

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148655.10 + phase: 0
(250 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
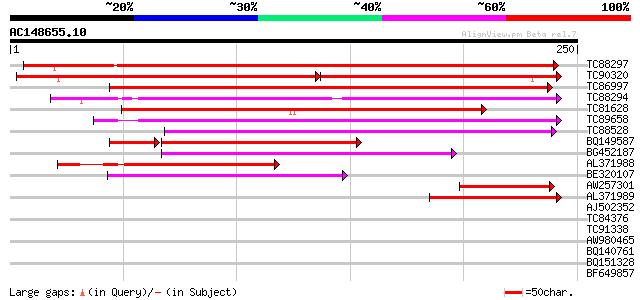
TC88297 similar to GP|15215586|gb|AAK91338.1 AT4g23630/F9D16_100... 281 2e-76
TC90320 similar to GP|17381218|gb|AAL36421.1 unknown protein {Ar... 154 2e-66
TC86997 similar to GP|6056199|gb|AAF02816.1| unknown protein {Ar... 186 7e-48
TC88294 weakly similar to GP|15215586|gb|AAK91338.1 AT4g23630/F9... 150 6e-37
TC81628 weakly similar to GP|12957724|gb|AAK09242.1 unknown prot... 141 3e-34
TC89658 weakly similar to PIR|A84527|A84527 hypothetical protein... 115 2e-26
TC88528 weakly similar to GP|4102690|gb|AAD01540.1| 24 kDa seed ... 110 4e-25
BQ149587 similar to GP|6056199|gb| unknown protein {Arabidopsis ... 92 7e-24
BG452187 89 1e-18
AL371988 similar to PIR|T47571|T475 hypothetical protein F24B22.... 82 3e-16
BE320107 weakly similar to PIR|T47571|T475 hypothetical protein ... 74 5e-14
AW257301 similar to GP|16323069|gb| AT5g41600/MBK23_13 {Arabidop... 54 6e-08
AL371989 weakly similar to GP|9279659|dbj| seed maturation prote... 50 8e-07
AJ502352 31 0.51
TC84376 30 1.1
TC91338 29 1.9
AW980465 PIR|A26014|A26 histone H3 - wheat, partial (98%) 28 2.5
BQ140761 weakly similar to GP|18076245|emb phosphophoryn {Rattus... 28 3.3
BQ151328 similar to PIR|G86292|G86 hypothetical protein AAF82153... 28 3.3
BF649857 27 7.4
>TC88297 similar to GP|15215586|gb|AAK91338.1 AT4g23630/F9D16_100
{Arabidopsis thaliana}, partial (75%)
Length = 1243
Score = 281 bits (719), Expect = 2e-76
Identities = 138/237 (58%), Positives = 177/237 (74%), Gaps = 1/237 (0%)
Frame = +3
Query: 7 ESFDDKIEDKFH-GSDSDSFSDSEDDHNINNRPPIPLAKSNTVYRLFGRDRPVHKVLGGG 65
+S +KI +K H DS S SDS ++ +P I K V+RLFGR++PVH VLGGG
Sbjct: 135 DSLLEKITEKLHFEDDSSSDSDSHSHSELDKKPVIESVKEK-VFRLFGREKPVHSVLGGG 311
Query: 66 KPADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNASVFI 125
KPAD+ LW+NKK +A L TA+WVFFEL++Y+ +TL+CH+ IL LA LFLWSNA F+
Sbjct: 312 KPADVFLWKNKKLSAGTLATATAIWVFFELLEYHFLTLICHISILVLALLFLWSNAHTFV 491
Query: 126 HKSPLQIPHVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVI 185
HK+P IP V +P+E + AS LRIEIN+ F+ LR+IGTG+D+KKFL VI GLW +S++
Sbjct: 492 HKTPPHIPIVHLPEEPFLQIASALRIEINRGFSALRDIGTGKDVKKFLIVIFGLWILSIV 671
Query: 186 GSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKVLSQI 242
GS NFLTLFYI +V LFT+P VY+K ED++D LAEKA IEIKKQYAV D KVLS++
