

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148607.14 - phase: 0
(359 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
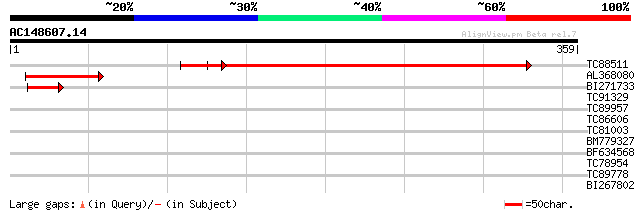
Sequences producing significant alignments: (bits) Value
TC88511 weakly similar to GP|20259567|gb|AAM14126.1 unknown prot... 391 e-109
AL368080 similar to GP|20259567|gb unknown protein {Arabidopsis ... 99 3e-21
BI271733 similar to GP|10177721|db gene_id:MPL12.20~pir||T00964~... 45 3e-05
TC91329 similar to GP|7682783|gb|AAF67364.1| Hypothetical protei... 32 0.49
TC89957 weakly similar to PIR|D96810|D96810 hypothetical protein... 30 1.4
TC86606 similar to SP|Q42777|MCCA_SOYBN Methylcrotonyl-CoA carbo... 30 1.9
TC81003 homologue to GP|18375499|gb|AAK09427.2 sucrose-phosphate... 29 3.2
BM779327 weakly similar to GP|18700131|gb AT4g24270/T22A6_100 {A... 28 4.2
BF634568 weakly similar to SP|Q38882|PDA1 Phospholipase D alpha ... 28 4.2
TC78954 weakly similar to PIR|D96810|D96810 hypothetical protein... 28 5.5
TC89778 similar to PIR|T00747|T00747 RING-H2 finger protein RHC1... 28 7.1
BI267802 similar to GP|14517514|gb AT5g52010/MSG15_9 {Arabidopsi... 27 9.3
>TC88511 weakly similar to GP|20259567|gb|AAM14126.1 unknown protein
{Arabidopsis thaliana}, partial (22%)
Length = 1060
Score = 391 bits (1005), Expect = e-109
Identities = 195/205 (95%), Positives = 198/205 (96%)
Frame = +3
Query: 126 IAIEKGKEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEA 185
+ + K K + LPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEA
Sbjct: 54 LPLRKEKNIHKHLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEA 233
Query: 186 IQPQPKRVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFGDVQSIKRAE 245
IQPQPKRVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFGDVQSIKRAE
Sbjct: 234 IQPQPKRVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFGDVQSIKRAE 413
Query: 246 DRHAKLFLPNRGLSELKKRHAEDFLASDKTKVSRAYSAQSPAQSVAGAYPNGPNQWPNYG 305
DRHAKLFLPNRGLSELKKRHAEDFLASDKTKVSRAYSAQSPAQSVAGAYPNGPNQWPNYG
Sbjct: 414 DRHAKLFLPNRGLSELKKRHAEDFLASDKTKVSRAYSAQSPAQSVAGAYPNGPNQWPNYG 593
Query: 306 VQPQTWPATTQAQGQQWPAGYTQQQ 330
VQPQTWPATTQAQGQQWPAGYTQQQ
Sbjct: 594 VQPQTWPATTQAQGQQWPAGYTQQQ 668
Score = 61.2 bits (147), Expect = 6e-10
Identities = 29/29 (100%), Positives = 29/29 (100%)
Frame = +2
Query: 109 EHRLGKLEDAFSLYEQAIAIEKGKEHSQT 137
EHRLGKLEDAFSLYEQAIAIEKGKEHSQT
Sbjct: 2 EHRLGKLEDAFSLYEQAIAIEKGKEHSQT 88
>AL368080 similar to GP|20259567|gb unknown protein {Arabidopsis thaliana},
partial (15%)
Length = 389
Score = 98.6 bits (244), Expect = 3e-21
Identities = 46/49 (93%), Positives = 48/49 (97%)
Frame = +3
