

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148532.15 + phase: 1 /partial
(178 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
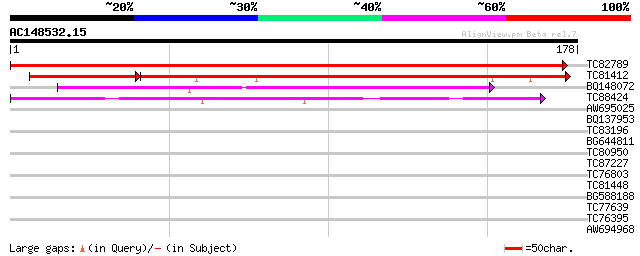
Score E
Sequences producing significant alignments: (bits) Value
TC82789 363 e-101
TC81412 weakly similar to PIR|T48506|T48506 hypothetical protein... 150 4e-50
BQ148072 weakly similar to GP|12382016|dbj hypothetical protein ... 115 9e-27
TC88424 similar to GP|22136724|gb|AAM91681.1 unknown protein {Ar... 42 2e-04
AW695025 similar to GP|2865416|gb|A chromomethylase {Arabidopsis... 37 0.004
BQ137953 weakly similar to GP|22136724|gb unknown protein {Arabi... 31 0.23
TC83196 similar to GP|7270830|emb|CAB80511.1 protein kinase like... 30 0.39
BG644811 similar to GP|15866773|gb| origin recognition complex s... 30 0.66
TC80950 similar to GP|15187169|gb|AAK91319.1 Unknown protein {Or... 30 0.66
TC87227 similar to GP|17064926|gb|AAL32617.1 calcium-dependent p... 28 1.9
TC76803 PIR|T09622|T09622 protein kinase MMK4 (EC 2.7.1.-) cold... 27 4.3
TC81448 weakly similar to OMNI|NT01MC3624 toxin secretion ATP-bi... 26 7.3
BG588188 weakly similar to GP|20260382|gb| unknown protein {Arab... 26 7.3
TC77639 similar to GP|10178280|emb|CAC08338. alpha-galactosidase... 26 9.6
TC76395 similar to SP|Q9ATF5|RL1X_CASSA Ribosomal protein L18a. ... 26 9.6
AW694968 similar to PIR|T48518|T485 transcription factor like pr... 26 9.6
>TC82789
Length = 841
Score = 363 bits (933), Expect = e-101
Identities = 174/175 (99%), Positives = 174/175 (99%)
Frame = +3
Query: 1 KSNSPPAPSPESETLEFKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLFDNVYVKNDFAEP 60
KSNSPPAPSPESETLEFKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLFDNVYVKNDFAEP
Sbjct: 84 KSNSPPAPSPESETLEFKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLFDNVYVKNDFAEP 263
Query: 61 RIGKIIKIWETPTLERKIKVQWFFRPIEVSKCLTWIKIYFNELFFACGDGDGLATIHPLE 120
RIGKIIKIWETPTLERKIKVQWFFRPIEVSKCLTWIKIYFNELFFACGDGDGLATIHPLE
Sbjct: 264 RIGKIIKIWETPTLERKIKVQWFFRPIEVSKCLTWIKIYFNELFFACGDGDGLATIHPLE 443
Query: 121 SIAGKCNIVCISKDSRNPQLSDKVTWSAAFVFYRYFDVGQRKIVEEVDDKSIGIE 175
SIAGKCNIVCISKDSRNPQLSDKVTWSAAFVFYRYFD GQRKIVEEVDDKSIGIE
Sbjct: 444 SIAGKCNIVCISKDSRNPQLSDKVTWSAAFVFYRYFDDGQRKIVEEVDDKSIGIE 608
>TC81412 weakly similar to PIR|T48506|T48506 hypothetical protein F15N18.60
- Arabidopsis thaliana, partial (14%)
Length = 716
Score = 150 bits (380), Expect(2) = 4e-50
Identities = 83/143 (58%), Positives = 103/143 (71%), Gaps = 8/143 (5%)
Frame = +3
Query: 42 GVDYSLFDNVYVKNDF---AEPRIGKIIKIWETPTLER--KIKVQWFFRPIEVSKCLTWI 96
