

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148652.13 - phase: 0 /pseudo
(395 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
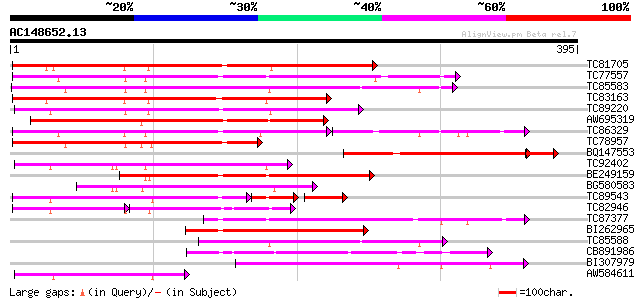
Score E
Sequences producing significant alignments: (bits) Value
TC81705 weakly similar to GP|6721520|dbj|BAA89562.1 EST AU029428... 258 2e-69
TC77557 similar to PIR|JC7519|JC7519 subtilisin-like serine prot... 228 3e-60
TC85583 similar to PIR|T51335|T51335 subtilisin-like proteinase ... 222 2e-58
TC83163 weakly similar to GP|6721520|dbj|BAA89562.1 EST AU029428... 221 3e-58
TC89220 similar to PIR|T05768|T05768 subtilisin-like proteinase ... 203 8e-53
AW695319 similar to GP|6708179|gb| subtilisin-type serine endope... 190 9e-49
TC86329 similar to PIR|T07171|T07171 subtilisin-like proteinase ... 169 1e-48
TC78957 similar to GP|14091078|gb|AAK53589.1 subtilisin-like pro... 181 3e-46
BQ147553 similar to GP|9759240|dbj| serine protease-like protein... 155 2e-43
TC92402 similar to GP|20198252|gb|AAM15483.1 subtilisin-like ser... 164 7e-41
BE249159 weakly similar to GP|20198252|gb subtilisin-like serine... 153 1e-37
BG580583 weakly similar to GP|9757901|dbj serine protease-like p... 148 4e-36
TC89543 similar to PIR|T06580|T06580 subtilisin-like proteinase ... 123 7e-35
TC82946 weakly similar to GP|20160863|dbj|BAB89802. putative sub... 91 6e-31
TC87377 weakly similar to PIR|T07171|T07171 subtilisin-like prot... 113 1e-25
BI262965 similar to PIR|A84473|A84 probable serine proteinase [i... 112 2e-25
TC85588 similar to GP|20198169|gb|AAM15440.1 subtilisin-like ser... 109 2e-24
CB891986 similar to PIR|S52769|S52 subtilisin-like proteinase ag... 107 8e-24
BI307979 weakly similar to GP|20521377|db putative subtilisin-li... 102 2e-22
AW584611 similar to PIR|T07184|T071 subtilisin-like proteinase (... 88 5e-18
>TC81705 weakly similar to GP|6721520|dbj|BAA89562.1 EST AU029428(E30359)
corresponds to a region of the predicted gene.~Similar
to antifreeze-like, partial (33%)
Length = 1443
Score = 258 bits (660), Expect = 2e-69
Identities = 136/278 (48%), Positives = 179/278 (63%), Gaps = 24/278 (8%)
Frame = +1
Query: 3 GVISVFPSEEFHIQTTRSWDFL----GLRQS--IKRDQIIETDLVIGVIDIRIWPESESF 56
GV+SVFP + TTRSWDFL G++ S + Q D++I +ID IWPES SF
Sbjct: 283 GVVSVFPDPILELHTTRSWDFLDSDLGMKPSTNVLTHQHSSNDIIIALIDTGIWPESPSF 462
Query: 57 NDKGLGPIPKKWRGVCAGGGNF---SCNNKIIGARFYGDGDV-------------SARYS 100
D+G+G IP W+G+C G +F +CN K+IGAR+Y D S R +
Sbjct: 463 TDEGIGKIPSVWKGICMEGHDFKKSNCNRKLIGARYYNTQDTFGSNKTHIGGAKGSPRDT 642
Query: 101 FGHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAA 160
GH THTASTA G V + ++YG AKGTARGG PS+RIAAYK C + CSG IL A
Sbjct: 643 VGHGTHTASTAAGVNVNNANYYGLAKGTARGGSPSTRIAAYKTCSEEG---CSGSTILKA 813
Query: 161 FDDAISDGVDVITVSLGPEHA--SDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVC 218
DDAI DGVD+I++S+G SD+LNDPIAIG+FHA ++G+ +AGN GP P++V
Sbjct: 814 MDDAIKDGVDIISISIGLSSLMQSDYLNDPIAIGAFHAEQRGVTAVCSAGNDGPDPNTVV 993
Query: 219 SGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKSINIT 256
