31/54 (57%), Positives = 38/54 (69%)
Frame = +3
Query: 284 FETLKRYRHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAG 337
F + Y++ +F QGTAGVA+AGLLG VRAQGRP+ DF K KIV+ GAG
Sbjct: 39 FNQVLCYKYDIILFFT--QGTAGVAIAGLLGAVRAQGRPMIDFPKMKIVVAGAG 194
>TC87628 similar to SP|P37224|MAOM_AMAHP NAD-dependent malic enzyme 65 kDa
isoform mitochondrial precursor (EC 1.1.1.39)
(NAD-ME)., partial (5%)
Length = 661
Score = 37.4 bits (85), Expect = 0.017
Identities = 16/32 (50%), Positives = 21/32 (65%)
Frame = +3
Query: 577 SKELEHMSEDDTVEYVRGNMWYPEYCPLVHEK 608
++EL +S D+ EYV NMW PEY LV+ K
Sbjct: 489 ARELRKLSRDEIKEYVINNMWNPEYPTLVYRK 584
>TC79960 similar to PIR|T52419|T52419 CTF2A protein [imported] - Arabidopsis
thaliana, partial (53%)
Length = 907
Score = 30.0 bits (66), Expect = 2.7
Identities = 24/77 (31%), Positives = 38/77 (49%), Gaps = 11/77 (14%)
Frame = +1
Query: 315 TVRAQGRPLSDFAKQKIVIVGAGSAG---------LGVLNMAVQAVS--RMSGCSETAAK 363
T++AQ SD K+ +VIVG G AG LGV ++ ++ R G S T +K
Sbjct: 160 TIKAQS---SDVRKEHVVIVGGGIAGLATALSLHRLGVRSLVLEQSESLRTGGTSLTLSK 330
Query: 364 SQFFLIDKNGLVTTERS 380
+ + +D G+ R+
Sbjct: 331 NGWSALDSIGVANYLRT 381
>TC77252 homologue to GP|13122294|dbj|BAB32888. Dim1 homolog {Arabidopsis
thaliana}, complete
Length = 1158
Score = 30.0 bits (66), Expect = 2.7
Identities = 14/46 (30%), Positives = 26/46 (56%)
Frame = -1
Query: 445 IFAMSNPTMNAECTAIDAFNHAGENIVFASGSPFENVDLGNGRAGH 490
IF ++ T + CT++D+FN E ++F SP + V + + + H
Sbjct: 894 IFGSNDETTSLPCTSVDSFNDVNE-LLFVLHSPIDFVVVSSSKIYH 760
>BE325200 similar to GP|19920182|gb| Putative Glucan 1 3-beta-glucosidase
precursor {Oryza sativa (japonica cultivar-group)},
partial (14%)
Length = 364
Score = 29.6 bits (65), Expect = 3.5
Identities = 10/29 (34%), Positives = 19/29 (65%)
Frame = +2
Query: 32 RVWFINVVLIFFMILGSIRGFPLTERDRL 60
++WF+N++++F MIL G P+ R+
Sbjct: 59 KLWFLNLIMLFCMILSMTHGRPINPEFRV 145
>TC79538 similar to GP|21594039|gb|AAM65957.1 carbonate dehydratase-like
protein {Arabidopsis thaliana}, partial (57%)
Length = 1260
Score = 29.6 bits (65), Expect = 3.5
Identities = 31/101 (30%), Positives = 49/101 (47%)
Frame = -3
Query: 445 IFAMSNPTMNAECTAIDAFNHAGENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIG 504
IF+M+NPT + CTAI H G A+ + N D+ N ++ H ++
Sbjct: 781 IFSMNNPTFDKACTAIILHTHKGAYTPTAAVA--NN*DVLNQKSVHCKF*CSICFG---- 620
Query: 505 LGSLLSGAHHITDGMLQAASECLASYMTEEDIQKGILYPSI 545
GS L G + I++ A EC+ S++ ++I G YP I
Sbjct: 619 -GSTLKGRNQISN---IADCECV-SWLKSQNI--GGAYPGI 518
>TC90272 similar to PIR|T45686|T45686 receptor-like protein kinase homolog -
Arabidopsis thaliana, partial (58%)
Length = 941
Score = 29.3 bits (64), Expect = 4.6
Identities = 19/64 (29%), Positives = 34/64 (52%), Gaps = 1/64 (1%)
Frame = +3
Query: 392 NPRDLEGLAEGASIIEVVKKVKPHVLLGLSGVGGVFNEQVL-KAMRESVSTKPAIFAMSN 450
NPR +GLAE + IE++ + + L+ L G NE +L E+ + K ++ S
Sbjct: 24 NPRSHQGLAEFRTEIEMLSQFRHRHLVSLIGYCDERNEMILIYEYMENGTVKSHLYGSSL 203
Query: 451 PTMN 454
P+++
Sbjct: 204 PSLS 215
>CB065334 weakly similar to PIR|B83279|B832 hypothetical protein PA2929
[imported] - Pseudomonas aeruginosa (strain PAO1),
partial (28%)
Length = 599
Score = 28.5 bits (62), Expect = 7.8
Identities = 15/26 (57%), Positives = 17/26 (64%)
Frame = -1
Query: 297 FNDDIQGTAGVALAGLLGTVRAQGRP 322
F DD A +A AGL GTVR +GRP
Sbjct: 218 FGDDF---AQIAQAGLTGTVRLRGRP 150
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.139 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,502,943
Number of Sequences: 36976
Number of extensions: 224936
Number of successful extensions: 985
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 965
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 968
length of query: 608
length of database: 9,014,727
effective HSP length: 102
effective length of query: 506
effective length of database: 5,243,175
effective search space: 2653046550
effective search space used: 2653046550
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148817.2


