

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148607.15 - phase: 0
(174 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
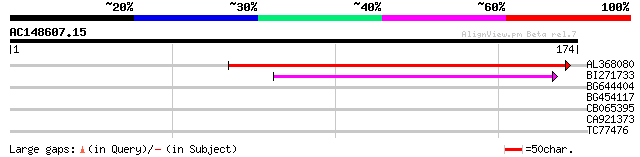
AL368080 similar to GP|20259567|gb unknown protein {Arabidopsis ... 181 1e-46
BI271733 similar to GP|10177721|db gene_id:MPL12.20~pir||T00964~... 75 1e-14
BG644404 similar to GP|21593919|gb unknown {Arabidopsis thaliana... 30 0.49
BG454117 similar to GP|6437558|gb putative ATPase (ISW2-like) {A... 27 3.2
CB065395 similar to GP|23343453|emb repressor {Bacteriophage Nil... 27 5.4
CA921373 weakly similar to PIR|T14519|T14 probable S-receptor ki... 26 9.3
TC77476 homologue to GP|22296820|gb|AAM94349.1 pyruvate kinase {... 26 9.3
>AL368080 similar to GP|20259567|gb unknown protein {Arabidopsis thaliana},
partial (15%)
Length = 389
Score = 181 bits (459), Expect = 1e-46
Identities = 88/105 (83%), Positives = 92/105 (86%)
Frame = +3
Query: 68 HPDGADNSPKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGFETTIRRPYFHVRPL 127
HPDGADNSPKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGFETTIRRPYFHVRPL
Sbjct: 3 HPDGADNSPKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGFETTIRRPYFHVRPL 182
Query: 128 NIGELENWHNYLDFIEREGDLSKVTKLLSYKGTFCSIFSCYRLTY 172
NIGELENWHNYLDFIEREGDLSK+ KL C+ + Y + Y
Sbjct: 183 NIGELENWHNYLDFIEREGDLSKIVKLYERCVIACANYPEYWIRY 317
>BI271733 similar to GP|10177721|db gene_id:MPL12.20~pir||T00964~strong
similarity to unknown protein {Arabidopsis thaliana},
partial (5%)
Length = 383
Score = 75.1 bits (183), Expect = 1e-14
Identities = 39/87 (44%), Positives = 49/87 (55%)
Frame = +3
Query: 82 GLTEAEELEKYIAIREEMYKKAKEFDSKIIGFETTIRRPYFHVRPLNIGELENWHNYLDF 141
GLT + L+KY I E++Y A E SKI FE I+R YF RPL+ +L+NWH YLDF
Sbjct: 114 GLTSSTALKKYRIIGEQLYHNACELYSKISSFEANIQRYYFDFRPLDANQLQNWHAYLDF 293
Query: 142 IEREGDLSKVTKLLSYKGTFCSIFSCY 168
IE GD KL C+ + Y
Sbjct: 294 IELHGDFDWAVKLYERCLIVCANYPDY 374
>BG644404 similar to GP|21593919|gb unknown {Arabidopsis thaliana}, partial
(46%)
Length = 478
Score = 30.0 bits (66), Expect = 0.49
Identities = 23/74 (31%), Positives = 32/74 (43%), Gaps = 5/74 (6%)
Frame = +1
Query: 41 ELRTADEAAAVAGVVSEGIDQGVEGEVHPDGADNSPKPASAGLTEAEE-----LEKYIAI 95
E+R D A VAG S + E P N +++A++ L K AI
Sbjct: 61 EVRLVDNAVNVAGESSSPL-----AEAKPTSPSNGRVNGKVYISKAQDVVSNVLAKGSAI 225
Query: 96 REEMYKKAKEFDSK 109
R++ KAK FD K
Sbjct: 226 RQDAMNKAKAFDEK 267
>BG454117 similar to GP|6437558|gb putative ATPase (ISW2-like) {Arabidopsis
thaliana}, partial (16%)
Length = 658
Score = 27.3 bits (59), Expect = 3.2
Identities = 30/110 (27%), Positives = 40/110 (36%), Gaps = 6/110 (5%)
Frame = +2
Query: 1 MQQDWARLAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVVSEGID 60
+Q+ A+L+ E P L +E N ++E +E AVA VS D
Sbjct: 5 IQESMAKLSKQQPSSDEEPVNSLS------EEEQVNEEINEEEDQEELEAVARAVSSDDD 166
Query: 61 QGVEGEVHPD------GADNSPKPASAGLTEAEELEKYIAIREEMYKKAK 104
V GE PD G D G E + EK + KK K
Sbjct: 167 DEVAGENPPDSDADVAGEDGDDDGEGEGGPEISKREKERLREMQKLKKQK 316
>CB065395 similar to GP|23343453|emb repressor {Bacteriophage Nil2}, partial
(12%)
Length = 610
Score = 26.6 bits (57), Expect = 5.4
Identities = 10/26 (38%), Positives = 17/26 (64%)
Frame = +1
Query: 126 PLNIGELENWHNYLDFIEREGDLSKV 151
P +ENW+++L +RE DLS++
Sbjct: 277 PYRCALMENWNSFLKRYKREHDLSQI 354
>CA921373 weakly similar to PIR|T14519|T14 probable S-receptor kinase (EC
2.7.1.-) SFR1 - wild cabbage, partial (13%)
Length = 779
Score = 25.8 bits (55), Expect = 9.3
Identities = 11/33 (33%), Positives = 18/33 (54%), Gaps = 6/33 (18%)
Frame = -3
Query: 136 HNYLDFIEREGDLSKVTKL------LSYKGTFC 162
H Y D++E GD+ +T + LSY ++C
Sbjct: 171 HTYDDYVENSGDIIVITNIHCLYHCLSYLPSYC 73
>TC77476 homologue to GP|22296820|gb|AAM94349.1 pyruvate kinase {Glycine max},
complete
Length = 1970
Score = 25.8 bits (55), Expect = 9.3
Identities = 12/37 (32%), Positives = 21/37 (56%)
Frame = -1
Query: 39 LSELRTADEAAAVAGVVSEGIDQGVEGEVHPDGADNS 75
L +RTAD+A G++ G+D + ++ P D+S
Sbjct: 1322 LLAVRTADDARLSNGLIGNGVDLIISLKIAP*SRDDS 1212
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.135 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,830,368
Number of Sequences: 36976
Number of extensions: 54246
Number of successful extensions: 305
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 305
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 305
length of query: 174
length of database: 9,014,727
effective HSP length: 89
effective length of query: 85
effective length of database: 5,723,863
effective search space: 486528355
effective search space used: 486528355
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 55 (25.8 bits)
Medicago: description of AC148607.15


