

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148607.5 - phase: 0
(122 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
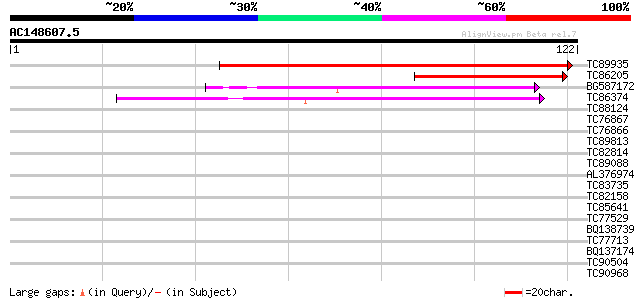
Score E
Sequences producing significant alignments: (bits) Value
TC89935 similar to GP|16604683|gb|AAL24134.1 putative ABC transp... 88 7e-19
TC86205 similar to PIR|A86289|A86289 probable ABC transporter [i... 59 6e-10
BG587172 similar to PIR|T45888|T4 ABC transporter-like protein -... 44 1e-05
TC86374 similar to GP|18377973|gb|AAL67129.1 putative ABC transp... 40 2e-04
TC88124 weakly similar to PIR|T45888|T45888 ABC transporter-like... 35 0.005
TC76867 similar to GP|2909420|emb|CAA12026.1 LEA protein {Cicer ... 32 0.060
TC76866 similar to GP|2909420|emb|CAA12026.1 LEA protein {Cicer ... 32 0.060
TC89813 similar to GP|18377973|gb|AAL67129.1 putative ABC transp... 30 0.30
TC82814 homologue to GP|10176814|dbj|BAB10022. endosomal protein... 28 0.66
TC89088 homologue to PIR|H85135|H85135 hypothetical protein AT4g... 27 1.5
AL376974 similar to PIR|B90137|B901 sulfate permease [imported] ... 27 1.9
TC83735 similar to PIR|A84509|A84509 probable ABC transporter [i... 27 2.5
TC82158 weakly similar to GP|19920013|gb|AAM08453.1 hypothetical... 27 2.5
TC85641 weakly similar to GP|8777295|dbj|BAA96885.1 cytochrome P... 27 2.5
TC77529 homologue to PIR|T52063|T52063 ran GTPase-activating pro... 26 3.3
BQ138739 26 3.3
TC77713 similar to PIR|S62667|S62667 Nramp1 protein - rice, part... 26 3.3
BQ137174 26 4.3
TC90504 similar to PIR|T48329|T48329 hypothetical protein F15A17... 26 4.3
TC90968 similar to SP|O81971|C7D9_SOYBN Cytochrome P450 71D9 (EC... 26 4.3
>TC89935 similar to GP|16604683|gb|AAL24134.1 putative ABC transporter
protein {Arabidopsis thaliana}, partial (34%)
Length = 1244
Score = 88.2 bits (217), Expect = 7e-19
Identities = 36/76 (47%), Positives = 53/76 (69%)
Frame = +3
Query: 46 LFYGCIVILFIGVAELSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWV 105
+ + I+ +F G +EL + ++RLPVFYK R +LF PPW Y LP ++L+IP++ E VWV
Sbjct: 3 ILFTMIMNMFNGFSELPLTIARLPVFYKHRDHLFHPPWTYTLPNFLLRIPISIFEAIVWV 182
Query: 106 ILTYYVIGFDPYIGRY 121
++TYY IGF P R+
Sbjct: 183 LITYYTIGFAPEASRF 230
>TC86205 similar to PIR|A86289|A86289 probable ABC transporter [imported] -
Arabidopsis thaliana, partial (52%)
Length = 2721
Score = 58.5 bits (140), Expect = 6e-10
Identities = 23/33 (69%), Positives = 30/33 (90%)
Frame = +1
Query: 88 PAWILKIPLTFVEVAVWVILTYYVIGFDPYIGR 120
P+WILKIP++ +EV++WV LTYYVIGFDP +GR
Sbjct: 1 PSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGR 99
>BG587172 similar to PIR|T45888|T4 ABC transporter-like protein - Arabidopsis
thaliana, partial (8%)
Length = 577