Sbjct: 672 GSWSNFLTLFYITFVLLFTVPFVYDKYEDKIDPLAEKAFIEIKKQYAVFDEKVLSKV 842
>TC90320 similar to GP|17381218|gb|AAL36421.1 unknown protein {Arabidopsis
thaliana}, partial (81%)
Length = 911
Score = 154 bits (388), Expect(2) = 2e-66
Identities = 77/135 (57%), Positives = 94/135 (69%), Gaps = 1/135 (0%)
Frame = +3
Query: 4 VDTESFDDKIEDKFHGS-DSDSFSDSEDDHNINNRPPIPLAKSNTVYRLFGRDRPVHKVL 62
V +ES DKI K H DS S S D+ + P + N V+RLFGR++P+H VL
Sbjct: 162 VKSESLLDKISGKIHDHHDSSSSSSDSDNDKKEKKVSSPTSLKNKVFRLFGREKPLHNVL 341
Query: 63 GGGKPADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNAS 122
GGGKPAD+ LWRNKK +A LG TALWV FEL++Y+L+TL+ HL ILALA LFLWSNAS
Sbjct: 342 GGGKPADVFLWRNKKISATTLGVATALWVLFELLEYHLLTLISHLAILALAVLFLWSNAS 521
Query: 123 VFIHKSPLQIPHVVI 137
FI+KSP +IP V I
Sbjct: 522 TFINKSPPKIPQVHI 566
Score = 116 bits (290), Expect(2) = 2e-66
Identities = 58/107 (54%), Positives = 81/107 (75%), Gaps = 1/107 (0%)
Frame = +1
Query: 138 PQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVIGSCFNFLTLFYI 197
P+E V + AS +RIEIN+ FAILR+I +GRD+K+FL+VIAGLW +S++GS NFLTLFYI
Sbjct: 568 PEEPVLQIASAIRIEINRAFAILRDIASGRDLKQFLSVIAGLWVLSIVGSWTNFLTLFYI 747
Query: 198 FYVSLFTLPLVYEKNEDQVDALAEKAMIEIKK-QYAVLDAKVLSQIP 243
+ L T+P++YEK ED+VD+ EKA+ +++ D VLS+IP
Sbjct: 748 AFXLLHTVPVLYEKYEDRVDSFGEKALS*VQRSSMRGFDEXVLSKIP 888
>TC86997 similar to GP|6056199|gb|AAF02816.1| unknown protein {Arabidopsis
thaliana}, partial (67%)
Length = 1010
Score = 186 bits (472), Expect = 7e-48
Identities = 91/195 (46%), Positives = 129/195 (65%)
Frame = +3
Query: 45 SNTVYRLFGRDRPVHKVLGGGKPADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLV 104
S+ + RLFGR++PVH +LGGGK AD+LLWRNK +A L T +WV FE + YN I+L+
Sbjct: 174 SSEINRLFGREKPVHHLLGGGKSADVLLWRNKTISASFLTGATIVWVLFEWLNYNFISLL 353
Query: 105 CHLMILALAALFLWSNASVFIHKSPLQIPHVVIPQECVFEAASVLRIEINQVFAILREIG 164
C +++L L A FLWSNAS F++ +P Q+P V+P+E A+V+ E+N+ L+++
Sbjct: 354 CFVLVLGLLAQFLWSNASGFLNSTPSQVPRFVVPEELFVNIATVIGNEVNRGLRFLQDVS 533
Query: 165 TGRDIKKFLTVIAGLWFISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAM 224
++K FL V+ LW SVIGS NFLT+ YI V+ TLP++YE+ ED+VD K
Sbjct: 534 CEGNLKTFLIVVVSLWAGSVIGSWCNFLTVIYIGIVAAHTLPVLYERYEDEVDNFVLKVF 713
Query: 225 IEIKKQYAVLDAKVL 239
+I+ Y LDA VL
Sbjct: 714 GQIQNNYRKLDAGVL 758
>TC88294 weakly similar to GP|15215586|gb|AAK91338.1 AT4g23630/F9D16_100
{Arabidopsis thaliana}, partial (18%)
Length = 845
Score = 150 bits (378), Expect = 6e-37
