

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148816.5 + phase: 0 /pseudo
(211 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
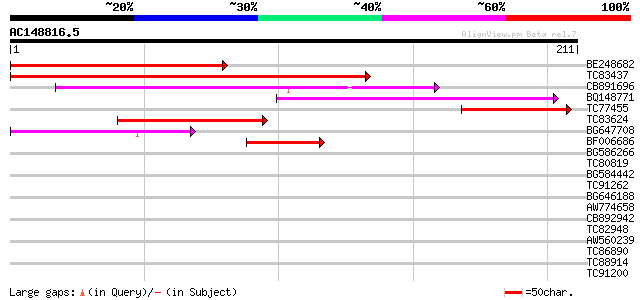
BE248682 similar to GP|18568269|gb putative gag-pol polyprotein ... 121 2e-28
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 120 5e-28
CB891696 59 2e-09
BQ148771 52 1e-07
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 52 2e-07
TC83624 homologue to PIR|G84581|G84581 copia-like retroelement p... 49 2e-06
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 42 2e-04
BF006686 40 5e-04
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 35 0.021
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 35 0.027
BG584442 34 0.047
TC91262 similar to GP|20804797|dbj|BAB92481. putative amino acid... 34 0.047
BG646188 similar to PIR|T48467|T48 aspartyl aminopeptidase-like ... 31 0.40
AW774658 similar to GP|2808681|emb| Hcr9-4B {Lycopersicon hirsut... 27 0.54
CB892942 similar to GP|13569548|gb unknown {Arabidopsis thaliana... 29 1.2
TC82948 28 2.0
AW560239 weakly similar to GP|14495231|dbj hypothetical protein~... 27 4.4
TC86890 weakly similar to GP|4406801|gb|AAD20110.1| unknown prot... 27 5.7
TC88914 weakly similar to PIR|F96672|F96672 Similar to Flavonol ... 26 9.8
TC91200 weakly similar to GP|10177640|dbj|BAB10787. contains sim... 26 9.8
>BE248682 similar to GP|18568269|gb putative gag-pol polyprotein {Zea mays},
partial (1%)
Length = 441
Score = 121 bits (303), Expect = 2e-28
Identities = 57/81 (70%), Positives = 65/81 (79%)
Frame = +3
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
VSHLQYADDTLCIG +V NLWT+KA+L+GF+M SGLK+NF KSSL+GINV DFM AC
Sbjct: 159 VSHLQYADDTLCIGMPTVDNLWTLKALLQGFEMASGLKVNFHKSSLIGINVPRDFMEAAC 338
Query: 61 DFLNCSAGSITFKYLGLPVGA 81
FLNC SI F YLGLP G+
Sbjct: 339 RFLNCREESIPFIYLGLPGGS 401
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 120 bits (300), Expect = 5e-28
Identities = 58/134 (43%), Positives = 88/134 (65%)
Frame = +2
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
VSHLQ+A+DTL + + N+ ++A L F +SGLK+NF KS LV +N++ +++ A
Sbjct: 542 VSHLQFANDTLLLETKNWANIRALRAALVIF*AMSGLKVNFHKSGLVCVNIAPSWLSEAA 721
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
L+ G + F YLG+P+ N R +S WEP+V I RL W++R++SFGGR+VLL SV
Sbjct: 722 SVLSWKVGKVPFLYLGMPIEGNSRRLSFWEPIVNRIKARLTGWNSRFLSFGGRLVLLKSV 901
Query: 121 LNSMPIFYLSFLKM 134
L S+ ++ L K+
Sbjct: 902 LTSLSVYALPSSKL 943
>CB891696
Length = 638
Score = 58.5 bits (140), Expect = 2e-09
Identities = 45/144 (31%), Positives = 76/144 (52%), Gaps = 1/144 (0%)
Frame = +1