G DYSLFD VY++N +EP IGKIIKIWE P E+ K+K+QWFFRP EVSK L I
Sbjct: 120 GEDYSLFDTVYLQNGNEPQSEPHIGKIIKIWEVPNREKAKKVKIQWFFRPREVSKFLKRI 299
Query: 97 KIYFNELFFACGDGDGLATIHPLESIAGKCNIVCISKDSRNPQLSDKVTWSAAF-VFYRY 155
+IY+NELFFA G G+GL I+P+ESIAGKCN+VCISKD+RNPQ SD+ +A V
Sbjct: 300 QIYYNELFFATGVGNGLTNINPVESIAGKCNVVCISKDARNPQPSDEAVQNADLCVLSLP 479
Query: 156 FDVGQRK--IVEEVDDKSIGIEG 176
+G K +EVD+KS+GI+G
Sbjct: 480 LMLGSVK*WRRKEVDEKSVGIDG 548
Score = 63.9 bits (154), Expect(2) = 4e-50
Identities = 27/35 (77%), Positives = 30/35 (85%)
Frame = +2
Query: 7 APSPESETLEFKWGQKRGKGGKKRDIQFYQSFTLR 41
+P P ETLEFKWG+KRGKGGKKRD QFY+SFT R
Sbjct: 14 SPPPSPETLEFKWGKKRGKGGKKRDTQFYESFTFR 118
>BQ148072 weakly similar to GP|12382016|dbj hypothetical protein {Oryza
sativa (japonica cultivar-group)}, partial (7%)
Length = 673
Score = 115 bits (288), Expect = 9e-27
Identities = 63/138 (45%), Positives = 81/138 (58%), Gaps = 1/138 (0%)
Frame = +2
Query: 16 EFKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLFDNVYVKN-DFAEPRIGKIIKIWETPTL 74
+FKWG KRG G K +D QFY SF GV+Y L D VY + D E IG ++K++E
Sbjct: 254 DFKWGNKRGIGVKNKDTQFYDSFVYEGVEYFLHDCVYFYHTDHVETSIGMLVKMFENGR- 430
Query: 75 ERKIKVQWFFRPIEVSKCLTWIKIYFNELFFACGDGDGLATIHPLESIAGKCNIVCISKD 134
+ I+V WFFRP E+ + +NELF A G G GL ++ +ESI GKC IVC S D
Sbjct: 431 RKMIRVVWFFRPSEIRSFHRSYQPSWNELFLASGKGKGLTNVNSVESILGKCCIVCSSXD 610
Query: 135 SRNPQLSDKVTWSAAFVF 152
RNP+ S+ A F F
Sbjct: 611 KRNPKPSETELKRANFFF 664
>TC88424 similar to GP|22136724|gb|AAM91681.1 unknown protein {Arabidopsis
thaliana}, partial (50%)
Length = 1635
Score = 41.6 bits (96), Expect = 2e-04
Identities = 45/172 (26%), Positives = 71/172 (41%), Gaps = 4/172 (2%)
Frame = +2
Query: 1 KSNSPPAPSPESETLEFKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLFDNVYVKNDFAE- 59
K +SP P E +G+G K+ Y+SF G YSL D V + + E
Sbjct: 212 KKDSPQTPEDAKPIGEPLRVSGKGRGRKRH----YESFEFDGNQYSLEDPVMLVPEDKEQ 379
Query: 60 -PRIGKIIKIWETPTLERKIKVQWFFRPIEVSK--CLTWIKIYFNELFFACGDGDGLATI 116
P + I I + + + QWF+RP E K +W ELF++ +
Sbjct: 380 KPYVAIIKDIIQYFSGSIMVAGQWFYRPEEAEKKGGGSWKSCDTRELFYSFHRDE----- 544
Query: 117 HPLESIAGKCNIVCISKDSRNPQLSDKVTWSAAFVFYRYFDVGQRKIVEEVD 168
P ES+ KC + + + + P K F+ R +D +RK+ + D
Sbjct: 545 VPAESVMHKCVVHFVPLNKQFP----KRKQHPGFIVQRVYDTLERKLWKLTD 688
>AW695025 similar to GP|2865416|gb|A chromomethylase {Arabidopsis thaliana},
partial (2%)
Length = 652
Score = 37.0 bits (84), Expect = 0.004
Identities = 31/101 (30%), Positives = 47/101 (45%), Gaps = 1/101 (0%)
Frame = +2
Query: 35 YQSFTLRGVDYSLFDNVYVKNDFA-EPRIGKIIKIWETPTLERKIKVQWFFRPIEVSKCL 93
Y+ + GV Y L DN YVK + E I I++++ T E QWF+R +