Identities = 85/227 (37%), Positives = 134/227 (58%), Gaps = 2/227 (0%)
Frame = +2
Query: 19 GSDSDSFSDSED--DHNINNRPPIPLAKSNTVYRLFGRDRPVHKVLGGGKPADILLWRNK 76
G++ D+F+ S+ + N N+ + +SN LFGR P+H VLGGGK ADILLW++
Sbjct: 11 GTN*DNFNSSKKKKEDNFNH*SIVMPIRSNQP-GLFGR--PLHAVLGGGKIADILLWKDM 181
Query: 77 KCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNASVFIHKSPLQIPHVV 136
K +A + + +W FE+++YNL+TL+CH++I + LF+W NA+ I I
Sbjct: 182 KSSAAIVAGFSMVWFLFEVVEYNLVTLLCHILIALMLILFIWYNAAGLITWKLPDIYDFE 361
Query: 137 IPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVIGSCFNFLTLFY 196
IP V + + N +I TG+D++ F IAGLW +S IG+ F+ + L Y
Sbjct: 362 IPDSTV----RFIHKKFNLFLRKFYDISTGKDLRFFFVTIAGLWIMSTIGNFFSTVNLLY 529
Query: 197 IFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKVLSQIP 243
++ L TLP++YE+ E +VD LA K ++K+ + LD+ VL++IP
Sbjct: 530 TTFLCLVTLPIMYERYEHEVDYLASKGNQDVKRLFNKLDSTVLNRIP 670
>TC81628 weakly similar to GP|12957724|gb|AAK09242.1 unknown protein {Oryza
sativa (japonica cultivar-group)}, partial (67%)
Length = 704
Score = 141 bits (355), Expect = 3e-34
Identities = 66/163 (40%), Positives = 107/163 (65%), Gaps = 2/163 (1%)
Frame = +1
Query: 50 RLFGRDRPVHKVLGGGKPADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMI 109
+LFGR++PVH +LGGGK AD+LLWR+KK +A L A T +W+ FE + YN ++L+C ++
Sbjct: 199 KLFGREKPVHHILGGGKSADVLLWRDKKISAAVLTAATTIWLLFEWLNYNFLSLLCLALV 378
Query: 110 LALAALFLWSNAS-VF-IHKSPLQIPHVVIPQECVFEAASVLRIEINQVFAILREIGTGR 167
L ++ FLW+NAS VF + P + P +V+P++ A+ + E+N+ L+ + G
Sbjct: 379 LVMSVQFLWTNASGVFSSDRKPSKAPRLVLPKDFFVNIATAVGAEVNRGLRFLQNVSCGX 558
Query: 168 DIKKFLTVIAGLWFISVIGSCFNFLTLFYIFYVSLFTLPLVYE 210
++K+F + LW +VIG+ FNF T YI + + TLP++Y+
Sbjct: 559 NLKQFXIGVVCLWAGAVIGNWFNFFTXXYIGFGAAHTLPVLYK 687
>TC89658 weakly similar to PIR|A84527|A84527 hypothetical protein At2g15280
[imported] - Arabidopsis thaliana, partial (59%)
Length = 1217
Score = 115 bits (287), Expect = 2e-26
Identities = 66/206 (32%), Positives = 108/206 (52%)
Frame = +1
Query: 38 PPIPLAKSNTVYRLFGRDRPVHKVLGGGKPADILLWRNKKCTAIALGAGTALWVFFELMQ 97
PPI + + + VH LG G AD+LLW+N + L + T LW FE
Sbjct: 100 PPIAMVEPQRI--------SVHHALGSGLVADVLLWKNWRGGVTLLISATTLWYLFERAG 255
Query: 98 YNLITLVCHLMILALAALFLWSNASVFIHKSPLQIPHVVIPQECVFEAASVLRIEINQVF 157
YN ++ V ++++L + LFLW+ A+ +++ +P + I ++ + + + VL+I +N+
Sbjct: 256 YNFLSFVANVILLLVVILFLWAKAANLLNRPLPPLPDLEISEKTIAKLSDVLQIWLNRAL 435
Query: 158 AILREIGTGRDIKKFLTVIAGLWFISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVD 217