+ APW+ +VAA++IDR F ++LGNGK+F G IN +
Sbjct: 994 NTAPWIFTVAASNIDRNFQSTIVLGNGKSFQGAGINFS 1107
>TC77557 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase (EC
3.4.21.-) - Arabidopsis thaliana, partial (88%)
Length = 2682
Score = 228 bits (581), Expect = 3e-60
Identities = 135/333 (40%), Positives = 186/333 (55%), Gaps = 21/333 (6%)
Frame = +3
Query: 3 GVISVFPSEEFHIQTTRSWDFLGLRQSIKR--DQIIETDLVIGVIDIRIWPESESFNDKG 60
G+++V P ++ + TTR+ FLGL +S + ++V+GV+D +WPES+SFND G
Sbjct: 513 GILAVLPEVKYELHTTRTPQFLGLDKSADMFPESSSGNEVVVGVLDTGVWPESKSFNDAG 692
Query: 61 LGPIPKKWRGVCAGGGNFS---CNNKIIGARFYGDG-------------DVSARYSFGHR 104
GPIP W+G C G NF+ CN K+IGARF+ G S R GH
Sbjct: 693 FGPIPTTWKGACESGTNFTAANCNKKLIGARFFSKGVEAMLGPIDETTESKSPRDDDGHG 872
Query: 105 THTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDA 164
THT+STA G V D S +G+A GTARG +R+A YK+C G C ILAA D A
Sbjct: 873 THTSSTAAGSVVPDASLFGYASGTARGMATRARVAVYKVCW---KGGCFSSDILAAIDKA 1043
Query: 165 ISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWL 224
ISD V+V+++SLG SD+ D +AIG+F AMEKGIL + +AGN GP S+ + APW+
Sbjct: 1044ISDNVNVLSLSLGGG-MSDYFRDSVAIGAFSAMEKGILVSCSAGNAGPSAYSLSNVAPWI 1220
Query: 225 VSVAATSIDRQFIDKVILGNGKTFVGKSI---NITPSNGKKIPIAVRNAQACLAGGNASP 281
+V A ++DR F V LGNG + G S+ N P + P+ + A N +
Sbjct: 1221TTVGAGTLDRDFPASVSLGNGLNYSGVSLYRGNALPES----PLPLIYAGNATNATNGNL 1388
Query: 282 EMCDCIDENMVKGKLVLCGSRNGKDLAYANGAI 314
M + +V GK+VLC G + GA+
Sbjct: 1389CMTGTLSPELVAGKIVLCD--RGMNARVQKGAV 1481
>TC85583 similar to PIR|T51335|T51335 subtilisin-like proteinase AIR3
auxin-induced [imported] - Arabidopsis thaliana
(fragment), partial (40%)
Length = 2691
Score = 222 bits (565), Expect = 2e-58
Identities = 133/339 (39%), Positives = 186/339 (54%), Gaps = 28/339 (8%)
Frame = +2
Query: 2 KGVISVFPSEEFHIQTTRSWDFLGLRQSIKRDQIIE----TDLVIGVIDIRIWPESESFN 57
+ V+SVF S+ + TTRSW+FLGLR++ K + + +I ID +WPES+SFN
Sbjct: 332 RNVVSVFLSKPHKLHTTRSWEFLGLRRNAKNTAWQKGKFGENTIIANIDTGVWPESKSFN 511
Query: 58 DKGLGPIPKKWRGVCAGG-GNFS------CNNKIIGARFYGDG-----------DVSARY 99
DKG GP+P KWRG A FS CN K+IGARF+ + +AR
Sbjct: 512 DKGYGPVPSKWRGGKACEISKFSKYKKNPCNRKLIGARFFSNAYEAYNDKLPSWQRTARD 691
Query: 100 SFGHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDI-DNGRCSGDAIL 158
GH THT STAGG V D S + GT +GG P +R+A YK+C + D C G +L
Sbjct: 692 FLGHGTHTLSTAGGNFVPDASVFAIGNGTVKGGSPRARVATYKVCWSLLDLEDCFGADVL 871
Query: 159 AAFDDAISDGVDVITVSLGPE---HASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPS 215
AA D AISDGVD+I++SL + D D ++IG+FHA+ + IL +AGN GP
Sbjct: 872 AAIDQAISDGVDIISLSLAGHSLVYPEDIFTDEVSIGAFHALSRNILLVASAGNEGPTGG 1051
Query: 216 SVCSGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKSINITPSNGKKIPIAVRNAQACLA 275
SV + APW+ ++AA+++DR F + +GN +T G S+ + + P+ V
Sbjct: 1052SVVNVAPWVFTIAASTLDRDFSSTITIGN-QTIRGASLFVNLPPNQAFPLIVSTDGKLAN 1228
Query: 276 GGNASPEMC--DCIDENMVKGKLVLCGSRNGKDLAYANG 312
N + C +D + VKGK+V C R G + A G
Sbjct: 1229ATNHDAQFCKPGTLDPSKVKGKIVEC-IREGNIKSVAEG 1342
>TC83163 weakly similar to GP|6721520|dbj|BAA89562.1 EST AU029428(E30359)