Score = 44.3 bits (103), Expect = 1e-05
Identities = 28/76 (36%), Positives = 43/76 (55%), Gaps = 4/76 (5%)
Frame = -1
Query: 43 VGALFYGCIVILFIGVAELSMVVSRLP----VFYKQRGYLFFPPWAYALPAWILKIPLTF 98
VGA+ YG + LF+G+ S + L V Y++R + AYAL + +IP F
Sbjct: 295 VGAI-YGAV--LFLGINNCSSAIRNLETERNVMYRERFAGMYSATAYALGQVVTEIPYLF 125
Query: 99 VEVAVWVILTYYVIGF 114
++ A +VI+TY +IGF
Sbjct: 124 IQAAEFVIITYPMIGF 77
>TC86374 similar to GP|18377973|gb|AAL67129.1 putative ABC transporter
protein {Arabidopsis thaliana}, partial (38%)
Length = 1011
Score = 40.0 bits (92), Expect = 2e-04
Identities = 19/96 (19%), Positives = 47/96 (48%), Gaps = 4/96 (4%)
Frame = +2
Query: 24 TIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAELS----MVVSRLPVFYKQRGYLF 79
T+F + +++S + +GA++ ++F+G+ +V VFY++R
Sbjct: 5 TVFWKVGENKESSTDLTLVIGAMY---AAVIFVGINNCQTVQPVVAIERTVFYRERAAGM 175
Query: 80 FPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFD 115
+ P YAL ++++P + + ++ Y ++ F+
Sbjct: 176 YAPLPYALAQVLIEVPFVLFQACYYSLIVYAMVSFE 283
>TC88124 weakly similar to PIR|T45888|T45888 ABC transporter-like protein -
Arabidopsis thaliana, partial (27%)
Length = 1641
Score = 35.4 bits (80), Expect = 0.005
Identities = 20/76 (26%), Positives = 36/76 (47%)
Frame = +3
Query: 39 GDIYVGALFYGCIVILFIGVAELSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTF 98
G +Y+ +F G I L V + V Y+++ + AY+ ++IP
Sbjct: 744 GSMYIAVIFLGINYCSTI----LPYVATERSVLYREKFAGMYSSMAYSFAQVAIEIPYIL 911
Query: 99 VEVAVWVILTYYVIGF 114
V+ ++V +TY +IGF
Sbjct: 912 VQAIIYVAITYPMIGF 959
>TC76867 similar to GP|2909420|emb|CAA12026.1 LEA protein {Cicer arietinum},
partial (85%)
Length = 828
Score = 32.0 bits (71), Expect = 0.060
Identities = 19/56 (33%), Positives = 27/56 (47%), Gaps = 1/56 (1%)
Frame = -2
Query: 52 VILFIGVAELSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFV-EVAVWVI 106
V L VA ++ ++ PV K L FP WA + AW + PL+ V WV+
Sbjct: 503 VFLTASVAPCAIPLTFSPVCCKNPAELSFPDWAVSFVAWPIS*PLSLA*PVVSWVL 336
>TC76866 similar to GP|2909420|emb|CAA12026.1 LEA protein {Cicer arietinum},
partial (85%)
Length = 781
Score = 32.0 bits (71), Expect = 0.060
Identities = 19/56 (33%), Positives = 27/56 (47%), Gaps = 1/56 (1%)
Frame = -2
Query: 52 VILFIGVAELSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFV-EVAVWVI 106
V L VA ++ ++ PV K L FP WA + AW + PL+ V WV+
Sbjct: 483 VFLTASVAPCAIPLTFSPVCCKNPAELSFPDWAVSFVAWPIS*PLSLA*PVVSWVL 316
>TC89813 similar to GP|18377973|gb|AAL67129.1 putative ABC transporter
protein {Arabidopsis thaliana}, partial (30%)
Length = 837
Score = 29.6 bits (65), Expect = 0.30
Identities = 13/46 (28%), Positives = 24/46 (51%)
Frame = +1
Query: 70 VFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFD 115
