

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148652.15 + phase: 0 /pseudo
(121 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
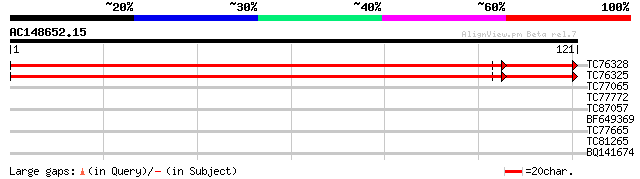
TC76328 homologue to PIR|C96814|C96814 hypothetical protein T30F... 150 2e-39
TC76325 similar to PIR|C96814|C96814 hypothetical protein T30F21... 149 7e-39
TC77065 similar to PIR|A85356|A85356 nucleotide sugar epimerase-... 35 0.005
TC77772 similar to GP|22136300|gb|AAM91228.1 unknown protein {Ar... 30 0.22
TC87057 similar to PIR|T48135|T48135 nucleotide sugar epimerase-... 30 0.22
BF649369 27 1.9
TC77665 similar to SP|Q9SNY3|GMD1_ARATH GDP-mannose 4 6 dehydrat... 26 3.2
TC81265 weakly similar to PIR|T45969|T45969 mRNA capping enzyme-... 26 4.2
BQ141674 26 4.2
>TC76328 homologue to PIR|C96814|C96814 hypothetical protein T30F21.10
[imported] - Arabidopsis thaliana, partial (56%)
Length = 1492
Score = 150 bits (378), Expect(2) = 2e-39
Identities = 75/106 (70%), Positives = 83/106 (77%)
Frame = +2
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
MHFAAQTHVDNS+ NSFEFT+ + Q+K FIHVSTD+ YGETDE+AV
Sbjct: 362 MHFAAQTHVDNSFGNSFEFTKNNIYGTHVLLEACKVTGQIKRFIHVSTDEVYGETDEDAV 541
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
VGNHEASQLL TNPYSA K GAEMLVM+YGRSYGLPVITTRG+ VY
Sbjct: 542 VGNHEASQLLPTNPYSATKAGAEMLVMAYGRSYGLPVITTRGNNVY 679
Score = 27.7 bits (60), Expect(2) = 2e-39
Identities = 10/18 (55%), Positives = 12/18 (66%)
Frame = +1
Query: 104 RVYPFSHERKESIDSWRW 121
+VYP + RK S DSW W
Sbjct: 712 KVYPLGNARKSSSDSW*W 765
>TC76325 similar to PIR|C96814|C96814 hypothetical protein T30F21.10
[imported] - Arabidopsis thaliana, partial (97%)
Length = 2416
Score = 149 bits (375), Expect(2) = 7e-39
Identities = 74/106 (69%), Positives = 83/106 (77%)
Frame = +1
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
MHFAAQTHVDNS+ NSFEFT+ + Q++ FIHVSTD+ YGETDE+AV
Sbjct: 274 MHFAAQTHVDNSFGNSFEFTKNNIYGTHVLLEACKVTGQIRRFIHVSTDEVYGETDEDAV 453
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
VGNHEASQLL TNPYSA K GAEMLVM+YGRSYGLPVITTRG+ VY
Sbjct: 454 VGNHEASQLLPTNPYSATKAGAEMLVMAYGRSYGLPVITTRGNNVY 591
Score = 26.9 bits (58), Expect(2) = 7e-39
Identities = 9/18 (50%), Positives = 13/18 (72%)
Frame = +3
Query: 104 RVYPFSHERKESIDSWRW 121
+V+P + RK+S DSW W
Sbjct: 624 KVHPSGYARKDSSDSWGW 677
>TC77065 similar to PIR|A85356|A85356 nucleotide sugar epimerase-like
protein [imported] - Arabidopsis thaliana, partial (97%)
Length = 1679
Score = 35.4 bits (80), Expect = 0.005
Identities = 25/106 (23%), Positives = 42/106 (39%)
Frame = +2
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
MH AAQ V + EN + ++ L++ + N + S+ YG ++
Sbjct: 602 MHLAAQAGVRYAMENPMSYVNSNIAGLVTLLEACKTANPQPSIVWASSSSVYGLNEKVPF 781
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
+ Q + Y+A K E + +Y YGL + R VY
Sbjct: 782 SESDRTDQ--PASLYAATKKAGEEITHTYNHIYGLSITGLRFFTVY 913
>TC77772 similar to GP|22136300|gb|AAM91228.1 unknown protein {Arabidopsis
thaliana}, partial (62%)
Length = 2554
Score = 30.0 bits (66), Expect = 0.22
Identities = 25/109 (22%), Positives = 45/109 (40%), Gaps = 9/109 (8%)
Frame = +2