corresponds to a region of the predicted gene.~Similar
to antifreeze-like, partial (29%)
Length = 990
Score = 221 bits (564), Expect = 3e-58
Identities = 119/234 (50%), Positives = 153/234 (64%), Gaps = 12/234 (5%)
Frame = +2
Query: 3 GVISVFPSEEFHIQTTRSWDFL----GLR-QSIKRDQIIETDLVIGVIDIRIWPESESFN 57
GV+SVF + TTRSWDFL G+R I + Q D++IGVID IWPES SF
Sbjct: 293 GVVSVFEDPFLELHTTRSWDFLESDLGMRPHGILKHQHSSNDIIIGVIDTGIWPESPSFK 472
Query: 58 DKGLGPIPKKWRGVCAGGGNF---SCNNKIIGARFYGDGDV--SARYSFGHRTHTASTAG 112
D+G+G IP +W+GVC +F +CN K+IGAR+Y D S R GH THTASTA
Sbjct: 473 DEGIGKIPSRWKGVCMEAHDFKKSNCNRKLIGARYYNKKDPKGSPRDFNGHGTHTASTAA 652
Query: 113 GREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVI 172
G V + S+YG AKGTARGG PS+RIAAYK C CSG +L A DDAI DGVD+I
Sbjct: 653 GVIVNNASYYGLAKGTARGGSPSARIAAYKAC---SGEGCSGGTLLKAIDDAIKDGVDII 823
Query: 173 TVSLG--PEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWL 224
++S+G E S++L+DPIAIG+FHA ++G++ +AGN GP +V + PW+
Sbjct: 824 SISIGFSSEFLSEYLSDPIAIGAFHAEQRGVMVVCSAGNEGPDHYTVVNTTPWI 985
>TC89220 similar to PIR|T05768|T05768 subtilisin-like proteinase (EC
3.4.21.-) - Arabidopsis thaliana, partial (39%)
Length = 1149
Score = 203 bits (517), Expect = 8e-53
Identities = 113/269 (42%), Positives = 157/269 (58%), Gaps = 26/269 (9%)
Frame = +3
Query: 4 VISVFPSEEFHIQTTRSWDFLGLR--QSIKRDQIIETDLVIGVIDIRIWPESESFNDKGL 61
+++VF + TTRS FLGLR + + + +D+++GV D IWPE SF+D L
Sbjct: 315 ILAVFEDRRRQLHTTRSPQFLGLRNQRGLWSESDYGSDVIVGVFDTGIWPERRSFSDMNL 494
Query: 62 GPIPKKWRGVCAGGGNFS---CNNKIIGARFYGDGDV-------------------SARY 99
GPIP++W+GVC G FS CN K+IGAR++ G S R
Sbjct: 495 GPIPRRWKGVCESGEKFSPRNCNRKLIGARYFSKGHEVGAGSAGPLNPINETVEFRSPRD 674
Query: 100 SFGHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILA 159
+ GH THTASTA GR + G+A G A+G P +R+A YK+C N C ILA
Sbjct: 675 ADGHGTHTASTAAGRYAFQANMSGYASGIAKGVAPKARLAVYKVCWK--NSGCFDSDILA 848
Query: 160 AFDDAISDGVDVITVSLGPEH--ASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSV 217
AFD A++DGVDVI++S+G AS + DPIAIGS+ A+ +G+ + +AGN GP SV
Sbjct: 849 AFDAAVNDGVDVISISIGGGDGIASPYYLDPIAIGSYGAVSRGVFVSSSAGNDGPSGMSV 1028
Query: 218 CSGAPWLVSVAATSIDRQFIDKVILGNGK 246
+ APWL +V A +IDR F ++I+ + K
Sbjct: 1029TNLAPWLTTVGAGTIDRDFPSQIIIXDXK 1115
>AW695319 similar to GP|6708179|gb| subtilisin-type serine endopeptidase XSP1
{Arabidopsis thaliana}, partial (27%)
Length = 648
Score = 190 bits (482), Expect = 9e-49
Identities = 107/216 (49%), Positives = 137/216 (62%), Gaps = 8/216 (3%)
Frame = +2
Query: 15 IQTTRSWDFLGLRQSIKRDQIIETDLVIGVIDIRIWPESESFNDKGLGPIPKKWRGVCAG 74
+ TTRSWDF+GL + KR E D ++ ++D I PE +SF D G GP P KW+G C
Sbjct: 8 LHTTRSWDFIGLPLTAKRKLKSEGDTIVALLDTGITPEFQSFKDDGFGPPPAKWKGTCDK 187
Query: 75 GGNFS-CNNKIIGARFY------GDGDVSARYSF-GHRTHTASTAGGREVEDVSFYGFAK 126
NFS CNNKIIGA+++ D+ + GH THTASTA G V + S +G AK
Sbjct: 188 YVNFSGCNNKIIGAKYFKLDGRSNPSDILSPIDVEGHGTHTASTAAGNIVPNASLFGLAK 367
Query: 127 GTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHASDFLN 186
G ARG V S+R+A YKIC D C+ ILAAF+ AI DGVDVI+VSLG + ++