VFY++R + YAL I +IP F + + ++ Y ++ F+
Sbjct: 19 VFYRERAAGMYSALPYALAQVICEIPYVFGQTIFFSVIVYPMVSFE 156
>TC82814 homologue to GP|10176814|dbj|BAB10022. endosomal protein-like
{Arabidopsis thaliana}, partial (36%)
Length = 1050
Score = 28.5 bits (62), Expect = 0.66
Identities = 15/40 (37%), Positives = 20/40 (49%), Gaps = 5/40 (12%)
Frame = +3
Query: 80 FPPW-----AYALPAWILKIPLTFVEVAVWVILTYYVIGF 114
+P W A LP L I L F+ ++W+ YYV GF
Sbjct: 285 YPSWLLVLGAGTLPFGTLFIELFFIMSSIWMGRVYYVFGF 404
>TC89088 homologue to PIR|H85135|H85135 hypothetical protein AT4g12650
[imported] - Arabidopsis thaliana, partial (53%)
Length = 1275
Score = 27.3 bits (59), Expect = 1.5
Identities = 15/40 (37%), Positives = 20/40 (49%), Gaps = 5/40 (12%)
Frame = +2
Query: 80 FPPW-----AYALPAWILKIPLTFVEVAVWVILTYYVIGF 114
+P W A LP L I L F+ ++W+ YYV GF
Sbjct: 401 YPSWLLVLGAGTLPFGTLFIELFFILSSIWLGRFYYVFGF 520
>AL376974 similar to PIR|B90137|B901 sulfate permease [imported] - Guillardia
theta nucleomorph, partial (2%)
Length = 437
Score = 26.9 bits (58), Expect = 1.9
Identities = 12/28 (42%), Positives = 17/28 (59%)
Frame = +2
Query: 2 KRNSFVYIFKLCQLAIMAMIAMTIFLRT 29
KRN+FVYIF LC ++ + +L T
Sbjct: 188 KRNAFVYIFDLCHARLVFDVINFEYLAT 271
>TC83735 similar to PIR|A84509|A84509 probable ABC transporter [imported] -
Arabidopsis thaliana, partial (34%)
Length = 872
Score = 26.6 bits (57), Expect = 2.5
Identities = 15/68 (22%), Positives = 32/68 (47%), Gaps = 7/68 (10%)
Frame = +1
Query: 55 FIGVAELSMVVSRLPVFYKQRGYLF-------FPPWAYALPAWILKIPLTFVEVAVWVIL 107
FI LS + LP+F ++R L + +YA+ ++ +P + ++ +
Sbjct: 28 FILTFLLSSAIEALPIFLQEREILMKETSSGSYRVSSYAIANGLVYLPFLLILAILFAVP 207
Query: 108 TYYVIGFD 115
Y++IG +
Sbjct: 208 LYWIIGLN 231
>TC82158 weakly similar to GP|19920013|gb|AAM08453.1 hypothetical protein
{Dictyostelium discoideum}, partial (4%)
Length = 1230
Score = 26.6 bits (57), Expect = 2.5
Identities = 18/57 (31%), Positives = 28/57 (48%), Gaps = 2/57 (3%)
Frame = -2
Query: 68 LPVFYKQRGYLFFPPWAY-ALPAWILKI-PLTFVEVAVWVILTYYVIGFDPYIGRYE 122
LP++ Q +L+ + +LP LKI P+ F ++ L FDP+ G YE
Sbjct: 1097 LPIYPSQMAFLWIVDYHLRSLPKRTLKIIPIFFFQIYHKQTLQEDPPNFDPFFGEYE 927
>TC85641 weakly similar to GP|8777295|dbj|BAA96885.1 cytochrome P450-like
{Arabidopsis thaliana}, partial (71%)
Length = 1941
Score = 26.6 bits (57), Expect = 2.5
Identities = 14/42 (33%), Positives = 21/42 (49%)
Frame = +1
Query: 47 FYGCIVILFIGVAELSMVVSRLPVFYKQRGYLFFPPWAYALP 88
FY +++LF+ LS+ +FYKQ+ L PP P
Sbjct: 124 FYLSLLLLFVTFISLSLFF----IFYKQKSPLNLPPGKMGYP 237
>TC77529 homologue to PIR|T52063|T52063 ran GTPase-activating protein
[imported] - alfalfa, partial (64%)
Length = 1392
Score = 26.2 bits (56), Expect = 3.3
Identities = 18/57 (31%), Positives = 30/57 (52%)