Query: 11 VDQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVF 59
+ +IVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVF
Sbjct: 243 LSKIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVF 389
>BI271733 similar to GP|10177721|db gene_id:MPL12.20~pir||T00964~strong
similarity to unknown protein {Arabidopsis thaliana},
partial (5%)
Length = 383
Score = 45.4 bits (106), Expect = 3e-05
Identities = 17/23 (73%), Positives = 20/23 (86%)
Frame = +3
Query: 12 DQIVKLYERCVIACANYPEYWIR 34
D VKLYERC+I CANYP+YW+R
Sbjct: 315 DWAVKLYERCLIVCANYPDYWMR 383
>TC91329 similar to GP|7682783|gb|AAF67364.1| Hypothetical protein T32B20.g
{Arabidopsis thaliana}, partial (38%)
Length = 1068
Score = 31.6 bits (70), Expect = 0.49
Identities = 36/142 (25%), Positives = 59/142 (41%), Gaps = 6/142 (4%)
Frame = +3
Query: 41 ASESMDLANNVLARASQ-VFVKRQPEIHL--FCARFKEQAGDIVGARAAYQLVHTEISPG 97
A S+++ V A +Q V +K + L F +E G + R Y+ + ++
Sbjct: 222 AEPSVEVKRKVAADGNQPVQMKLHKSLRLWTFFVDLEESLGSLESTREVYERI-LDLRIA 398
Query: 98 LLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKGKEHSQTLPMLFAQYSRFV-YLASGNS 156
+ II +A EDAF +YE+ + I K + + S+FV
Sbjct: 399 TPQIIINYAYFLEEHKYFEDAFKVYERGVKIFK---YPHVKDIWVTYLSKFVKRYGRTKL 569
Query: 157 EKAREILVGGLENASLS--KPL 176
E+ARE+ +E A KPL
Sbjct: 570 ERARELFENAVETAPADQVKPL 635
>TC89957 weakly similar to PIR|D96810|D96810 hypothetical protein T11I11.6
[imported] - Arabidopsis thaliana, partial (51%)
Length = 1843
Score = 30.0 bits (66), Expect = 1.4
Identities = 13/33 (39%), Positives = 21/33 (63%)
Frame = +3
Query: 100 EAIIRHANMEHRLGKLEDAFSLYEQAIAIEKGK 132
E + N ++ GK E+A +LY++AIAI+ K
Sbjct: 609 EGLKSMGNEAYKKGKFEEALALYDKAIAIDSNK 707
>TC86606 similar to SP|Q42777|MCCA_SOYBN Methylcrotonyl-CoA carboxylase alpha
chain mitochondrial precursor (EC 6.4.1.4), partial
(95%)
Length = 2532
Score = 29.6 bits (65), Expect = 1.9
Identities = 31/137 (22%), Positives = 56/137 (40%), Gaps = 15/137 (10%)
Frame = +1
Query: 69 FCARFKEQA----GDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQ 124
+C K Q+ G I+G + L T I P + + +++ N L KL+D+ + ++
Sbjct: 1213 YCTIIKFQSHQGSGWILGLKKEMLLACTTI-PMIAKLVVQGENRAAALVKLKDSLTKFQV 1389
Query: 125 A-----------IAIEKGKEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLS 173
A +A + E+ Y +++ + NSE A+E +ASL
Sbjct: 1390 AGLPTNVNFLLKLANHRAFENGNVETHFIDNYKEDLFVDATNSESAKEAYEAARRSASLV 1569
Query: 174 KPLLEALLHFEAIQPQP 190
L HF + + P
Sbjct: 1570 AACLIEKEHFISARNPP 1620
>TC81003 homologue to GP|18375499|gb|AAK09427.2 sucrose-phosphate synthase
{Medicago sativa}, partial (30%)
Length = 1164
Score = 28.9 bits (63), Expect = 3.2
Identities = 15/39 (38%), Positives = 21/39 (53%)
Frame = -1