Sbjct: 368 GMARGAVHSARLAIYKICWTEDG--CADMDILAAFEAAIHDGVDVISVSLGGGN-ENYAQ 538
Query: 187 DPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAP 222
D IAIG+FHAM KGI+T +AGN GP ++V + AP
Sbjct: 539 DSIAIGAFHAMRKGIITVASAGNGGPTMATVVNNAP 646
>TC86329 similar to PIR|T07171|T07171 subtilisin-like proteinase (EC
3.4.21.-) 1 - tomato, partial (91%)
Length = 2671
Score = 169 bits (427), Expect(2) = 1e-48
Identities = 94/242 (38%), Positives = 137/242 (55%), Gaps = 20/242 (8%)
Frame = +2
Query: 3 GVISVFPSEEFHIQTTRSWDFLGLRQSIKR--DQIIETDLVIGVIDIRIWPESESFNDKG 60
G++SV P + + TTR+ +FLGL ++I ++++++GVID +WPE +SF+D G
Sbjct: 365 GILSVIPDVRYELHTTRTPEFLGLEKTITLLPSSGKQSEVIVGVIDTGVWPELKSFDDTG 544
Query: 61 LGPIPKKWRGVCAGGGNF---SCNNKIIGARFYGDG-------------DVSARYSFGHR 104
LGP+PK W+G C G F +CN K++GARF+ G S R GH
Sbjct: 545 LGPVPKSWKGECETGKTFNSSNCNKKLVGARFFAKGYEAAFGPIDENTESKSPRDDDGHG 724
Query: 105 THTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDA 164
+HT++TA G V S +GFA GTA+G +R+AAYK+C G C I AA D A
Sbjct: 725 SHTSTTAAGSAVAGASLFGFASGTAKGMATQARVAAYKVCW---LGGCFTSDIAAAIDKA 895
Query: 165 ISDGVDVIT--VSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAP 222
I VS +D+ D +A+G+F A+E GIL + +AGN GP +S+ + AP
Sbjct: 896 IDTQARTAAYKVSRLGGGLTDYYKDTVAMGTFAAIEHGILVSSSAGNGGPSKASLANVAP 1075
Query: 223 WL 224
W+
Sbjct: 1076WI 1081
Score = 42.4 bits (98), Expect(2) = 1e-48
Identities = 43/150 (28%), Positives = 68/150 (44%), Gaps = 13/150 (8%)
Frame = +3
Query: 226 SVAATSIDRQFIDKVILGNGKTFVGKSINITPSNGKKIPIA-VRNAQACLAGGNASPEMC 284
+V A +IDR F + LGNG + G S+ NGK P + + A ++S +C
Sbjct: 1086 TVGARTIDRDFPAYITLGNGNRYNGVSL----YNGKLPPNSPLPLVYAANVSQDSSDNLC 1253
Query: 285 --DCIDENMVKGKLVLCGSRNGKDLAYAN------GAIGSI----HNVTKSQLGASFVTP 332
D + + V GK+V+C R G A + G IG I + + + SF+ P
Sbjct: 1254 STDSLIPSKVSGKIVIC-DRGGNPRAEKSLVVKRAGGIGMILANNQDYGEELVADSFLLP 1430
Query: 333 RPSLNLKTNDFVHIQSYTNSSKYPVADFLF 362
+L K ++ I+ Y +S+ P A F
Sbjct: 1431 AAALGEKASN--EIKKYASSAPNPTAKIAF 1514
>TC78957 similar to GP|14091078|gb|AAK53589.1 subtilisin-like protein
{Glycine max}, partial (28%)
Length = 838
Score = 181 bits (460), Expect = 3e-46
Identities = 97/191 (50%), Positives = 125/191 (64%), Gaps = 17/191 (8%)
Frame = +1
Query: 3 GVISVFPSEEFHIQTTRSWDFLGLRQSIKRDQIIE--------TDLVIGVIDIRIWPESE 54
GV+SVFP + TT SWDFL L+ +K D + +D+VIG++D IWPE+
Sbjct: 274 GVVSVFPDPILKLHTTHSWDFLKLQTHVKIDSTLSNSSSQSSSSDIVIGMLDSGIWPEAT 453
Query: 55 SFNDKGLGPIPKKWRGVCAGGGNFS---CNNKIIGARFY----GDGDVSA--RYSFGHRT 105
SF+D G+ PIP W+G+C +F+ CN KIIGAR+Y GD V+A R + GH T
Sbjct: 454 SFSDNGMDPIPSGWKGICMTSNDFNSSNCNRKIIGARYYPNLEGDDRVAATTRDTVGHGT 633
Query: 106 HTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAI 165
HTASTA G V S+YG A+G A+GG P SR+A K+C +I CSG AILAAFDDAI
Sbjct: 634 HTASTAAGNAVSGASYYGLAEGIAKGGSPESRLAI*KVCSNIG---CSGSAILAAFDDAI 804
Query: 166 SDGVDVITVSL 176
SDGVDV+++SL
Sbjct: 805 SDGVDVLSLSL 837
>BQ147553 similar to GP|9759240|dbj| serine protease-like protein
{Arabidopsis thaliana}, partial (7%)
Length = 494