++ +I R++ V+ L IS IGS FNFLTL YI + +LP+ Y+K +D +D
Sbjct: 436 SVAHDIAINRNLFLCAEVVGVLLTISYIGSLFNFLTLIYICVLLSLSLPVAYDKYQDLID 615
Query: 218 ALAEKAMIEIKKQYAVLDAKVLSQIP 243
I QY + VLS IP
Sbjct: 616 EKIHVVHGIIHPQYQKIRRIVLSIIP 693
>TC88528 weakly similar to GP|4102690|gb|AAD01540.1| 24 kDa seed maturation
protein {Glycine max}, partial (88%)
Length = 796
Score = 110 bits (276), Expect = 4e-25
Identities = 58/173 (33%), Positives = 93/173 (53%)
Frame = +1
Query: 69 DILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNASVFIHKS 128
DI+LWR KK +AI L T+ W+ E+ Q+N +TL+ L I + ++FL+SN F K
Sbjct: 109 DIVLWRRKKLSAIVLIVATSSWMLLEVCQFNFLTLISWLAIFVVTSIFLYSNMLTFFGKE 288
Query: 129 PLQIPHVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVIGSC 188
P + + + +E A +R I + L + D F+ V+ L IS +G+C
Sbjct: 289 PPNLLRLELKEETATRMAKTVRAWIEKSIRWLFVVSIKEDWPVFVGVMVRLLAISYVGTC 468
Query: 189 FNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKVLSQ 241
+FLT YI ++ TLP+ Y KNED++ E + KK Y ++D K +++
Sbjct: 469 MDFLTFIYIGILTGMTLPITYMKNEDKIKRCMEWLREKYKKSYEIIDEKAINK 627
>BQ149587 similar to GP|6056199|gb| unknown protein {Arabidopsis thaliana},
partial (41%)
Length = 600
Score = 91.7 bits (226), Expect(2) = 7e-24
Identities = 41/88 (46%), Positives = 59/88 (66%)
Frame = +3
Query: 68 ADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNASVFIHK 127
AD+LLWRNK +A L T +WV FE + YN I+L+C +++L L A FLWSNAS F++
Sbjct: 321 ADVLLWRNKTISASFLTGATIVWVLFEWLNYNFISLLCFVLVLGLLAQFLWSNASGFLNS 500
Query: 128 SPLQIPHVVIPQECVFEAASVLRIEINQ 155
+P Q+P V+P E A+V+ E+N+
Sbjct: 501 TPSQVPRFVVPXELFVNIATVIGNEVNR 584
Score = 35.8 bits (81), Expect(2) = 7e-24
Identities = 14/22 (63%), Positives = 19/22 (85%)
Frame = +1
Query: 45 SNTVYRLFGRDRPVHKVLGGGK 66
S+ + RLFGR++PVH +LGGGK
Sbjct: 145 SSEINRLFGREKPVHHLLGGGK 210
>BG452187
Length = 619
Score = 89.4 bits (220), Expect = 1e-18
Identities = 44/130 (33%), Positives = 71/130 (53%)
Frame = +2
Query: 68 ADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNASVFIHK 127
+D+LLW+ + + + T W+ E +T+ +++L + LFL SN + +K
Sbjct: 179 SDVLLWKRWQVSFGVIVVSTVAWLLLEWTDLPFLTICSDVLLLLIVLLFLNSNYAALRNK 358
Query: 128 SPLQIPHVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVIGS 187
P +P +V+ +E V A+ R++IN V I +I G+D F V+ LW +SVIGS
Sbjct: 359 QPPTLPELVVSEEMVNNVAASFRVKINNVLLIAHDITVGKDFXIFFKVVVCLWLLSVIGS 538
Query: 188 CFNFLTLFYI 197
F+F TL YI
Sbjct: 539 IFSFFTLAYI 568