Query: 18 VQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMACDFLNCSAGSITFKYLGL 77
V+N+ TMK I+ F++ S L +NF KS L+ +NV F ++ C + FKYLG+
Sbjct: 4 VENILTMKTIVSYFELASSLWVNFLKSGLINLNVIGHF*GW*NIYIKCKVH*VIFKYLGI 183
Query: 78 PVGANMRSMSTWEPLVETIGGRLNT-WSTRYISFGGRIVLLNSVLNSMPIFYLSFLKMLV 136
VG N ++ E L++ + L + W+T+ + +++ S I Y S +K+ V
Sbjct: 184 LVGENPCRVNM*ELLLKLLTN*LGSWWNTK*LWTQNGFSQIHAK*ISQNI-YFSLMKIPV 360
Query: 137 GVWKRIVRIQR*FLWGGVGGGKKI 160
V + I +++ FL G + KKI
Sbjct: 361 KV*ELISQLKTQFL*GNLKVTKKI 432
>BQ148771
Length = 680
Score = 52.4 bits (124), Expect = 1e-07
Identities = 25/105 (23%), Positives = 53/105 (49%)
Frame = -3
Query: 100 LNTWSTRYISFGGRIVLLNSVLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKK 159
L W ++S R+ L SV+ ++P++ + + + I ++QR F+WG ++
Sbjct: 573 LANWKANHLSLARRVTLAKSVIEAVPLYPMMTTIIPKACIEEIQKLQRKFVWGDTEVSRR 394
Query: 160 ISWVNWKSVCQQKENGGVRVKDIRVMNISLLAKWRWRLIDGREAL 204
V W+++ + K G+ ++ + VMN + + K W + G +L
Sbjct: 393 YHAVGWETMSKPKTIYGLGLRRLDVMNKACIMKLGWSIYSGSNSL 259
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 52.0 bits (123), Expect = 2e-07
Identities = 22/41 (53%), Positives = 31/41 (74%)
Frame = -3
Query: 169 CQQKENGGVRVKDIRVMNISLLAKWRWRLIDGREALWK*VL 209
C + GG+ V+DIR++N+SLLAKW WRL+ + +LWK VL
Sbjct: 969 CLPRCKGGLGVRDIRLVNVSLLAKWWWRLLQDQSSLWKEVL 847
>TC83624 homologue to PIR|G84581|G84581 copia-like retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(1%)
Length = 831
Score = 48.5 bits (114), Expect = 2e-06
Identities = 19/56 (33%), Positives = 38/56 (66%)
Frame = +1
Query: 41 FSKSSLVGINVSEDFMAMACDFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETI 96
FS+ + + +N+ E F+ + +FL C+ + F +LGLP+GAN + ST +P+++++
Sbjct: 256 FSRVNFMALNLEESFVEASPNFLLCNVNEVPFCFLGLPIGANPKRSSTRKPVLDSL 423
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 41.6 bits (96), Expect = 2e-04
Identities = 22/70 (31%), Positives = 35/70 (49%), Gaps = 1/70 (1%)
Frame = +1
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSL-VGINVSEDFMAMA 59
++HL +ADD+L +A++ T+ +L +Q SG +NF KS + NV M
Sbjct: 145 ITHLLFADDSLLFARANLTEAATIMQVLHSYQSASGQLVNFEKSEVSYSQNVPNQEKEMI 324
Query: 60 CDFLNCSAGS 69
C + GS
Sbjct: 325 CQQIAIKTGS 354
>BF006686
Length = 325
Score = 40.4 bits (93), Expect = 5e-04
Identities = 16/29 (55%), Positives = 21/29 (72%)
Frame = +3
Query: 89 WEPLVETIGGRLNTWSTRYISFGGRIVLL 117
WEPL+E + L +W + +SFGGRIVLL
Sbjct: 237 WEPLLEHVNKMLKSWGNKLLSFGGRIVLL 323
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 35.0 bits (79), Expect = 0.021
Identities = 22/77 (28%), Positives = 40/77 (51%), Gaps = 1/77 (1%)
Frame = -3