Score = 155 bits (391), Expect(2) = 2e-43
Identities = 79/131 (60%), Positives = 95/131 (72%)
Frame = +1
Query: 233 DRQFIDKVILGNGKTFVGKSINITPSNGKKIPIAVRNAQACLAGGNASPEMCDCIDENMV 292
DRQFIDK++LGNGKT +GKSIN PSNG K PI +C A GNAS EM DC+D+NMV
Sbjct: 1 DRQFIDKLVLGNGKTLIGKSINTFPSNGTKFPIVY----SCPARGNASHEMYDCMDKNMV 168
Query: 293 KGKLVLCGSRNGKDLAYANGAIGSIHNVTKSQLGASFVTPRPSLNLKTNDFVHIQSYTNS 352
GK+VLCG + A NGA GSI TK+ L A VTP+PS+ L +N+FVH+QSYTNS
Sbjct: 169 NGKIVLCGKGGDEIFADQNGAFGSIIKATKNNLDAPPVTPKPSIYLGSNEFVHVQSYTNS 348
Query: 353 SKYPVADFLFS 363
+KYPVA+ L S
Sbjct: 349 TKYPVAEILKS 381
Score = 38.9 bits (89), Expect(2) = 2e-43
Identities = 18/23 (78%), Positives = 18/23 (78%)
Frame = +3
Query: 360 FLFSWPKSFGPINYETRYKCPRS 382
FLFS PKS NYETRYKCPRS
Sbjct: 420 FLFSRPKSCHS*NYETRYKCPRS 488
>TC92402 similar to GP|20198252|gb|AAM15483.1 subtilisin-like serine
protease AIR3 {Arabidopsis thaliana}, partial (16%)
Length = 704
Score = 164 bits (414), Expect = 7e-41
Identities = 93/220 (42%), Positives = 123/220 (55%), Gaps = 26/220 (11%)
Frame = +1
Query: 4 VISVFPSEEFHIQTTRSWDFLGLR----QSIKRDQIIETDLVIGVIDIRIWPESESFNDK 59
V+SVF S+E + TTRSW+FLGL S + + +I ID +WPES SF+D+
Sbjct: 40 VVSVFLSKEHKLHTTRSWEFLGLHGNDINSAWQKGRFGENTIIANIDTGVWPESRSFSDR 219
Query: 60 GLGPIPKKWRG--VCA-----GGGNFSCNNKIIGARFYGDG-----------DVSARYSF 101
G+GPIP KWRG VC G CN K+IGARF+ D +AR
Sbjct: 220 GIGPIPAKWRGGNVCQINKLRGSKKVPCNRKLIGARFFSDAYERYNGKLPTSQRTARDFV 399
Query: 102 GHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDI-DNGRCSGDAILAA 160
GH THT STAGG V S + GT +GG P +R+A YK+C + D C G +L+A
Sbjct: 400 GHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDAASCFGADVLSA 579
Query: 161 FDDAISDGVDVITVSLG---PEHASDFLNDPIAIGSFHAM 197
D AI DGVD+I VS G ++ + D ++IG+FHA+
Sbjct: 580 IDQAIDDGVDIIFVSAGGPSSTNSEEIFTDEVSIGAFHAL 699
>BE249159 weakly similar to GP|20198252|gb subtilisin-like serine protease
AIR3 {Arabidopsis thaliana}, partial (24%)
Length = 661
Score = 153 bits (387), Expect = 1e-37
Identities = 89/189 (47%), Positives = 117/189 (61%), Gaps = 11/189 (5%)
Frame = +1
Query: 77 NFSCNNKIIGARFYGDG--------DVS---ARYSFGHRTHTASTAGGREVEDVSFYGFA 125
N S + K+IGAR + G D S AR + GH +HT STAGG V+ VS YG
Sbjct: 70 NCSTSRKLIGARAFYKGYEAYVGKLDASFYTARDTIGHGSHTLSTAGGNFVQGVSVYGNG 249
Query: 126 KGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHASDFL 185
GTA+GG P + +AAYK+C G CS +LA F+ AISDGVDV++VSLG + +
Sbjct: 250 NGTAKGGSPKAHVAAYKVCW---KGGCSDADVLAGFEAAISDGVDVLSVSLGMK-THNLF 417
Query: 186 NDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKVILGNG 245
D I+IGSFHA+ GI+ +AGN GP +V + APWL +VAA++IDR F V LG+
Sbjct: 418 TDSISIGSFHAVANGIVVVASAGNSGPYFGTVSNVAPWLFTVAASTIDRDFASYVTLGDN 597
Query: 246 KTFVGKSIN 254
K F G S++
Sbjct: 598 KHFKGTSLS 624
>BG580583 weakly similar to GP|9757901|dbj serine protease-like protein
{Arabidopsis thaliana}, partial (13%)
Length = 592
Score = 148 bits (373), Expect = 4e-36
Identities = 82/190 (43%), Positives = 109/190 (57%), Gaps = 22/190 (11%)
Frame = +3
Query: 47 IRIWPESESFNDKGLGPIPKKWRG--VCA-----GGGNFSCNNKIIGARFYG-------- 91