Sbjct: 125 YREVNVDGVIYKLNDNAYVKGEEGKENYIAAIVELFVTHENEYYFTAQWFYRAED----- 289
Query: 94 TWIKIYFNELFFACGDGDGLATIHPLESIAGKCNIVCISKD 134
T IK + + + + +PL+ + K NIV IS D
Sbjct: 290 TVIKNHGDLIDEKRIFKSDVKDENPLDCLVKKVNIVQISPD 412
>BQ137953 weakly similar to GP|22136724|gb unknown protein {Arabidopsis
thaliana}, partial (21%)
Length = 657
Score = 31.2 bits (69), Expect = 0.23
Identities = 26/89 (29%), Positives = 43/89 (48%), Gaps = 6/89 (6%)
Frame = +2
Query: 24 GKGGKKRDIQFYQSFTLRGVDYSLFDNVYVKNDFAE--PRIGKIIKIWETPTLERKIKV- 80
GKG +++ +QSF G YSL D V + + + P + I I ++ + +
Sbjct: 368 GKGRGRKN--HFQSFDFDGDQYSLEDPVLLVPEVKDQKPYVAIIKDITQSINGNGSLMIT 541
Query: 81 -QWFFRPIEVSK--CLTWIKIYFNELFFA 106
QWF+RP E K +W + ELF++
Sbjct: 542 GQWFYRPDEAEKKGGGSWQSVDTRELFYS 628
>TC83196 similar to GP|7270830|emb|CAB80511.1 protein kinase like protein
{Arabidopsis thaliana}, partial (66%)
Length = 1484
Score = 30.4 bits (67), Expect = 0.39
Identities = 15/63 (23%), Positives = 27/63 (42%), Gaps = 2/63 (3%)
Frame = -2
Query: 70 ETPTLERKIKVQWFFRPIEVSKCLTWIKIYFNELFFACGDGDGLATIHPLESIAGK--CN 127
E P++ K W F P SK + W++ + GL+++ + S G+ C
Sbjct: 652 ENPSVVEKACASWMFSPTSASKAVNWLRSF------------GLSSVETVISCMGREYCT 509
Query: 128 IVC 130
+ C
Sbjct: 508 LAC 500
>BG644811 similar to GP|15866773|gb| origin recognition complex subunit 1
{Zea mays}, partial (4%)
Length = 784
Score = 29.6 bits (65), Expect = 0.66
Identities = 12/30 (40%), Positives = 19/30 (63%)
Frame = +3
Query: 25 KGGKKRDIQFYQSFTLRGVDYSLFDNVYVK 54
KGGK +Q+Y+ G ++ + D+VYVK
Sbjct: 591 KGGKIAKVQYYKKVVYDGGEFEVGDDVYVK 680
>TC80950 similar to GP|15187169|gb|AAK91319.1 Unknown protein {Oryza
sativa}, partial (15%)
Length = 941
Score = 29.6 bits (65), Expect = 0.66
Identities = 13/31 (41%), Positives = 18/31 (57%)
Frame = +2
Query: 118 PLESIAGKCNIVCISKDSRNPQLSDKVTWSA 148
P+ S GK + C +D NPQLSD ++ A
Sbjct: 539 PMPSAGGKPPLKCTCRDGENPQLSDANSFRA 631
>TC87227 similar to GP|17064926|gb|AAL32617.1 calcium-dependent protein kinase
{Arabidopsis thaliana}, partial (95%)
Length = 2319
Score = 28.1 bits (61), Expect = 1.9
Identities = 14/35 (40%), Positives = 20/35 (57%), Gaps = 2/35 (5%)
Frame = +3
Query: 69 WETPTLERKIKVQWFF--RPIEVSKCLTWIKIYFN 101
WE LE K+ +WFF R VS+CL ++F+
Sbjct: 1731 WEIQ*LELKVNERWFFKLRK*LVSECLLIFNLFFH 1835
>TC76803 PIR|T09622|T09622 protein kinase MMK4 (EC 2.7.1.-) cold- and
drought-induced - alfalfa, complete
Length = 1709
Score = 26.9 bits (58), Expect = 4.3
Identities = 11/48 (22%), Positives = 23/48 (47%)
Frame = -2
Query: 77 KIKVQWFFRPIEVSKCLTWIKIYFNELFFACGDGDGLATIHPLESIAG 124
K+K +W +S + + K++ ++ FF C + H ++ I G