Query: 282 SAQSPAQSVAGAYPNGPNQWPNYGVQPQTWPATTQAQGQ 320
+A+SPA + N P+ PNY PQ P+T+ GQ
Sbjct: 1158 AAESPASXIMAVNGNXPDSMPNYQFAPQE-PSTSV*CGQ 1045
>BM779327 weakly similar to GP|18700131|gb AT4g24270/T22A6_100 {Arabidopsis
thaliana}, partial (19%)
Length = 518
Score = 28.5 bits (62), Expect = 4.2
Identities = 14/39 (35%), Positives = 21/39 (52%), Gaps = 1/39 (2%)
Frame = +3
Query: 9 LFVDQIVK-LYERCVIACANYPEYWIRYVLCMEASESMD 46
L V +IV +Y R C E W+RY+LC+E + +
Sbjct: 321 LKVGKIVSDVYSRATKNCPWVGELWVRYMLCLERGHASE 437
>BF634568 weakly similar to SP|Q38882|PDA1 Phospholipase D alpha 1 (EC
3.1.4.4) (AtPLDalpha1) (PLD alpha 1) (Choline
phosphatase 1), partial (13%)
Length = 687
Score = 28.5 bits (62), Expect = 4.2
Identities = 13/40 (32%), Positives = 21/40 (52%)
Frame = +1
Query: 119 FSLYEQAIAIEKGKEHSQTLPMLFAQYSRFVYLASGNSEK 158
+ L AI+I +EH+Q F SR V+ +GN+ +
Sbjct: 220 YKLLPDAISISATREHAQAWGRKFCHKSRVVFCVNGNASQ 339
>TC78954 weakly similar to PIR|D96810|D96810 hypothetical protein T11I11.6
[imported] - Arabidopsis thaliana, partial (45%)
Length = 2329
Score = 28.1 bits (61), Expect = 5.5
Identities = 12/33 (36%), Positives = 20/33 (60%)
Frame = +3
Query: 100 EAIIRHANMEHRLGKLEDAFSLYEQAIAIEKGK 132
E + N ++ G+ +A +LYE+AIAI+ K
Sbjct: 807 EVLKSMGNEAYKQGRFSEALALYERAIAIDSNK 905
>TC89778 similar to PIR|T00747|T00747 RING-H2 finger protein RHC1a
[imported] - Arabidopsis thaliana, partial (29%)
Length = 989
Score = 27.7 bits (60), Expect = 7.1
Identities = 18/62 (29%), Positives = 29/62 (46%), Gaps = 3/62 (4%)
Frame = -3
Query: 114 KLEDAFSLYEQAIAIEKGKEHSQTLPMLFAQYS---RFVYLASGNSEKAREILVGGLENA 170
K + FS + A+AI GK+ QY+ +VY+A + +I +G L+N
Sbjct: 699 KTKKGFSFFHSAVAILDGKQGRCYAEPTMVQYNPSETYVYMAFVLHQTQAQIYLGILDNV 520
Query: 171 SL 172
L
Sbjct: 519 DL 514
>BI267802 similar to GP|14517514|gb AT5g52010/MSG15_9 {Arabidopsis thaliana},
partial (35%)
Length = 606
Score = 27.3 bits (59), Expect = 9.3
Identities = 14/36 (38%), Positives = 20/36 (54%)
Frame = +1
Query: 235 FGDVQSIKRAEDRHAKLFLPNRGLSELKKRHAEDFL 270
FGDV I +RHA + LP L++ ++R D L
Sbjct: 274 FGDVTDISAYANRHAFIHLPQWVLNQRRERKNLDIL 381
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,038,926
Number of Sequences: 36976
Number of extensions: 149461
Number of successful extensions: 739
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 734
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 738
length of query: 359
length of database: 9,014,727
effective HSP length: 97
effective length of query: 262
effective length of database: 5,428,055
effective search space: 1422150410
effective search space used: 1422150410
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC148607.14