I +WPESESFND+G+GPIP +WRG +C CN K+IGARF+
Sbjct: 18 IGVWPESESFNDRGIGPIPLRWRGGNICQLDKLNTSKKVPCNRKLIGARFFNKAYEAFHG 197
Query: 92 ---DGDVSARYSFGHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDI- 147
+AR GH THT STAGG V++ + +G GT +GG P SR+A YK C +
Sbjct: 198 KLPSSQQTARDFVGHGTHTLSTAGGNFVQNATIFGIGNGTIKGGSPRSRVATYKACWSLT 377
Query: 148 DNGRCSGDAILAAFDDAISDGVDVITVSLGPEHASD---FLNDPIAIGSFHAMEKGILTT 204
D C G +LAA D +I DG D+I+VS G + ++ D I+IG+FHA+ + IL
Sbjct: 378 DVVDCFGADVLAAIDQSIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLV 557
Query: 205 QAAGNFGPIP 214
+AGN GP P
Sbjct: 558 ASAGNEGPTP 587
>TC89543 similar to PIR|T06580|T06580 subtilisin-like proteinase (EC
3.4.21.-) p69f - tomato, partial (14%)
Length = 1042
Score = 123 bits (309), Expect(3) = 7e-35
Identities = 70/170 (41%), Positives = 93/170 (54%), Gaps = 4/170 (2%)
Frame = +1
Query: 3 GVISVFPSEEFHIQTTRSWDFLGLR--QSIKRDQIIETDLVIGVIDIRIWPESESFNDKG 60
G++ P + TT S FLGL+ Q + D + ++IGVID I+P SFND+G
Sbjct: 337 GILLARPERTLSLHTTHSPTFLGLKHGQGLWNDDNLGKGVIIGVIDSGIYPYHPSFNDEG 516
Query: 61 LGPIPKKWRGVCAGGGNFSCNNKIIGARFYGDGDVSAR--YSFGHRTHTASTAGGREVED 118
+ P P KW+G C G CNNK+IGAR + + H THTA+ A GR ++D
Sbjct: 517 MPPPPAKWKGHCEFNGRKICNNKLIGARSLVKSTIQEPPFENIFHGTHTAAEAAGRFIKD 696
Query: 119 VSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDG 168
S +G AKG G P++ +A YK+C D C AILAA D AI DG
Sbjct: 697 ASVFGNAKGVXAGMAPNAHLAIYKVCN--DKIECPESAILAAMDIAIEDG 840
Score = 32.0 bits (71), Expect(3) = 7e-35
Identities = 13/30 (43%), Positives = 22/30 (73%)
Frame = +3
Query: 206 AAGNFGPIPSSVCSGAPWLVSVAATSIDRQ 235
+A N GP S++ + APW+++V A+ IDR+
Sbjct: 951 SAANSGPEYSTLSNEAPWILTVGASXIDRK 1040
Score = 30.0 bits (66), Expect(3) = 7e-35
Identities = 16/33 (48%), Positives = 23/33 (69%)
Frame = +2
Query: 169 VDVITVSLGPEHASDFLNDPIAIGSFHAMEKGI 201
V+V+++SLG + F DPIAIG+F A + GI
Sbjct: 842 VNVLSLSLGLX-SLPFFEDPIAIGAFAATQNGI 937
>TC82946 weakly similar to GP|20160863|dbj|BAB89802. putative
subtilisin-like protease {Oryza sativa (japonica
cultivar-group)}, partial (21%)
Length = 921
Score = 90.9 bits (224), Expect(2) = 6e-31
Identities = 56/127 (44%), Positives = 74/127 (58%), Gaps = 11/127 (8%)
Frame = +1
Query: 84 IIGARFYGDGDVS-----------ARYSFGHRTHTASTAGGREVEDVSFYGFAKGTARGG 132
+IGA+F+ G S R S GH THT+STA G V+D SF+G+A GTA G
Sbjct: 553 LIGAKFFNKGLQSKYPNITLGLNSTRDSHGHGTHTSSTAAGNRVDDASFFGYAPGTASGI 732
Query: 133 VPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHASDFLNDPIAIG 192
+SR+A YK D+ G S D ++AA D AISDGVDV+++S G + P+AI
Sbjct: 733 ASNSRVAMYKAIWDV--GVLSSD-VIAAIDAAISDGVDVLSLSFGINDV-PLYDXPVAIA 900
Query: 193 SFHAMEK 199
+F MEK
Sbjct: 901 TFAXMEK 921
Score = 61.2 bits (147), Expect(2) = 6e-31
Identities = 34/86 (39%), Positives = 42/86 (48%), Gaps = 5/86 (5%)
Frame = +3
Query: 3 GVISVFPSEEFHIQTTRSWDFLGLR--QSIKRDQIIETDLVIGVIDIRIWPESESFNDKG 60
G IS + TT S FLGL + D D+++G+ID IWPESESF D
Sbjct: 294 GYISSIKDSHMKLDTTHSPQFLGLNPNKGAWHDSNFGNDVIVGLIDTGIWPESESFKDNL 473
Query: 61 LGPIPKKWRGVCAGGGNFS---CNNK 83