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLV-GINVSEDFMAMA 59
++HL +ADDT+ GK++ + + +I+ ++ SG IN +KS++ S+ +
Sbjct: 253 INHLLFADDTMFFGKSNASSCAILLSIMDKYRAASGRCIN*TKSAITFSSKTSQAIIDRV 74
Query: 60 CDFLNCSAGSITFKYLG 76
L + T KYLG
Sbjct: 73 KGELKIAKEGGTGKYLG 23
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 34.7 bits (78), Expect = 0.027
Identities = 17/35 (48%), Positives = 23/35 (65%), Gaps = 2/35 (5%)
Frame = +1
Query: 177 VRVKDIRV--MNISLLAKWRWRLIDGREALWK*VL 209
+R+K + V N+SLL KW WRL+ +E LW VL
Sbjct: 19 LRMKGLGVGAFNLSLLGKWCWRLLVDKEGLWHRVL 123
>BG584442
Length = 775
Score = 33.9 bits (76), Expect = 0.047
Identities = 21/79 (26%), Positives = 36/79 (44%), Gaps = 1/79 (1%)
Frame = +1
Query: 115 VLLNSVLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKK-ISWVNWKSVCQQKE 173
V++ L S+ + +S +L I +I F W VG +K + W++ + + K
Sbjct: 430 VMIKYALQSISSYVMSIFLLLNSQVDEIEKIMNTFSWVHVGENRKGMHWMS*EKLFVHKN 609
Query: 174 NGGVRVKDIRVMNISLLAK 192
GG+ D NI +L K
Sbjct: 610 YGGMGFTDFTTFNIPMLGK 666
>TC91262 similar to GP|20804797|dbj|BAB92481. putative amino acid or GABA
permease {Oryza sativa (japonica cultivar-group)},
partial (36%)
Length = 904
Score = 33.9 bits (76), Expect = 0.047
Identities = 14/32 (43%), Positives = 22/32 (68%)
Frame = +1
Query: 108 ISFGGRIVLLNSVLNSMPIFYLSFLKMLVGVW 139
I+ G I++L+ ++NS+PI +LSFL L W
Sbjct: 520 IAIHGGILVLHGIINSLPISWLSFLGQLAAFW 615
>BG646188 similar to PIR|T48467|T48 aspartyl aminopeptidase-like protein -
Arabidopsis thaliana, partial (30%)
Length = 776
Score = 30.8 bits (68), Expect = 0.40
Identities = 24/69 (34%), Positives = 36/69 (51%), Gaps = 1/69 (1%)
Frame = +3
Query: 78 PVGANMRSMSTWEPLVETIGGRL-NTWSTRYISFGGRIVLLNSVLNSMPIFYLSFLKMLV 136
P A++++ S V+T GG L +TW R +S GR++L S SF+ LV
Sbjct: 393 PKTASLKASSYMMVNVQTYGGGLWHTWFDRDLSVAGRVILKRS--------DKSFVHKLV 548
Query: 137 GVWKRIVRI 145
V + I+RI
Sbjct: 549 KVSRPILRI 575
>AW774658 similar to GP|2808681|emb| Hcr9-4B {Lycopersicon hirsutum}, partial
(4%)
Length = 665
Score = 27.3 bits (59), Expect(2) = 0.54
Identities = 12/19 (63%), Positives = 14/19 (73%)
Frame = -1
Query: 2 SHLQYADDTLCIGKASVQN 20
SHLQ+ADDTL +G S N
Sbjct: 383 SHLQFADDTLLLGVKSWAN 327
Score = 21.6 bits (44), Expect(2) = 0.54
Identities = 8/24 (33%), Positives = 17/24 (70%)
Frame = -3
Query: 24 MKAILRGFQMVSGLKINFSKSSLV 47
+++IL F+ +SGLK+N + ++
Sbjct: 321 LRSILVIFENMSGLKVNLREEVII 250
>CB892942 similar to GP|13569548|gb unknown {Arabidopsis thaliana}, partial
(45%)
Length = 767
Score = 29.3 bits (64), Expect = 1.2
Identities = 15/32 (46%), Positives = 19/32 (58%), Gaps = 2/32 (6%)
Frame = -1
Query: 174 NGGVRVKDIRVMNISLLA--KWRWRLIDGREA 203