>AL371988 similar to PIR|T47571|T475 hypothetical protein F24B22.80 -
Arabidopsis thaliana, partial (10%)
Length = 331
Score = 81.6 bits (200), Expect = 3e-16
Identities = 36/98 (36%), Positives = 64/98 (64%)
Frame = +1
Query: 22 SDSFSDSEDDHNINNRPPIPLAKSNTVYRLFGRDRPVHKVLGGGKPADILLWRNKKCTAI 81
+++ SDS+D+ + KS + +F ++P+H++LGGGK AD+LLWR++ +A
Sbjct: 10 NNTSSDSDDE----------IGKSRAM--IFPHEKPLHEILGGGKVADVLLWRDRNVSAA 153
Query: 82 ALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWS 119
L T +W FE+++YN++TL+CH+ I + ++LWS
Sbjct: 154 FLLGITLIWFLFEVVEYNIVTLLCHISITTMLVIYLWS 267
>BE320107 weakly similar to PIR|T47571|T475 hypothetical protein F24B22.80 -
Arabidopsis thaliana, partial (42%)
Length = 372
Score = 73.9 bits (180), Expect = 5e-14
Identities = 35/106 (33%), Positives = 62/106 (58%)
Frame = +2
Query: 44 KSNTVYRLFGRDRPVHKVLGGGKPADILLWRNKKCTAIALGAGTALWVFFELMQYNLITL 103
K + RLF R R +H++LG G+ AD++LWR K T + L A +V FE Y L++L
Sbjct: 53 KMGSSQRLFNRKRTLHEILGSGQVADLILWRRKNQTVMILLVTLAAFVVFERSGYTLLSL 232
Query: 104 VCHLMILALAALFLWSNASVFIHKSPLQIPHVVIPQECVFEAASVL 149
V ++++L + LFLW+ ++ +++ +P + + E E A+ +
Sbjct: 233 VSNVLLLLVVILFLWAKSAAILNRPAPPLPQLHLSDEMTNEMAAFI 370
>AW257301 similar to GP|16323069|gb| AT5g41600/MBK23_13 {Arabidopsis
thaliana}, partial (16%)
Length = 132
Score = 53.9 bits (128), Expect = 6e-08
Identities = 25/42 (59%), Positives = 33/42 (78%)
Frame = -2
Query: 199 YVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKVLS 240
+V L T+P++YEK ED+VD+ EKA+ E KKQY+V D KVLS
Sbjct: 128 FVLLHTVPVLYEKYEDRVDSFGEKALHEFKKQYSVFDEKVLS 3
>AL371989 weakly similar to GP|9279659|dbj| seed maturation protein-like
{Arabidopsis thaliana}, partial (19%)
Length = 400
Score = 50.1 bits (118), Expect = 8e-07
Identities = 20/58 (34%), Positives = 40/58 (68%)
Frame = +3
Query: 186 GSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKVLSQIP 243
G+ FNF+ L YI ++ L TLP+VY++ E++++ LA +++++++Y L++IP
Sbjct: 3 GTYFNFINLLYIGFLCLQTLPIVYDRYEEEINNLAGHVIVDLRRKYRRFKKSYLNKIP 176
>AJ502352
Length = 627
Score = 30.8 bits (68), Expect = 0.51
Identities = 12/26 (46%), Positives = 17/26 (65%)
Frame = +3
Query: 188 CFNFLTLFYIFYVSLFTLPLVYEKNE 213
C NFL +F +SLF LPL++ +E
Sbjct: 345 CLNFLLVFLCILISLFLLPLIFLNHE 422
>TC84376
Length = 672
Score = 29.6 bits (65), Expect = 1.1
Identities = 21/79 (26%), Positives = 40/79 (50%), Gaps = 2/79 (2%)
Frame = +1
Query: 64 GGKPADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCH--LMILALAALFLWSNA 121