Sbjct: 1150 KLK*EWLHADRFISYVMQFFKVWMSQCFFNCNSSGRINCQHFIDKING 1007
>TC81448 weakly similar to OMNI|NT01MC3624 toxin secretion ATP-binding
protein putative {Magnetococcus sp. MC-1}, partial (4%)
Length = 655
Score = 26.2 bits (56), Expect = 7.3
Identities = 9/24 (37%), Positives = 17/24 (70%)
Frame = -2
Query: 30 RDIQFYQSFTLRGVDYSLFDNVYV 53
R ++Y F +G+D+++F NVY+
Sbjct: 627 RSRRYYYLFFKQGIDHNIFYNVYI 556
>BG588188 weakly similar to GP|20260382|gb| unknown protein {Arabidopsis
thaliana}, partial (6%)
Length = 670
Score = 26.2 bits (56), Expect = 7.3
Identities = 13/38 (34%), Positives = 20/38 (52%)
Frame = -3
Query: 70 ETPTLERKIKVQWFFRPIEVSKCLTWIKIYFNELFFAC 107
ET ++ KIK PI S L+ ++ NE++ AC
Sbjct: 644 ETIKIK*KIKKAMMVHPIRTSANLSLHSLFLNEMWVAC 531
>TC77639 similar to GP|10178280|emb|CAC08338. alpha-galactosidase-like
protein {Arabidopsis thaliana}, partial (90%)
Length = 1442
Score = 25.8 bits (55), Expect = 9.6
Identities = 13/41 (31%), Positives = 22/41 (52%)
Frame = +3
Query: 22 KRGKGGKKRDIQFYQSFTLRGVDYSLFDNVYVKNDFAEPRI 62
K+ G + Q ++F G+DY +DN Y ND ++P +
Sbjct: 468 KQMPGSLGHEFQDAKTFASWGIDYLKYDNCY--NDESKPTV 584
>TC76395 similar to SP|Q9ATF5|RL1X_CASSA Ribosomal protein L18a. [Sweet
chestnut] {Castanea sativa}, complete
Length = 1640
Score = 25.8 bits (55), Expect = 9.6
Identities = 22/92 (23%), Positives = 38/92 (40%), Gaps = 5/92 (5%)
Frame = +3
Query: 18 KWGQKRGKGGKKR-----DIQFYQSFTLRGVDYSLFDNVYVKNDFAEPRIGKIIKIWETP 72
K Q+RG K R ++ +Q T + Y + P+I ++ K+W T
Sbjct: 816 KRDQRRGLQDKLRHNEGAELLPFQMATFKYHQYQVVGRALPTEKDEHPKIYRM-KLWATN 992
Query: 73 TLERKIKVQWFFRPIEVSKCLTWIKIYFNELF 104
+ K K +F R ++ K + NE+F
Sbjct: 993 EVRAKSKFWYFLRKLKKVKKSNGQVLAINEIF 1088
>AW694968 similar to PIR|T48518|T485 transcription factor like protein -
Arabidopsis thaliana, partial (37%)
Length = 590
Score = 25.8 bits (55), Expect = 9.6
Identities = 9/15 (60%), Positives = 13/15 (86%)
Frame = +2
Query: 2 SNSPPAPSPESETLE 16
SNSPP PSP+S+ ++
Sbjct: 89 SNSPPPPSPQSQQIK 133
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.140 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,640,369
Number of Sequences: 36976
Number of extensions: 103370
Number of successful extensions: 517
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 507
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 512
length of query: 178
length of database: 9,014,727
effective HSP length: 90
effective length of query: 88
effective length of database: 5,686,887
effective search space: 500446056
effective search space used: 500446056
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC148532.15