+ IP KW G C F+ CN K
Sbjct: 474 MSEIPSKWNGQCENSILFNSSLCNKK 551
>TC87377 weakly similar to PIR|T07171|T07171 subtilisin-like proteinase (EC
3.4.21.-) 1 - tomato, partial (54%)
Length = 2071
Score = 113 bits (282), Expect = 1e-25
Identities = 82/238 (34%), Positives = 122/238 (50%), Gaps = 11/238 (4%)
Frame = +2
Query: 136 SRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHASDFLNDPIAIGSFH 195
+R+AAYK+C C I A D AI DGV+++++S+G D+ D IAIG+F
Sbjct: 11 ARVAAYKVCW---LSGCFTSDIAAGMDKAIEDGVNILSMSIGGS-IMDYYRDIIAIGAFT 178
Query: 196 AMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKSI-N 254
AM GIL + +AGN GP S+ + APW+ +V A +IDR F + LGNGKT+ G S+ N
Sbjct: 179 AMSHGILVSSSAGNGGPSAESLSNVAPWITTVGAGTIDRDFPSYITLGNGKTYTGASLYN 358
Query: 255 ITPSNGKKIPIA-VRNAQACLAGGNASPEMCDCIDENMVKGKLVLC----GSRNGKDLAY 309
PS+ +P+ N G P D + + V GK+V+C SR K L
Sbjct: 359 GKPSSDSLLPVVYAGNVSESSVGYLCIP---DSLTSSKVLGKIVICERGGNSRVEKGLVV 529
Query: 310 AN-GAIGSI----HNVTKSQLGASFVTPRPSLNLKTNDFVHIQSYTNSSKYPVADFLF 362
N G +G I + + S + P +L K++ ++ Y ++K P A +F
Sbjct: 530 KNAGGVGMILVNNEAYGEELIADSHLLPAAALGQKSSTV--LKDYVFTTKNPRAKLVF 697
>BI262965 similar to PIR|A84473|A84 probable serine proteinase [imported] -
Arabidopsis thaliana, partial (16%)
Length = 394
Score = 112 bits (281), Expect = 2e-25
Identities = 59/128 (46%), Positives = 80/128 (62%)
Frame = +3
Query: 123 GFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHAS 182
G+A GTARG P +RIA YK+C C ILA D AI DGVDV+++SLG ++
Sbjct: 18 GYATGTARGMAPQARIAVYKVCW---TDGCFASDILAGIDQAIQDGVDVLSLSLGGSSST 188
Query: 183 DFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKVIL 242
+ D IAIG+F A+E+GI + +AGN GP S+ + APW+++V A ++DR F L
Sbjct: 189 PYYFDTIAIGAFAAVERGIFVSCSAGNTGPRSGSLSNVAPWIMTVGAGTLDRDFPAYATL 368
Query: 243 GNGKTFVG 250
GNGK G
Sbjct: 369 GNGKRXXG 392
>TC85588 similar to GP|20198169|gb|AAM15440.1 subtilisin-like serine
protease AIR3 {Arabidopsis thaliana}, partial (7%)
Length = 572
Score = 109 bits (273), Expect = 2e-24
Identities = 63/180 (35%), Positives = 97/180 (53%), Gaps = 6/180 (3%)
Frame = +1
Query: 132 GVPSSRIAAYKICGDI-DNGRCSGDAILAAFDDAISDGVDVITVSLGPE---HASDFLND 187
G P +R+A YK+C + D C G +LAA D AISDGVD+I++SL + D D
Sbjct: 1 GSPRARVATYKVCWSLLDLEDCFGADVLAAIDQAISDGVDIISLSLAGHSLVYPEDIFTD 180
Query: 188 PIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKVILGNGKT 247
++IG+FHA+ + IL +AGN GP SV + PW+ ++AA+++DR F + +GN +T
Sbjct: 181 EVSIGAFHALSRNILLVASAGNEGPTGGSVVNVTPWVFTIAASTLDRDFSSTITIGN-QT 357
Query: 248 FVGKSINITPSNGKKIPIAVRNAQACLAGGNASPEMC--DCIDENMVKGKLVLCGSRNGK 305
G S+ + + P+ V N + C +D + VKG L+ + GK
Sbjct: 358 IRGASLFVNLPPNQAFPLIVSTDGKLANATNHDAQFCKPGTLDPSKVKGMLLSNQPKQGK 537
>CB891986 similar to PIR|S52769|S52 subtilisin-like proteinase ag12 (EC
3.4.21.-) - alder, partial (14%)
Length = 598
Score = 107 bits (267), Expect = 8e-24
Identities = 75/215 (34%), Positives = 123/215 (56%), Gaps = 2/215 (0%)
Frame = +1
Query: 124 FAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHASD 183
+AKG ARG P +R+A YK+ + G + D +LA D AI+DGVDVI++S+G +