+ G RV +RV SLL KW WR+ +G EA
Sbjct: 401 SSGTRVLCVRVQKTSLLQVKKWTWRVREGGEA 306
>TC82948
Length = 705
Score = 28.5 bits (62), Expect = 2.0
Identities = 11/44 (25%), Positives = 25/44 (56%)
Frame = +3
Query: 159 KISWVNWKSVCQQKENGGVRVKDIRVMNISLLAKWRWRLIDGRE 202
K+ V+W+ VC+ + G + ++ + +N +L K W ++ +E
Sbjct: 255 KVVKVSWEKVCRPIKEGSLGIRSLSKLNEALNLKLCWDMMISKE 386
>AW560239 weakly similar to GP|14495231|dbj hypothetical protein~similar to
Arabidopsis thaliana chromosome 1 T6A9.6, partial (10%)
Length = 630
Score = 27.3 bits (59), Expect = 4.4
Identities = 15/44 (34%), Positives = 25/44 (56%)
Frame = -3
Query: 166 KSVCQQKENGGVRVKDIRVMNISLLAKWRWRLIDGREALWK*VL 209
+ + Q+ E+G DIR+ N+ + A +W L+D EA K +L
Sbjct: 574 EKILQEWESGNTTF-DIRIPNMMITAYCKWGLLDKAEAYIKRLL 446
>TC86890 weakly similar to GP|4406801|gb|AAD20110.1| unknown protein
{Arabidopsis thaliana}, partial (54%)
Length = 2446
Score = 26.9 bits (58), Expect = 5.7
Identities = 10/22 (45%), Positives = 16/22 (72%)
Frame = +1
Query: 175 GGVRVKDIRVMNISLLAKWRWR 196
GGVR+K + +++I LL + WR
Sbjct: 790 GGVRIKTVPLLSIGLLCEVTWR 855
>TC88914 weakly similar to PIR|F96672|F96672 Similar to Flavonol
3-O-Glucosyltransferase [imported] - Arabidopsis
thaliana, partial (20%)
Length = 1051
Score = 26.2 bits (56), Expect = 9.8
Identities = 10/27 (37%), Positives = 16/27 (59%)
Frame = -3
Query: 125 PIFYLSFLKMLVGVWKRIVRIQR*FLW 151
PI + K L +WK I++I+ +LW
Sbjct: 647 PILKTFYAKFLSQIWKPILKIKENYLW 567
>TC91200 weakly similar to GP|10177640|dbj|BAB10787. contains similarity to
heparanase~gene_id:MGG23.2 {Arabidopsis thaliana},
partial (15%)
Length = 805
Score = 26.2 bits (56), Expect = 9.8
Identities = 17/64 (26%), Positives = 30/64 (46%), Gaps = 8/64 (12%)
Frame = +3
Query: 44 SSLVGINVSEDFMAMA--------CDFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVET 95
SS++G N+ +DF+ CD+ CS G + L L + ++ + PL
Sbjct: 477 SSVIG-NIDDDFICATLDWWPPQKCDYGTCSWGLASLLNLDLNNKIFLNAVKAFSPLKLR 653
Query: 96 IGGR 99
+GG+
Sbjct: 654 LGGK 665
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.330 0.142 0.465
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,787,115
Number of Sequences: 36976
Number of extensions: 115890
Number of successful extensions: 768
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 764
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 768
length of query: 211
length of database: 9,014,727
effective HSP length: 92
effective length of query: 119
effective length of database: 5,612,935
effective search space: 667939265
effective search space used: 667939265
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC148816.5