Frame = +1
Query: 51 IVILFIGVAELSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVIL 107
+ + F+GVAELS++ +L F W L L+I ++ +EV VW++L
Sbjct: 736 LFLTFLGVAELSLMGRKLQSF*NH-------*WDQTL----LQIYVSVIEVLVWMLL 873
>BQ138739
Length = 676
Score = 26.2 bits (56), Expect = 3.3
Identities = 8/24 (33%), Positives = 16/24 (66%)
Frame = -1
Query: 80 FPPWAYALPAWILKIPLTFVEVAV 103
FPP ++ LP ++K+P F++ +
Sbjct: 373 FPPPSFLLPPSVIKVPFPFIKCPI 302
>TC77713 similar to PIR|S62667|S62667 Nramp1 protein - rice, partial (84%)
Length = 1575
Score = 26.2 bits (56), Expect = 3.3
Identities = 19/61 (31%), Positives = 32/61 (52%), Gaps = 7/61 (11%)
Frame = +3
Query: 15 LAIMAMIAMTI--FLRTEMHRDSVAHGDIYVGALFYGCIVILFI-----GVAELSMVVSR 67
LA +A+IA I + T + + H ++ G L GC +LF+ GV +L +++S
Sbjct: 435 LAELAVIAADIPEVIGTAFALNILFHVPVWGGVLLTGCSTLLFLGLQRFGVRKLELLISI 614
Query: 68 L 68
L
Sbjct: 615 L 617
>BQ137174
Length = 977
Score = 25.8 bits (55), Expect = 4.3
Identities = 18/75 (24%), Positives = 34/75 (45%), Gaps = 2/75 (2%)
Frame = +1
Query: 8 YIFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAELSMVVSR 67
Y+F + + ++ I RT++HR + D+ YGC V+ + M++
Sbjct: 760 YMFIVILFVCVLLLLYCISCRTQLHRH*LIIADV------YGCDVLRVCVMI*WKMILCD 921
Query: 68 LPVFYKQR--GYLFF 80
+ V Y Q+ YL +
Sbjct: 922 IIVVYNQQIIAYLLY 966
>TC90504 similar to PIR|T48329|T48329 hypothetical protein F15A17.110 -
Arabidopsis thaliana, partial (54%)
Length = 611
Score = 25.8 bits (55), Expect = 4.3
Identities = 14/43 (32%), Positives = 21/43 (48%), Gaps = 7/43 (16%)
Frame = +3
Query: 15 LAIMAMIAMTIFLRTEMHRDSVAH-------GDIYVGALFYGC 50
L I ++ +T++ R + +VAH GD Y G L GC
Sbjct: 465 LFIWSLAILTVYSRVYLGYHNVAHVFCWY*FGDFYWGGLVLGC 593
>TC90968 similar to SP|O81971|C7D9_SOYBN Cytochrome P450 71D9 (EC 1.14.-.-)
(P450 CP3). [Soybean] {Glycine max}, partial (64%)
Length = 1112
Score = 25.8 bits (55), Expect = 4.3
Identities = 10/25 (40%), Positives = 14/25 (56%)
Frame = +1
Query: 95 PLTFVEVAVWVILTYYVIGFDPYIG 119
P +EV +W I T + GF Y+G
Sbjct: 409 PPELLEVLIWEICTLLINGFKTYLG 483
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.336 0.149 0.486
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,271,239
Number of Sequences: 36976
Number of extensions: 60093
Number of successful extensions: 463
Number of sequences better than 10.0: 58
Number of HSP's better than 10.0 without gapping: 459
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 463
length of query: 122
length of database: 9,014,727
effective HSP length: 84
effective length of query: 38
effective length of database: 5,908,743
effective search space: 224532234
effective search space used: 224532234
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC148607.5