Query: 2 HFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAVV 61
HF + + + + F+ +E + F +S++ H+S T +
Sbjct: 104 HFTPLSQAPHQWRHKCRFS----IERVLFSQNPISQSNQSQHSHISIKNPNL*TPHHHHQ 271
Query: 62 GNHEASQLLETNPYSAMKIGAEM---------LVMSYGRSYGLPVITTR 101
NH + Q+ +P+S KIGA LV+S+ + P+I TR
Sbjct: 272 QNHHSHQIQRQDPHSTTKIGATQFQKLHL*PNLVISHALQFLKPMIPTR 418
>TC87057 similar to PIR|T48135|T48135 nucleotide sugar epimerase-like
protein - Arabidopsis thaliana, partial (80%)
Length = 1655
Score = 30.0 bits (66), Expect = 0.22
Identities = 24/106 (22%), Positives = 37/106 (34%)
Frame = +2
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
MH AAQ V + +N + ++ S N + S+ YG +
Sbjct: 551 MHLAAQAGVRYAMQNPNSYVHSNLAGFTVLLEACKSANPQPAIVWASSSSVYGLNSKVPF 730
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
Q + Y+A K E + +Y YGL + R VY
Sbjct: 731 SEKDRTDQ--PASLYAATKKAGEGIAHTYNHIYGLSITALRFFTVY 862
>BF649369
Length = 631
Score = 26.9 bits (58), Expect = 1.9
Identities = 19/55 (34%), Positives = 25/55 (44%)
Frame = +2
Query: 63 NHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVYPFSHERKESID 117
+H LL YS G E++ YGRSY V T G+R E +E +D
Sbjct: 167 DHTC*DLLRCAEYSR*HEG-EIIPFEYGRSYDPLVQLTNGNRGRSIQGEIEEGVD 328
>TC77665 similar to SP|Q9SNY3|GMD1_ARATH GDP-mannose 4 6 dehydratase (EC
4.2.1.47) (GDP-D-mannose dehydratase) (GMD). [Mouse-ear
cress], partial (94%)
Length = 1538
Score = 26.2 bits (56), Expect = 3.2
Identities = 21/101 (20%), Positives = 44/101 (42%), Gaps = 7/101 (6%)
Frame = +1
Query: 2 HFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKG-------FIHVSTDKFYGE 54
+ AAQ+HV S+E ++T + + R L+ ++ + + + + +G
Sbjct: 472 NLAAQSHVAVSFEIP-DYT-ADVVATGALRLLEAVRSHIDATGRSHIRYYQAGSSEMFGS 645
Query: 55 TDENAVVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGL 95
T E + +PY+A K+ A ++Y +YG+
Sbjct: 646 TPPP----QSETTPFHPRSPYAASKVAAHWYTVNYREAYGI 756
>TC81265 weakly similar to PIR|T45969|T45969 mRNA capping enzyme-like
protein - Arabidopsis thaliana, partial (13%)
Length = 581
Score = 25.8 bits (55), Expect = 4.2
Identities = 13/27 (48%), Positives = 14/27 (51%)
Frame = +2
Query: 90 GRSYGLPVITTRGDRVYPFSHERKESI 116
GR GL + T R YP S RKE I
Sbjct: 431 GRELGLVIDLTNTTRYYPLSDWRKERI 511
>BQ141674
Length = 1043
Score = 25.8 bits (55), Expect = 4.2
Identities = 12/25 (48%), Positives = 16/25 (64%), Gaps = 1/25 (4%)
Frame = -1
Query: 8 HVDNSYENSFEFTQTSF-MELMSFR 31
H+DN+Y +S F +SF M LM R
Sbjct: 257 HIDNAYASSISFAYSSFHMSLMVVR 183
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.130 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,079,699
Number of Sequences: 36976
Number of extensions: 28052
Number of successful extensions: 143
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 141
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 141
length of query: 121
length of database: 9,014,727
effective HSP length: 84
effective length of query: 37
effective length of database: 5,908,743
effective search space: 218623491
effective search space used: 218623491
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 52 (24.6 bits)
Medicago: description of AC148652.15