G K + LW N C + + L++F+ ++ + ++C+ + IL +LF WSNA
Sbjct: 271 GNKKLKVSLWPNH-CLILVV-----LFLFYAILVFWSWLVLCYSCISILFAQSLFEWSNA 432
Query: 122 SVFIHKSPLQIPHVVIPQE 140
+H ++ H+V +E
Sbjct: 433 FYKVHWE--RVHHIVANKE 483
>TC91338
Length = 744
Score = 28.9 bits (63), Expect = 1.9
Identities = 19/62 (30%), Positives = 25/62 (39%)
Frame = +1
Query: 20 SDSDSFSDSEDDHNINNRPPIPLAKSNTVYRLFGRDRPVHKVLGGGKPADILLWRNKKCT 79
S SF+D +I N SN Y +G +RP H G G P + NK T
Sbjct: 142 SQPPSFNDYSTCKDIKNSYSCGSIISNISYPFWGENRPFH--CGAGNPYHLYCHNNKDTT 315
Query: 80 AI 81
+
Sbjct: 316 IL 321
>AW980465 PIR|A26014|A26 histone H3 - wheat, partial (98%)
Length = 607
Score = 28.5 bits (62), Expect = 2.5
Identities = 18/49 (36%), Positives = 31/49 (62%), Gaps = 2/49 (4%)
Frame = -2
Query: 166 GRDIKKFLTV-IAGLWFIS-VIGSCFNFLTLFYIFYVSLFTLPLVYEKN 212
GR++ F T +AG F++ ++ SCF L L +IF + ++ +PL E+N
Sbjct: 159 GRNLCGFFTPPVAGADFLAALVASCFPRLFLRWIFVLYVWYVPLGTERN 13
>BQ140761 weakly similar to GP|18076245|emb phosphophoryn {Rattus
norvegicus}, partial (9%)
Length = 620
Score = 28.1 bits (61), Expect = 3.3
Identities = 13/39 (33%), Positives = 19/39 (48%)
Frame = +1
Query: 6 TESFDDKIEDKFHGSDSDSFSDSEDDHNINNRPPIPLAK 44
++S D D DSD SDS D + ++ P+P K
Sbjct: 271 SDSSDGSDSDSDASDDSDDSSDSSSDDSSSDEEPVPAPK 387
>BQ151328 similar to PIR|G86292|G86 hypothetical protein AAF82153.1 [imported]
- Arabidopsis thaliana, partial (3%)
Length = 1425
Score = 28.1 bits (61), Expect = 3.3
Identities = 11/28 (39%), Positives = 16/28 (56%)
Frame = +1
Query: 12 KIEDKFHGSDSDSFSDSEDDHNINNRPP 39
+IE + H S F++ + HN NN PP
Sbjct: 1297 EIEREHHISQIPQFTEQQQSHNTNNPPP 1380
>BF649857
Length = 601
Score = 26.9 bits (58), Expect = 7.4
Identities = 9/22 (40%), Positives = 16/22 (71%)
Frame = -2
Query: 181 FISVIGSCFNFLTLFYIFYVSL 202
+I ++G ++F+T+F FY SL
Sbjct: 384 YIKILGFYYHFITIFMAFYASL 319
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.141 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,001,479
Number of Sequences: 36976
Number of extensions: 115744
Number of successful extensions: 1164
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 1146
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1156
length of query: 250
length of database: 9,014,727
effective HSP length: 94
effective length of query: 156
effective length of database: 5,538,983
effective search space: 864081348
effective search space used: 864081348
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC148655.10