Sbjct: 1 YAKGVARGIAPRARLAMYKVIWE--EGLLASD-VLAGMDQAIADGVDVISISMGFDGVPL 171
Query: 184 FLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKVILG 243
+ D IAI SF AMEKGI+ + +AGN GP ++ +G PW+++VAA +IDR F ++LG
Sbjct: 172 Y-EDAIAIASFAAMEKGIVVSSSAGNSGPKHGTLHNGIPWVLTVAAGTIDRTF-GSLVLG 345
Query: 244 NGKTFVGKSINITPSN-GKKIPIAVRNAQACLAGGNASPEMCDCIDENMVKGKLVLCGSR 302
NG+ +G ++ + S + +P+ N L+ N+ + + K +++C S
Sbjct: 346 NGQNIIGWTLFASNSTIVENLPLVYDNT---LSSCNSVKRL-----SQVNKQVIIICDS- 498
Query: 303 NGKDLAYANGAIGSIHNVTK-SQLGASFVTPRPSL 336
++ ++ I VT+ + LGA F++ P L
Sbjct: 499 ----ISNSSSVFDQIDVVTQTNMLGAVFLSDSPEL 591
>BI307979 weakly similar to GP|20521377|db putative subtilisin-like protease
{Oryza sativa (japonica cultivar-group)}, partial (18%)
Length = 777
Score = 102 bits (255), Expect = 2e-22
Identities = 70/213 (32%), Positives = 108/213 (49%), Gaps = 9/213 (4%)
Frame = +3
Query: 158 LAAFDDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSV 217
L A DDAI DGVDVI +S+G + +D IA G+ A+ K I+ +AGN GP P S+
Sbjct: 3 LKAIDDAIEDGVDVINLSIGFPAPLKYEDDVIAKGALQAVRKNIVVVCSAGNAGPSPHSL 182
Query: 218 CSGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKSINITPSNGKKIPIAVRN--AQACLA 275
+ +PW+++V A+++DR F+ + L NG T G+SI P+ + + A +
Sbjct: 183 SNPSPWIITVGASTVDRTFLAPIKLSNGTTIEGRSITPLRMGNSFCPLVLASDVEYAGIL 362
Query: 276 GGNASPEMCDCIDENMVKGKLVLC----GSRNGKDLAYAN-GAIGSIHNVTKSQLGASFV 330
N+S + + +D + VKGK+VLC G R K L G +G I K
Sbjct: 363 SANSSYCLDNTLDPSKVKGKIVLCMRGQGGRVKKSLEVQRAGGVGLILGNNKVYANDVPS 542
Query: 331 TPR--PSLNLKTNDFVHIQSYTNSSKYPVADFL 361
P P+ + + + + Y +SS P+A L
Sbjct: 543 DPYFIPTTGVTYENTLKLVQYIHSSPNPMAQLL 641
>AW584611 similar to PIR|T07184|T071 subtilisin-like proteinase (EC 3.4.21.-)
precursor P69B pathogenesis-related - tomato, partial
(11%)
Length = 706
Score = 88.2 bits (217), Expect = 5e-18
Identities = 49/126 (38%), Positives = 69/126 (53%), Gaps = 4/126 (3%)
Frame = +1
Query: 4 VISVFPSEEFHIQTTRSWDFLGLRQS--IKRDQIIETDLVIGVIDIRIWPESESFNDKGL 61
++S P + TT + FLGL+Q + D + ++IG+ID I+P SFND+G+
Sbjct: 328 IVSARPERTLELHTTHTPTFLGLKQGQGLWSDDNLGKGVIIGIIDTGIFPLHPSFNDEGM 507
Query: 62 GPIPKKWRGVCAGGGNFSCNNKIIGARFYGDGDVSAR--YSFGHRTHTASTAGGREVEDV 119
P P KW+G C G CNNK+IGAR + +F H THTA+ A GR +ED
Sbjct: 508 PPPPAKWKGHCEFTGGQVCNNKLIGARNLVKSAIQEPPFENFFHGTHTAAEAAGRFIEDA 687
Query: 120 SFYGFA 125
+G A
Sbjct: 688 KCFGNA 705
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.137 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,048,232
Number of Sequences: 36976
Number of extensions: 168444
Number of successful extensions: 836
Number of sequences better than 10.0: 70
Number of HSP's better than 10.0 without gapping: 757
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 762
length of query: 395
length of database: 9,014,727
effective HSP length: 98
effective length of query: 297
effective length of database: 5,391,079
effective search space: 1601150463
effective search space used: 1601150463
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC148652.13


